Abstract
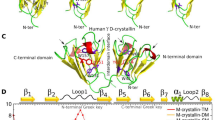
The crystallins are a family of monomeric proteins present in the mammalian lens and mutations in these proteins cause various forms of cataracts. The aim of our current study is to emphasize the structural characterization of aggregation propensity of mutation R58H on γD crystallin using molecular dynamics (MD) approach. MD result revealed that difference in the sequence level display a wide variation in the backbone atomic position, and thus exhibits rigid conformational dynamics. Changes in the flexibility of residues favoured to increase the number of intra-molecular hydrogen bonds in mutant R58H. Moreover, notable changes in the hydrogen bonding interaction resulted to cause the misfolding of mutant R58H by introducing α-helix. Principal component analysis (PCA) result suggested that mutant R58H showed unusual conformational dynamics along the two principal components when compared to the wild-type (WT)-γD crystallin. In a nutshell, the increased surface hydrophobicity could be the cause of self-aggregation of mutant R58H leading to aculeiform cataract.







Similar content being viewed by others
Abbreviations
- RMSD:
-
Root-mean-square deviation
- RMSF:
-
Root-mean-square fluctuation
- ASA:
-
Accessible surface area
- DSSP:
-
Dictionary of secondary structure of protein
- PCA:
-
Principal component analysis
- FEL:
-
Free energy landscape
References
Ji F, Jung J, Gronenborn AM (2012) Structural and biochemical characterization of the childhood cataract-associated R76S mutant of human gD-crystallin. Biochemistry 51:2588–2596
Zhenzhen L, Allen T, Yizhi L, Mingxing W, Xiaohua G, Fu S (2012) Enhancement of ubiquitin conjugation activity reduces intracellular aggregation of V76D mutant γD-Crystallin. Invest Ophthalmol 53(10):6655–6665
Moran SD, Woys AM, Buchanan LE, Bixby E, Decatur SM, Zanni MT (2012) Two-dimensional IR spectroscopy and segmental 13C labeling reveals the domain structure of human γD-crystallin amyloid fibrils. Proc Natl Acad Sci USA 109:3329–3334
Moreau KL, King JA (2012) Protein misfolding and aggregation in cataract disease and prospects for prevention. Trends Mol Med 18:273–282
Hains PG, Truscott RJ (2007) Post-translational modifications in the nuclear region of young, aged, and cataract human lenses. J Proteome Res 6:3935–3943
Hains PG, Truscott RJ (2008) Proteomic analysis of the oxidation of cysteine residues in human age-related nuclear cataract lenses. Biochim Biophys Acta 1784:1959–1964
Lampi KJ, Ma Z, Hanson SR, Azuma M, Shih M, Shearer TR, Smith DL (1998) Age-related changes in human lens crystallins identified by two-dimensional electrophoresis and mass spectrometry. Exp Eye Res 67:31–43
Ma Z, Hanson SR, Lampi KJ, David LL, Smith DL, Smith JB (1998) Age-related changes in human lens crystallins identified by HPLC and mass spectrometry. Exp Eye Res 67:21–30
Zhang Z, Smith DL, Smith JB (2003) Human beta-crystallins modified by backbone cleavage, deamidation and oxidation are prone to associate. Exp Eye Res 77:259–272
Wilmarth PA, Tanner S, Dasari S, Nagalla SR, Riviere MA, Bafna V, Pevzner PA, David LL (2006) Age-related changes in human crystallins determined from comparative analysis of post-translational modifications in young and aged lens: does deamidation contribute to crystallin insolubility? J Proteome Res 5:2554–2566
Hains PG, Truscott RJ (2010) Age-dependent deamidation of lifelong proteins in the human lens. Invest. Ophthalmol Vis Sci 51:3107–3114
Srikumar PS, Rohini K, Rajesh PK (2014) Molecular dynamics simulations and principal component analysis on human laforin mutation W32G and W32G/K87A. Protein J 33(3):289–295
Rohini K, Srikumar PS (2013) Structural insights on Mycobacterium tuberculosis thiazole synthase—A molecular dynamics/docking approach. Appl Biochem Biotechnol 169(6):1790–1798
Ji F, Jung J, Koharudin LM, Gronenborn AM (2013) The human W42R γD-crystallin mutant structure provides a link between congenital and age-related cataracts. J Biol Chem 288:99–109
Lee S, Mahler B, Toward J, Jones B, Wyatt K, Dong L, Wistow G, Wu Z (2010) A single destabilizing mutation (F9S) promotes concerted unfolding of an entire globular domain in gamma S-crystallin. J Mol Biol 399:320–330
Rajendran V, Gopalakrishnan C, Sethumadhavan R (2017) Pathological role of a point mutation (T315I) in BCR-ABL1 protein-A computational insight. J Cell Biochem 119(1):918–925
Kamaraj B, Purohit R (2016) Mutational analysis on membrane associated transporter protein (MATP) and their structural consequences in oculocutaeous albinism type 4 (OCA4)-a molecular dynamics approach. J Cell Biochem 117(11):2608–2619
Rajendran V (2016) Structural analysis of oncogenic mutation of isocitrate dehydrogenase 1. Mol Biosyst 2(7):2276–2287
Rajendran V, Purohit R, Sethumadhavan R (2012) In silico investigation of molecular mechanism of laminopathy caused by a point mutation (R482W) in lamin A/C protein. Amino Acids 43(2):603–615
Berman HM, Westbrook J, Feng Z, Gilliland GG, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Berendsen K, Van der Spoel HJC, Van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56
Kaminski M, Friesner GA, Tirado-Rives RA, Jorgensen LW (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J Phys Chem B 105:6474–6487
Gopalakrishnan C, Kalsi N, Jethi S, Purohit R (2015) Computational investigation of molecular mechanism and neuropathological implications in Huntington disease. Mol Cell Biochem 409(1–2):1–11
Darden N, York T, Pedersen L (1993) Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Anbarasu K, Jayanthi S (2014) Structural modeling and molecular dynamics studies on the human LMTK3 domain and the mechanism of ATP binding. Mol Biosyst 10:1139–1145
Purohit R (2014) Role of ELA region in auto-activation of mutant KIT receptor: a molecular dynamics simulation insight. J Biomol Struct Dyn 32(7):1033–1046
Haider S, Parkinson GN, Neidle S (2008) Molecular dynamics and principal components analysis of human telomeric quadruplex multimers. Biophys J 95:296–311
Chandrasekaran P, Rajasekaran R (2016) A systematic molecular dynamics approach to the structural characterization of amyloid aggregation propensity of β2-microglobulin mutant D76N. Mol Biosyst 12:850–859
Rajendran V, Gopalakrishnan C, Purohit R (2016) Impact of point mutation P29S in RAC1 on tumorigenesis. Tumour Biol 37(11):15293–15304
Batista PR, Costa MG, Pascutti PG, Bisch PM, De Souza W (2011) High temperatures enhance cooperative motions between CBM and catalytic domains of a thermostable cellulase: mechanism insights from essential dynamics. Phys Chem Chem Phys 13:13709–13720
Wieczorek G, Zielenkiewicz P (2008) DeltaF508 mutation increases conformational flexibility of CFTR protein. J Cyst Fibros 7:295–300
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637
Lambrughi M, Papaleo E, Testa L, Brocca S, De Gioia L, Grandori R (2012) Intramolecular interactions stabilizing compact conformations of the intrinsically disordered kinase-inhibitor domain of Sic1: a molecular dynamics investigation. Front Physiol 3:435
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Karunakaran, R., Srikumar, P.S. A molecular dynamics approach to explore the structural characterization of cataract causing mutation R58H on human γD crystallin. Mol Cell Biochem 449, 55–62 (2018). https://doi.org/10.1007/s11010-018-3342-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-018-3342-8