Abstract
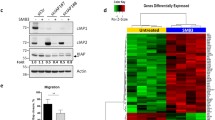
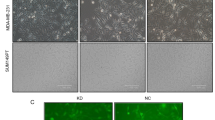
p53 protein is probably the best known tumor suppressor. Earlier reports proved that human breast cancer cells expressing mutant p53 displayed resistance to apoptosis. This study is intended to investigate, the potential applications of RNA interference (RNAi) to block p53 expression, as well as its subsequent effect on cell growth, apoptosis and migration on a triple negative human breast cancer cell line (Hs578T). p53siRNA significantly reduced cell index (CI) compared to the control and we observed an inhibition of cellular migration in the interval of time between 0 and 30 h, as shown in the data obtained by dynamic evaluation using the xCELLigence System. Also, by using PCR-array technology, a panel of 84 key genes involved in apoptosis was investigated. Our studies indicate that the knockdown of p53 expression by siRNA modulates several genes involved in cell death pathways and apoptosis, showing statistically significant gene expression differences for 22 genes, from which 18 were upregulated and 4 were downregulated. The present research also emphasizes the important role of BCL-2 pro-apoptotic family of genes (Bim, Bak, and Bax) in activating apoptosis and reducing cell proliferation by p53siRNA treatment. Death receptors cooperate with BCL-2 pro-apoptotic genes in reducing cell proliferation. The limited success may be due to the activation of the antiapoptotic gene Mcl-1, and it may be associated with the resistance of triple negative breast cancer cells to cancer treatment. Thus, targeting p53siRNA pathways using siRNA may serve as a promising therapeutic strategy for the treatment of breast cancers.




Similar content being viewed by others
References
Wang Z, Sun Y (2010) Targeting p53 for novel anticancer therapy. Transl Oncol 3(1):1–12
Finlay CA, Hinds PW, Levine AJ (1989) The p53 proto-oncogene can act as a suppressor of transformation. Cell 57(7):1083–1093
Bishayee A, Ahmed S, Brankov N, Perloff M (2011) Triterpenoids as potential agents for the chemoprevention and therapy of breast cancer. Front Biosci 16:980–996
Rahman M, Davis SR, Pumphrey JG, Bao J, Nau MM, Meltzer PS, Lipkowitz S (2009) TRAIL induces apoptosis in triple-negative breast cancer cells with a mesenchymal phenotype. Breast Cancer Res Treat 113(2):217–230
Liu Y-Y, Patwardhan G, Bhinge K, Gupta V, Gu X, Jazwinski M (2011) Suppression of glucosylceramide synthase restores p53-dependent apoptosis in mutant p53 cancer cells. Cancer Res 71(6):2276–2285
Norberg T, Klaar S, Karf G, Nordgren H, Holmberg L, Bergh J (2001) Increased p53 mutation frequency during tumor progression––results from a breast cancer cohort. Cancer Res 61:8317–8321
Sigal A, Rotter V (2000) Oncogenic mutations of the p53 tumor suppressor: the demons of the guardian of the genome. Cancer Res 60:6788–6793
Vikhanskaya F, Lee K, Mazzoletti M, Broggini M, Sabapathy K (2007) Cancer-derived p53 mutants suppress p53-target gene expression—potential mechanism for gain of function of mutant p53. Nucleic Acids Res 35(6):2093–2104
Kim DH, Rossi JJ (2007) Strategies for silencing human disease using RNA interference. Nat Rev Genet 8:173–184
Brummelkamp TR, Bernards R, Agami R (2002) Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell 2:243–247
Kim DH, Rossi JJ (2007) Strategies for silencing human disease using RNA interference. Nat Rev Genet 8:173–184
Rye PD, Stigbrand T (2004) Interfering with cancer: A brief outline of advances in RNA interference in oncology. Tumor Biology 25:5–6
Lacroix M, Toillon RA, Leclercq G (2006) p53 and breast cancer, an update. Endocr Relat Cancer 13(2):293–325
Gasco M, Shami S, Crook T (2002) The p53 pathway in breast cancer. Breast Cancer Res 4:70–76
Ma CX, Cai S, Li S, Ryan CE, Guo Z, Schaiff WT, Lin L, Hoog J, Goiffon RJ, Prat A, Aft RL, Ellis MJ, Piwnica-Worms H (2012) Targeting Chk1 in p53-deficient triple-negative breast cancer is therapeutically beneficial in human-in-mouse tumor Models. J Clin Invest 122(4):1541–1552
Zhao R, Gish K, Murphy M, Yin Y, Notterman D, Hoffman WH, Tom E, Mack DH, Levine AJ (2000) Analysis of p53-regulated gene expression patterns using oligonucleotide arrays. Genes Dev 14:981–993
Zhou M, Liu Z, Zhao Y, Ding Y, Liu H, Xi Y, Xiong W, Li G, Lu J, Fodstad O, Riker AI, Tan M (2010) MicroRNA-125b confers the resistance of breast cancer cells to paclitaxel through suppression of pro-apoptotic Bcl-2 antagonist killer 1 (Bak1) expression. J Biol Chem 285(28):21496–21507
Scott GB, Bowles PA, Wilson EB, Meade JL, Low BC, Davison A, Davison A, Blair GE, Cook GP (2010) Identification of the BCL2/adenovirus E1B–19K protein-interacting protein 2 (BNIP-2) as a granzyme B target during human natural killer cell-mediated killing. Biochem J 431:423–431
Morel C, Carlson SM, White FM, Davis RJ (2009) Mcl-1 integrates the opposing actions of signaling pathways that mediate survival and apoptosis. Mol Cell Biol 29(14):3845–3852
Plati J, Bucur O, Khosravi-Far R (2011) Apoptotic cell signaling in cancer progression and therapy. Integr Biol (Camb) 3(4):279–296
Bruey JM, Bruey-Sedano N, Luciano F, Zhai D, Balpai R, Xu C, Kress CL, Bailly-Maitre B, Li X, Osterman A, Mastsuzawa S, Terskikh AV, Faustin B, Reed JC (2007) Bcl-2 and Bcl-XL regulate proinflammatory caspase-1 activation by interaction with NALP1. Cell 129:45–56
Aggarwal BB, Shishodia S, Sandur SK, Pandey MK, Sethi G (2006) Inflammation and cancer: how hot is the link? Biochem Pharmacol 72(11):1605–1621
Elmore S (2007) Apoptosis: a review of programmed cell death. Toxicol Pathol 35(4):495–516
Booy EP, Henson ES, Gibson SB (2011) Epidermal growth factor regulates Mcl-1 expression through the MAPK-Elk-1 signalling pathway contributing to cell survival in breast cancer. Oncog 30:2367–2378
Woods NT, Yamaguchi H, Lee FY, Bhalla KN (2007) Wang Anoikis HG. initiated by Mcl-1 degradation and Bim induction, is deregulated during oncogenesis. Cancer Res 67:10744
Qin W, Hu J, Guo M, Xu J, Li J, Yao G et al (2003) BNIPL-2, a novel homologue of BNIP-2, interacts with Bcl-2 and Cdc42GAP in apoptosis. Biochem Biophys Res Commun 308:379–385
Debily MA, Marhomy SE, Boulanger V, Eveno E, Mariage-Samson R, Camarca A et al (2009) A functional and regulatory network associated with PIP expression in human breast cancer. PLoS One 4(3):4696
Lu Y, Chen GQ (2011) Effector caspases and leukemia. Int J of Cell Biol. doi:10.1155/2011/738301
Valentín-Acevedo A, Sinquett FL, Covey LR (2011) c-Rel deficiency increases caspase-4 expression and leads to ER stress and necrosis in EBV-transformed cells. PLoS One 6(10):25467
Smolnikar K, Löffek S, Schulz T, Michna H, Diel P (2000) Treatment with the pure antiestrogen faslodex (ICI 182780) induces tumor necrosis factor receptor 1 (TNFR1) expression in MCF-7 breast cancer cells. Breast Cancer Res Treat 63(3):249–259
Adams JM, Cory S (2007) The Bcl-2 apoptotic switch in cancer development and therapy Bcl-2 apoptotic switch in cancer. Oncog 26:1324–1337
Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A (2005) An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. PNAS 2(38):13550–13555
Acknowledgments
This study was partially financed by a grant from the Romanian National University Research Council project PD 533/28.07.2010 “Combining chemotherapeutic effects of flavan-3-ols with RNA interference target therapy in cancer” and partially by a POSCCE 709/2010 Grant with title: “Clinical and economical impact of proteome and transcriptome molecular profiling in neoadjuvant therapy of triple negative breast cancer (BREAST IMPACT)”.
Conflict of interest
The authors report no conflicts of interest in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Braicu, C., Pileczki, V., Irimie, A. et al. p53siRNA therapy reduces cell proliferation, migration and induces apoptosis in triple negative breast cancer cells. Mol Cell Biochem 381, 61–68 (2013). https://doi.org/10.1007/s11010-013-1688-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-013-1688-5