Abstract
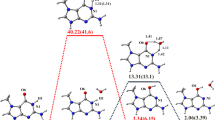
DNA hydroxymethylation plays a very important role in some biological processes, such as DNA methylation process. In addition, its presence can also cause some diseases. In this paper, the electrical properties of cytosine hydroxymethylated (Chm) DNA sequences are studied. The density functional theory (DFT) and Landauer–Büttiker framework are used to study the decoherence conductance and transmission of the Chm strands in different configurations, which provides a theoretical basis for the detection of Chm. The results show that the conductance of the hydroxymethylated DNA strand is smaller than that of the native and methylated strands. The length dependence of the Chm strands is also studied. With the length increasing, the conductance becomes larger. This study shows that DNA methylation can be detected electrically.



Similar content being viewed by others
Data availability
The data used to support the funding of this study are included within the article. The data and materials in the current study are available from the corresponding author on reasonable requests.
References
Arkin MR, Stemp EDA, Holmlin RE, Barton JK, Hormann A, Olson EJC (1996) PF Rates of DNA-mediated electron transfer between metallointercalators. Science 273:475–480. https://doi.org/10.1126/science.273.5274.475
Fink HW, Schnenberger C (1999) Electrical conduction through DNA molecules. Nature 398:407–410. https://doi.org/10.1038/18855
Yoo KH, Ha DH, Lee JO, Park JW, Kim J, Kim JJ, Lee HY, Kawai T, Choi HY (2001) Electrical Conduction through Poly(dA)-Poly(dT) and Poly(dG)-Poly(dC) DNA Molecules. Phys Rev Lett 87:198102. https://doi.org/10.1103/PhysRevLett.87.198102
de Pablo PJ, Moreno-Herrero F, Colchero J, Gomez-Herrero J, Herrero P, Baro AM, Ordejon P, Soler JM, Ec A (2000) Absence of dc-conductivity in lambda-DNA. Phys Rev Lett 85:4992. https://doi.org/10.1103/PhysRevLett.85.4992
Macfarlane RJ, Lee B, Hill HD, Senesi AJ, Seifert S, Mirkin CA (2009) Molecular recognition and self-assembly special feature Assembly and organization processes in DNA-directed colloidal crystallization. Proc Natl Acad Sci USA 106:10493. https://doi.org/10.1073/pnas.0900630106
Cuevas JC, Scheer E (2010) Molecular electronics : an introduction to theory and experiment. World Scientific. https://doi.org/10.1142/7434
Wang ZG, Wu HH, Li Q, Besenbacher F, Li YR, Zeng XC, Dong MD (2020) Reversing Interfacial Catalysis of Ambipolar Wse2 Single Crystal. Advanced Science 7:1901382. https://doi.org/10.1002/advs.201901382
Wang ZG, Li Q, Chen YF, Cui BX, Li YR, Besenbacher F, Dong MD (2018) The ambipolar transport behavior of WSe2 transistors and its analogue circuits. NPG Asia Materials 10:703–712. https://doi.org/10.1038/s41427-018-0062-1
Guo AM, Yang Z, Zhu HJ, Xiong SJ (2010) Influence of backbone on the charge transport properties of G4-DNA molecules: a model-based calculation. J Phys: Condens Matter 22:065102. https://doi.org/10.1088/0953-8984/22/6/065102
Wang XF, Chakraborty T, Berashevich J (2010) Quantum transport anomalies in DNA containing mispairs. Nanotechnology 21:485101. https://doi.org/10.1088/0957-4484/21/48/485101
Edirisinghe N, Apalkov V, Berashevich J, Chakraborty T (2010) Electrical current through DNA containing mismatched base pairs. Nanotechnology 21:245101. https://doi.org/10.1088/0957-4484/21/24/245101
Teschome B, Facsko S, Schönherr T, Kerbusch J, Keller A, Erbe A (2016) Temperature-dependent charge transport through individually contacted DNA origami-based Au nanowires. Langmuir 32:10159–10165. https://doi.org/10.1021/acs.langmuir.6b01961
Kypr J, Kejnovska I, Renciuk D, Vorlickova M (2009) Circular dichroism and conformational polymorphism of DNA. Nucleic Acids Res 37:1713–1725. https://doi.org/10.1093/nar/gkp026
Wolter M, Elstner M, Kubar T (2013) Charge transport in desolvated DNA. Journal of Chemical Physics 139:125102. https://doi.org/10.1063/1.4821594
Qi J, Anantram MP (2014a) Charge Transport through Methylated DNA Strand. J Chem Phys 143(9):094306. https://doi.org/10.1063/1.4929909
Fernandez AF, Bayón GF, Sierra MI, Urdinguio RG, Torano E, Garcia MG, Carella A, Lopez V, Santamarina P, Perez RF (2018) Loss of 5hmC identifies a new type of aberrant DNA hypermethylation in glioma. Hum Mol Genet 27:3046–3059. https://doi.org/10.1093/hmg/ddy214
Shi DQ, Iftikhar A, Tang J, Yang WC (2017) New Insights into 5hmC DNA Modification: Generation, Distribution and Function. Frontiers in Genetics 8:100–107. https://doi.org/10.3389/fgene.2017.00100
Sun W, Zang L, Shu Q, Li X (2014) From development to diseases: The role of 5hmC in brain. Genomics 104:347–351. https://doi.org/10.1016/j.ygeno.2014.08.021
Macke TJ, Case DA (1998) Modeling unusual nucleic acid structures. Am Chem Soc 682:379–393. https://doi.org/10.1021/bk-1998-0682.ch024
Flores-Moreno R, Posada E, Moncada F, Romero J, Charry J, Diaz-Tinoco M, Gonzalez SA, Aguirre NF, Reyes A (2014) LOWDIN: The any particle molecular orbital code. Int J Quantum Chem 114:50–56. https://doi.org/10.1002/qua.24500
Qi J, Edirisinghe N, Rabbani G, Anantram MP (2013) Unified model for conductance through DNA with the Landauer-Buttiker formalism. Phys Rev 8:109–115. https://doi.org/10.1103/PhysRevB.87.085404
Qi J, Anantram MP (2014b) Electron Flow in GC8 DNA Strand. ECS Trans 64:1–9. https://doi.org/10.1149/06412.0001ecst
Funding
This work was supported by the China Scholarship Council Study Program for Young Backbone Teachers under Grant No. 201707845017, the National Nature Science Foundation of China under Grant No. 61804020, the Scientific and Technological Research Foundation of Chongqing Municipal Education Commission under Grant No. KJQN201900643, the Raised Fund Plan of National Natural Science Foundation of China of Chongqing University of Posts and Telecommunications under Grant No. A2016-143, the PhD scientific research foundation of Chongqing University of Posts and Telecommunications under Grant Nos. A2015-11 and A2015-12, and SR Patil’s support from SERB, India under Grant No. TAR/2018/001188.
Author information
Authors and Affiliations
Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agreed to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
He, L., Zhang, J., He, C. et al. Effect of cytosine hydroxymethylation on DNA charge transport. Mol Cell Biochem 476, 1599–1603 (2021). https://doi.org/10.1007/s11010-020-03957-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-020-03957-7