Abstract
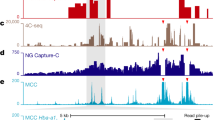
Hybridization with an oligonucleotide array was used to map the regions of DNA anchorage to the nuclear matrix. Matrix-associated DNA served as a hybridization probe. To obtain the oligonucleotide array, 60-mer oligonucleotides regularly distributed throughout the genome region of interest at 2-kb intervals were immobilized on a nylon filter. The organization of DNA into loop domains was studied in a 100-kb region of chicken chromosome 16, including the α -globin gene cluster. A 40-kb DNA loop, which was fixed to the nuclear matrix and harbored all α-globin genes, was observed in erythroid cells. One of its anchorage regions colocalized with matrix associated region (MAR) and an insulator found previously in the 5′ region of the chicken α-globin gene domain. The spatial (domain-loop) organization of the α-globin gene cluster in lymphoid cells proved to be strikingly different from that in erythroid cells.
Similar content being viewed by others
REFERENCES
Razin S.V., Petrov A., Hair A., Vassetzky Y.S. 2004. Chromatin domains and territories: Flexibly rigid. Crit. Rev. Eukaryot. Gene Expr. 14, 79–88.
Vassetzky Y., Hair A., Mechali M. 2000. Rearrangement of chromatin domains during development in Xenopus. Genes Dev. 14, 1541–1552.
Paulson J.R., Laemmli U.K. 1977. The structure of histone-depleted metaphase chromosomes. Cell. 12, 817–828.
Hancock R., Hughes M.E. 1982. Organization of DNA in the eukaryotic nucleus. Biol. Cell. 44, 201–212.
Buongiorno-Nardelli M., Gioacchino M., Carri M.T., Marilley M. 1982. A relationship between replicon size and supercoiled loop domains in the eukaryotic genome. Nature. 298, 100–102.
Razin S.V., Kekelidze M.G., Lukanidin E.M., Scherrer K., Georgiev G.P. 1986. Replication origins are attached to the nuclear skeleton. Nucleic Acids Res. 14, 8189–8207.
Vassetzky Y.S., Hair A., Razin S.V. 2000. Rearrangement of chromatin domains in cancer and development. J. Cell Biochem. S35, 54–60.
Svetlova E.Y., Razin S.V., Debatisse M. 2001. Mammalian recombination hot spot and DNA loop anchorage region: A model for the study of common fragile sites. J. Cell Biochem. S36, 170–178.
Iarovaia O.V., Bystritskiy A., Ravcheev D., Hancock R., Razin S.V. 2004. Visualization of individual DNA loops and a map of loop-domains in the human dystrophin gene. Nucleic Acids Res. 32, 2079–2086.
Razin S.V., Vassetzky Y.S., Hancock R. 1991. Nuclear matrix attachment regions and topoisomerase II binding and reaction sites in the vicinity of a chicken DNA replication origins. Biochem. Biophys. Res. Commun. 177, 265–270.
Boulikas T. 1993. Nature of DNA sequences at the attachment regions of genes to the nuclear matrix. J. Cell Biochem. 52, 14–22.
Vassetzky Y., Lemaitre J.M., Mechali M. 2000. Specification of chromatin domains and regulation of replication and transcription during development. Crit. Rev. Eukaryot. Gene Expr. 10, 31–38.
Linskens M.H., Eijsermans A., Dijkwel P.A. 1987. Comparative analysis of DNA loop length in nontransformed and transformed hamster cells. Mutat. Res. 178, 245–256.
Farache G., Razin S.V., Rzeszowska-Wolny J., Moreau J., Targa F.R., Scherrer K. 1990. Mapping of structural and transcription-related matrix attachment sites in the alpha-globin gene domain of avian erythroblasts and erythrocytes. Mol. Cell. Biol. 10, 5349–5358.
Basler J., Hastie N.D., Pietras D., Matsui S., Sandgerg A.A., Berezney R. 1981. Hybridization of nuclear matrix attached deoxyribonucleic acid fragments. Biochemistry. 20, 6921–6929.
Razin S.V., Mantieva V.L., Georgiev G.P. 1979. The similarity of DNA sequences remaining bound to scaffold upon nuclease treatment of interphase nuclei and metaphase chromosomes. Nucleic Acids Res. 7, 1713–1735.
Jackson D.A., Dickinson P., Cook P.R. 1990. The size of chromatin loops in HeLa cells. EMBO J. 9, 567–571.
Beug H., Doederlein G., Freudenstrein C., Graf T. 1979. Erythroblast cell lines transformed by a temperature sensitive mutant of avian erythroblastosis virus. A model system to study erythroid differentiation in vitro. J. Cell Physiol. 1, 195–207.
Rzeszowska-Wolny J., Razin S., Puvion E., Moreau J., Scherrer K. 1988. Isolation and characterization of stable nuclear matrix preparations and associated DNA from avian erythroblasts. Biol. Cell. 64, 13–22.
Razin S.V., Farrell C.M., Recillas-Targa F. 2003. Genomic domains and regulatory elements operating at the domain level. Int. Rev. Cytol. 226, 63–125.
Recillas-Targa F., Razin S.V. 2001. Chromatin domains and regulation of gene expression: Familiar and enigmatic clusters of chicken globin genes. Crit. Rev. Eukaryot. Gene Expr. 11, 227–242.
Razin S.V., Petrov P., Hancock R. 1991. Precise localization of the α-globin gene cluster within one of the 20-to 300-kilobase DNA fragment released by cleavage of chicken chromosomal DNA at topoisomerase II site in vivo: Evidence that the fragment are DNA loops or domains. Proc. Natl. Acad. Sci. USA. 88, 8515–8519.
Jackson D.A., McCready S.J., Cook P.R. 1981. RNA is synthesized at the nuclear cage. Nature. 292, 552–555.
Aelen J.M., Opstelsten R.J., Wanka F. 1983. Organization of DNA replication in Physarum polycephalum. Attachment of origins of replication and replication forks to the nuclear matrix. Nucleic Acids Res. 11, 1181–1195.
Bickmore W.A., Oghene K. 1996. Visualizing the spatial relationships between defined DNA sequences and the axial region of extracted metaphase chromosomes. Cell. 84, 95–104.
Iarovaia O.V., Shkumatov P., Razin S.V. 2004. Breakpoint cluster regions of the AML-1 and ETO genes contain MAR elements and are preferentially associated with the nuclear matrix in proliferating HEL cells. J. Cell Sci. 117, 4583–4590.
Cremer T., Cremer C. 2001. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nature Rev. Genet. 2, 292–301.
Ciejek E.M., Tsai M.-J., O'Malley B.W. 1983. Actively transcribed genes are associated with the nuclear matrix. Nature. 306, 607–609.
Cockerill P.N., Garrard W.T. 1986. Chromosomal loop anchorage of the kappa immunoglobulin gene occurs next to the enhancer in a region containing topo-isomerase II sites. Cell. 44, 273–282.
Cockerill P.N., Yuen M.-H., Garrard W.T. 1987. The enhancer of the immunoglobulin heavy chain locus is flanked by presumptive chromosomal loop anchorage elements. J. Biol. Chem. 262, 5394–5397.
Bode J., Schlake T., Rios-Ramirez M., Mielke C., Stengert M., Kay V., Klehr-Wirth D. 1995. Scaffold/matrix-attached regions: Structural properties creating transcriptionally active loci. Int. Rev. Cytol. 162A, 389–454.
Iarovaia O.V., Hancock R., Lagarkova M.A., Miassod R., Razin S.V. 1996. Mapping of genomic DNA loop organization in a 500-kilobase region of the Drosophila X chromosome using the topoisomerase II-mediated DNA loop excision protocol. Mol. Cell. Biol. 16, 302–308.
Razin S.V., Shen K., Ioudinkova E., Scherrer K. 1999. Functional analysis of DNA sequences located within a cluster of DNase I hypersensitive sites colocalising with MAR element at the upstream border of the chicken α-globin gene domain. J. Cell Biochem. 74, 38–49.
Valadez-Graham V., Razin S.V., Recillas-Targa F. 2004. CTCF-dependent enhancer blockers at the upstream region of the chicken alpha-globin gene domain. Nucleic Acids Res. 32, 1354–1362.
Razin S.V. 1987. DNA interaction with the nuclear matrix and spatial organization of replication and transcription. BioEssays. 6, 19–23.
Ratsch A., Joos S., Kioschis P., Lichter P. 2002. Topological organization of the MYC/IGK locus in Burkitt's lymphoma cells assessed by nuclear halo preparations. Exp. Cell Res. 273, 12–20.
Engel J.D., Dodgson J.B. 1980. Analysis of the closely linked adult chicken alpha-globin genes in recombinant DNAs. Proc. Natl. Acad. Sci. USA. 77, 2596–2600.
Farache G., Razin S.V., Recillas Targa F., Scherrer K. 1990. Organization of the 3′-boundary of the chicken α-globin gene domain and characterization of a CR1-specific protein binding site. Nucleic Acids Res. 18, 401–409.
Flint J., Tufarelli C., Peden J., Clark K., Daniels R.J., Hardison R., Miller W., Philipsen S., Tan-Un K.C., McMorrow T., Higgs D.R. 2001. Comparative genome analysis delimits a chromosomal domain and identifies key regulatory elements in the alpha globin cluster. Human Mol. Genet. 10, 371–382.
Ioudinkova E.S., Iarovaia O.V., Scherrer K., Razin S.V. 2001. Regulation of globin genes expression: New findings made with the chicken domain of alpha globin genes. Gene Ther. Mol. Biol. 6, 149–157.
Razin S.V., Rynditch A., Borunova V., Ioudinkova E., Smalko V., Scherrer K. 2004. The 33-kb transcript of the chicken alpha-globin gene domain is part of the nuclear matrix. J. Cell Biochem. 92, 445–457.
Sjakste N., Iarovaia O.V., Razin S.V., Linares-Cruz G., Sjakste T., Le Gac V., Zhao Z., Scherrer K. 2000. A novel gene is transcribed in the chicken alpha-globin gene domain in the direction opposite to the globin genes. Mol. Gen. Genet. 262, 1012–1021.
Author information
Authors and Affiliations
Additional information
__________
Translated from Molekulyarnaya Biologiya, Vol. 39, No. 6, 2005, pp. 971–977.
Original Russian Text Copyright © 2005 by Ioudinkova, Petrov, Vassetzky, Razin.
Rights and permissions
About this article
Cite this article
Ioudinkova, E.S., Petrov, A.V., Vassetzky, Y.S. et al. Spatial Organization of the Chicken α-Globin Gene Domain in Cells of Different Origins. Mol Biol 39, 851–856 (2005). https://doi.org/10.1007/s11008-005-0105-x
Received:
Issue Date:
DOI: https://doi.org/10.1007/s11008-005-0105-x