Abstract
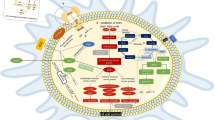
Knowledge of virulence factors is important to understand the microbial pathogenesis and find better antibiotics. Mammalian cell entry (mce) is a crucial protein family for the virulence of Mycobacterium tuberculosis (M. tuberculosis). This review summarized the advances on mce genes. The genomic organization, characteristics of mce genes, phylogeny of this family, and their roles in M. tuberculosis virulence are emphasized in this review.






Similar content being viewed by others
Abbreviations
- HIV:
-
Human immunodeficiency virus
- SBP:
-
Substrate-binding proteins
- ABC:
-
ATP-binding cassette
- CHP:
-
Conserved hypothetical protein
References
Flynn JL, Chan J (2001) Immunology of tuberculosis. Annu Rev Immunol 19:93–129. doi:10.1146/annurev.immunol.19.1.93
Shepard CC (1957) Use of HeLa cells infected with tubercle bacilli for the study of antituberculous drugs. J Bacteriol 73:494–498
Kress G (1995) Combining dental training with medical training. J Dent Educ 59:1061
Shepard WP (1957) Industrial medicine, a new specialty. Can Med Assoc J 77:206–211
Rosqvist R, Magnusson KE, Wolf-Watz H (1994) Target cell contact triggers expression and polarized transfer of Yersinia YopE cytotoxin into mammalian cells. EMBO J 13:964–972
Bliska JB, Galan JE, Falkow S (1993) Signal transduction in the mammalian cell during bacterial attachment and entry. Cell 73:903–920. doi:0092-8674(93)90270-Z[pii]
Arruda S, Bomfim G, Knights R, Huima-Byron T, Riley LW (1993) Cloning of an M. tuberculosis DNA fragment associated with entry and survival inside cells. Science 261:1454–1457
Casali N, Riley LW (2007) A phylogenomic analysis of the Actinomycetales mce operons. BMC Genomics 8:60. doi:10.1186/1471-2164-8-60
Kumar A, Bose M, Brahmachari V (2003) Analysis of expression profile of mammalian cell entry (mce) operons of Mycobacterium tuberculosis. Infect Immun 71:6083–6087. doi:10.1128/IAI.71.10.6083-6087.2003
Santangelo MD, Klepp L, Nunez-Garcia J et al (2009) Mce3R, a TetR-type transcriptional repressor, controls the expression of a regulon involved in lipid metabolism in Mycobacterium tuberculosis. Microbiol-Sgm 155:2245–2255. doi:10.1099/Mic.0.027086-0
Dunphy KY, Senaratne RH, Masuzawa M, Kendall LV, Riley LW (2010) Attenuation of Mycobacterium tuberculosis functionally disrupted in a fatty acyl-coenzyme A synthetase gene fadD5. J Infect Dis 201:1232–1239. doi:10.1086/651452
Casali N, White AM, Riley LW (2006) Regulation of the Mycobacterium tuberculosis mce1 operon. J Bacteriol 188:441–449. doi:10.1128/JB.188.2.441-449.2006
Joon M, Bhatia S, Pasricha R, Bose M, Brahmachari V (2010) Functional analysis of an intergenic non-coding sequence within mce1 operon of M. tuberculosis. BMC Microbiol 10:128. doi:10.1186/1471-2180-10-128
Dassa E, Bouige P (2001) The ABC of ABCs: a phylogenetic and functional classification of ABC systems in living organisms. Res Microbiol 152(3–4):211–229
Mourez M, Hofnung M, Dassa E (1997) Subunit interactions in ABC transporters: a conservedsequence in hydrophobic membrane proteins of periplasmic permeases defines an important site of interaction with the ATPase subunits. EMBO J 16(11):3066–3077
Joshi SM, Pandey AK, Capite N, Fortune SM, Rubin EJ, Sassetti CM (2006) Characterization of mycobacterial virulence genes through genetic interaction mapping. Proc Natl Acad Sci USA 103:11760–11765. doi:10.1073/pnas.0603179103
Liu PQ, Liu CE, Ames GF (1999) Modulation of ATPase activity by physical disengagement of the ATP-binding domains of an ABC transporter, the histidine permease. J Biol Chem 274:18310–18318
Chitale S, Ehrt S, Kawamura I et al (2001) Recombinant Mycobacterium tuberculosis protein associated with mammalian cell entry. Cell Microbiol 3:247–254. doi:cmi110[pii]
El-Shazly S, Ahmad S, Mustafa AS, Al-Attiyah R, Krajci D (2007) Internalization by HeLa cells of latex beads coated with mammalian cell entry (Mce) proteins encoded by the mce3 operon of Mycobacterium tuberculosis. J Med Microbiol 56:1145–1151. doi:10.1099/jmm.0.47095-0
Harboe M, Christensen A, Ahmad S et al (2002) Cross-reaction between mammalian cell entry (Mce) proteins of mycobacterium tuberculosis. Scand J Immunol 56:580–587
Casali N, Konieczny M, Schmidt MA, Riley LW (2002) Invasion activity of a Mycobacterium tuberculosis peptide presented by the Escherichia coli AIDA autotransporter. Infect Immun 70:6846–6852. doi:10.1128/IAI.70.12.6846-6852.2002
Haile Y, Caugant DA, Bjune G, Wiker HG (2002) Mycobacterium tuberculosis mammalian cell entry operon (mce) homologs in Mycobacterium other than tuberculosis (MOTT). FEMS Immunol Med Microbiol 33:125–132. doi:PiiS0928-8244(02)00302-4
Cole ST, Eiglmeier K, Parkhill J et al (2001) Massive gene decay in the leprosy bacillus. Nature 409:1007–1011. doi:10.1038/35059006
Zumarraga M, Bigi F, Alito A, Romano MI, Cataldi A (1999) A 12.7 kb fragment of the Mycobacterium tuberculosis genome is not present in Mycobacterium bovis. Microbiology 145:893–897
Kumar A, Chandolia A, Chaudhry U, Brahmachari V, Bose M (2005) Comparison of mammalian cell entry operons of mycobacteria: in silico analysis and expression profiling. FEMS Immunol Med Microbiol 43:185–195. doi:10.1016/j.femsim.2004.08.013
Cole ST, Brosch R, Parkhill J et al (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393:537–544. doi:10.1038/31159
Dirusso CC, Black PN (2004) Bacterial long chain fatty acid transport: gateway to a fatty acid-responsive signaling system. J Biol Chem 279:49563–49566. doi:10.1074/jbc.R400026200
Cheigh CI, Senaratne R, Uchida Y, Casali N, Kendall LV, Riley LW (2010) Posttreatment reactivation of tuberculosis in mice caused by Mycobacterium tuberculosis disrupted in mce1R. J Infect Dis 202:752–759. doi:10.1086/655224
Ahmad S, Akbar PK, Wiker HG, Harboe M, Mustafa AS (1999) Cloning, expression and immunological reactivity of two mammalian cell entry proteins encoded by the mce1 operon of Mycobacterium tuberculosis. Scand J Immunol 50(5):510–518. doi:sji631[pii]
Tekaia F, Gordon SV, Garnier T, Brosch R, Barrell BG, Cole ST (1999) Analysis of the proteome of Mycobacterium tuberculosis in silico. Tuber Lung Dis 79:329–342. doi:S0962847999902204[pii]
Santangelo Mde L, Blanco F, Campos E et al (2009) Mce2R from Mycobacterium tuberculosis represses the expression of the mce2 operon. Tuberculosis 89:22–28. doi:10.1016/j.tube.2008.09.002
Marjanovic O, Miyata T, Goodridge A, Kendall LV, Riley LW (2010) Mce2 operon mutant strain of Mycobacterium tuberculosis is attenuated in C57BL/6 mice. Tuberculosis 90:50–56. doi:10.1016/j.tube.2009.10.004
Gioffre A, Infante E, Aguilar D et al (2005) Mutation in mce operons attenuates Mycobacterium tuberculosis virulence. Microbes Infect 7:325–334. doi:10.1016/j.micinf.2004.11.007
Ahmad S, El-Shazly S, Mustafa AS, Al-Attiyah R (2005) The six mammalian cell entry proteins (Mce3A-F) encoded by the mce3 operon are expressed during in vitro growth of Mycobacterium tuberculosis. Scand J Immunol 62:16–24. doi:10.1111/j.1365-3083.2005.01639.x
Ahmad S, El-Shazly S, Mustafa AS, Al-Attiyah R (2004) Mammalian cell-entry proteins encoded by the mce3 operon of Mycobacterium tuberculosis are expressed during natural infection in humans. Scand J Immunol 60:382–391. doi:10.1111/j.0300-9475.2004.01490.x
Santangelo MP, Goldstein J, Alito A et al (2002) Negative transcriptional regulation of the mce3 operon in Mycobacterium tuberculosis. Microbiology 148:2997–3006
Rkenes TP, Lamark T, Strom AR (1996) DNA-binding properties of the BetI repressor protein of Escherichia coli: the inducer choline stimulates BetI-DNA complex formation. J Bacteriol 178:1663–1670
Saini N, Sharma M, Chandolia A, Pasricha R, Brahmachari V, Bose M (2008) Characterization of Mce 4 A protein of Mycobacterium tuberculosis: role in invasion and survival. BMC Microbiol 8:200. doi:10.1186/1471-2180-8-200
Senaratne RH, Sidders B, Sequeira P et al (2008) Mycobacterium tuberculosis strains disrupted in mce3 and mce4 operons are attenuated in mice. J Med Microbiol 57:164–170. doi:10.1099/jmm.0.47454-0
Kendall SL, Withers M, Soffair CN et al (2007) A highly conserved transcriptional repressor controls a large regulon involved in lipid degradation in Mycobacterium smegmatis and Mycobacterium tuberculosis. Mol Microbiol 65:684–699. doi:10.1111/j.1365-2958.2007.05827.x
Pandey AK, Sassetti CM (2008) Mycobacterial persistence requires the utilization of host cholesterol. Proc Natl Acad Sci USA 105:4376–4380. doi:10.1073/pnas.0711159105
MacMicking JD, Taylor GA, McKinney JD (2003) Immune control of tuberculosis by IFN-gamma-inducible LRG-47. Science 302:654–659. doi:10.1126/science.1088063
Kendall SL, Burgess P, Balhana R et al (2010) Cholesterol utilization in mycobacteria is controlled by two TetR-type transcriptional regulators: kstR and kstR2. Microbiology 156:1362–1371. doi:10.1099/mic.0.034538-0
Xu G, Li Y, Yang J et al (2008) Mycobacterium bovis Mce4E protein may play a role in modulating cytokine expression profile in alveolar macrophage. Int J Tuberc Lung D 12:664–669
McDermott MF (2001) TNF and TNFR biology in health and disease. Cell Mol Biol (Noisy-le-grand) 47:619–635
Nathan CF, Hibbs JB Jr (1991) Role of nitric oxide synthesis in macrophage antimicrobial activity. Curr Opin Immunol 3(1):65–70
Moncada S, Palmer RM, Higgs EA (1991) Nitric oxide: physiology, pathophysiology, and pharmacology. Pharmacol Rev 43:109–142
Corbel C, Melchers F (1984) The synergism of accessory cells and of soluble alpha-factors derived from them in the activation of B cells to proliferation. Immunol Rev 78:51–74
Pasare C, Medzhitov R (2003) Toll pathway-dependent blockade of CD4 + CD25 + T cell-mediated suppression by dendritic cells. Science 299:1033–1036. doi:10.1126/science.10782311078231[pii]
Pajon R, Yero D, Lage A, Llanes A, Borroto CJ (2006) Computational identification of beta-barrel outer-membrane proteins in Mycobacterium tuberculosis predicted proteomes as putative vaccine candidates. Tuberculosis (Edinb) 86:290–302. doi:10.1016/j.tube.2006.01.005
Fu LM, Shinnick TM (2007) Genome-wide exploration of the drug action of capreomycin on Mycobacterium tuberculosis using Affymetrix oligonucleotide GeneChips. J Infection 54:277–284. doi:10.1016/j.jinf.2006.05.012
Acknowledgments
The work is Supported by the National key infectious disease project (No. 2008ZX10003-006, No. 2008ZX10003-001), national natural science foundation (No. 81071316), Excellent PhD thesis fellowship of southwest university (No. kb2009010, No. ky2009009), The Fundamental Research Funds for the Central Universities (XDJK2009A003) and Natural Science Foundation Project of CQ CSTC (CSTC, 2010BB5002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, F., Xie, JP. Mammalian cell entry gene family of Mycobacterium tuberculosis . Mol Cell Biochem 352, 1–10 (2011). https://doi.org/10.1007/s11010-011-0733-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-011-0733-5