Abstract
We introduce propagation kernels, a general graph-kernel framework for efficiently measuring the similarity of structured data. Propagation kernels are based on monitoring how information spreads through a set of given graphs. They leverage early-stage distributions from propagation schemes such as random walks to capture structural information encoded in node labels, attributes, and edge information. This has two benefits. First, off-the-shelf propagation schemes can be used to naturally construct kernels for many graph types, including labeled, partially labeled, unlabeled, directed, and attributed graphs. Second, by leveraging existing efficient and informative propagation schemes, propagation kernels can be considerably faster than state-of-the-art approaches without sacrificing predictive performance. We will also show that if the graphs at hand have a regular structure, for instance when modeling image or video data, one can exploit this regularity to scale the kernel computation to large databases of graphs with thousands of nodes. We support our contributions by exhaustive experiments on a number of real-world graphs from a variety of application domains.
Similar content being viewed by others
1 Introduction
Learning from structured data is an active area of research. As domains of interest become increasingly diverse and complex, it is important to design flexible and powerful methods for analysis and learning. By structured data we refer to situations where objects of interest are structured and hence can naturally be represented using graphs. Real-world examples are molecules or proteins, images annotated with semantic information, text documents reflecting complex content dependencies, and manifold data modeling objects and scenes in robotics. The goal of learning with graphs is to exploit the rich information contained in graphs representing structured data. The main challenge is to efficiently exploit the graph structure for machine-learning tasks such as classification or retrieval. A popular approach to learning from structured data is to design graph kernels measuring the similarity between graphs. For classification or regression problems, the graph kernel can then be plugged into a kernel machine, such as a support vector machine or a Gaussian process, for efficient learning and prediction.
Several graph kernels have been proposed in the literature, but they often make strong assumptions as to the nature and availability of information related to the graphs at hand. The most simple of these proposals assume that graphs are unlabeled and have no structure beyond that encoded by their edges. However, graphs encountered in real-world applications often come with rich additional information attached to their nodes and edges. This naturally implies many challenges for representation and learning such as:
-
missing information leading to partially labeled graphs,
-
uncertain information arising from aggregating information from multiple sources, and
-
continuous information derived from complex and possibly noisy sensor measurements.
Images, for instance, often have metadata and semantic annotations which are likely to be only partially available due to the high cost of collecting training data. Point clouds captured by laser range sensors consist of continuous 3d coordinates and curvature information; in addition, part detectors can provide possibly noisy semantic annotations. Entities in text documents can be backed by entire Wikipedia articles providing huge amounts of structured information, themselves forming another network.
Surprisingly, existing work on graph kernels does not broadly account for these challenges. Most of the existing graph kernels (Shervashidze et al. 2009, 2011; Hido and Kashima 2009; Kashima et al. 2003; Gärtner et al. 2003) are designed for unlabeled graphs or graphs with a complete set of discrete node labels. Kernels for graphs with continuous node attributes have only recently gained greater interest (Borgwardt and Kriegel 2005; Kriege and Mutzel 2012; Feragen et al. 2013). These graph kernels have two major drawbacks: they can only handle graphs with complete label or attribute information in a principled manner and they are either efficient, but limited to specific graph types, or they are flexible, but their computation is memory and/or time consuming. To overcome these problems, we introduce propagation kernels. Their design is motivated by the observation that iterative information propagation schemes originally developed for within-network relational and semi-supervised learning have two desirable properties: they capture structural information and they can often adapt to the aforementioned issues of real-world data. In particular, propagation schemes such as diffusion or label propagation can be computed efficiently and they can be initialized with uncertain and partial information.
A high-level overview of the propagation kernel algorithm is as follows. We begin by initializing label and/or attribute distributions for every node in the graphs at hand. We then iteratively propagate this information along edges using an appropriate propagation scheme. By maintaining entire distributions of labels and attributes, we can accommodate uncertain information in a natural way. After each iteration, we compare the similarity of the induced node distributions between each pair of graphs. Structural similarities between graphs will tend to induce similar local distributions during the propagation process, and our kernel will be based on approximate counts of the number of induced similar distributions throughout the information propagation.
To achieve competitive running times and to avoid having to compare the distributions of all pairs of nodes between two given graphs, we will exploit locality sensitive hashing (lsh) to bin the label/attribute distributions into efficiently computable graph feature vectors in time linear in the total number of nodes. These new graph features will then be fed into a base kernel, a common scheme for constructing graph kernels. Whereas lsh is usually used to preserve the \(\ell ^1\) or \(\ell ^2\) distance, we are able to show that the hash values can preserve both the total variation and the Hellinger probability metrics. Exploiting explicit feature computation and efficient information propagation, propagation kernels allow for using graph kernels to tackle novel applications beyond the classical benchmark problems on datasets of chemical compounds and small- to medium-sized image or point-cloud graphs.
The present paper is a significant extension of a previously published conference paper (Neumann et al. 2012) and presents and extends a novel graph-kernel application already published as a workshop contribution (Neumann et al. 2013). Propagation kernels were originally defined and applied for graphs with discrete node labels (Neumann et al. 2012, 2013); here we extend their definition to a more general and flexible framework that is able to handle continuous node attributes. In addition to this expanded view of propagation kernels, we also introduce and discuss efficient propagation schemes for numerous classes of graphs. A central message of this paper is:
A suitable propagation scheme is the key to designing fast and powerful propagation kernels.
In particular, we will discuss propagation schemes applicable to huge graphs with regular structure, for example grid graphs representing images or videos. Thus, implemented with care, propagation kernels can easily scale to large image databases. The design of kernels for grids allows us to perform graph-based image analysis not only on the scene level (Neumann et al. 2012; Harchaoui and Bach 2007) but also on the pixel level opening up novel application domains for graph kernels.
We proceed as follows. We begin by touching upon related work on kernels and graphs. After introducing information propagation on graphs via random walks, we introduce the family of propagation kernels (Sect. 4). The following two sections discuss specific examples of the two main components of propagation kernels: node kernels for comparing propagated information (Sect. 5) and propagation schemes for various kinds of information (Sect. 6). We will then analyze the sensitivity of propagation kernels with respect to noisy and missing information, as well as with respect to the choice of their parameters. Finally, to demonstrate the feasibility and power of propagation kernels for large real-world graph databases, we provide experimental results on several challenging classification problems, including commonly used bioinformatics benchmark problems, as well as real-world applications such as image-based plant-disease classification and 3d object category prediction in the context of robotic grasping.
2 Kernels and graphs
Propagation kernels are related to three lines of research on kernels: kernels between graphs (graph kernels) developed within the graph mining community, kernels between nodes (kernels on graphs) established in the within-network relational learning and semi-supervised learning communities, and kernels between probability distributions.
2.1 Graph kernels
Propagation kernels are deeply connected to several graph kernels developed within the graph-mining community. Categorizing graph kernels with respect to how the graph structure is captured, we can distinguish four classes: kernels based on walks (Gärtner et al. 2003; Kashima et al. 2003; Vishwanathan et al. 2010; Harchaoui and Bach 2007) and paths (Borgwardt and Kriegel 2005; Feragen et al. 2013), kernels based on limited-size subgraphs (Horváth et al. 2004; Shervashidze et al. 2009; Kriege and Mutzel 2012), kernels based on subtree patterns (Mahé and Vert 2009; Ramon and Gärtner 2003), and kernels based on structure propagation (Shervashidze et al. 2011). However, there are two major problems with most existing graph kernels: they are often slow or overly specialized. There are efficient graph kernels specifically designed for unlabeled and fully labeled graphs (Shervashidze et al. 2009, 2011), attributed graphs (Feragen et al. 2013), or planar labeled graphs (Harchaoui and Bach 2007), but these are constrained by design. There are also more flexible but slower graph kernels such as the shortest path kernel (Borgwardt and Kriegel 2005) or the common subgraph matching kernel (Kriege and Mutzel 2012).
The Weisfeiler–Lehman (wl) subtree kernel, one instance of the recently introduced family of wl-kernels (Shervashidze et al. 2011), computes count features for each graph based on signatures arising from iterative multi-set label determination and compression steps. In each kernel iteration, these features are then fed into a base kernel. The wl-kernel is finally the sum of those base kernels over the iterations.
Although wl-kernels are usually competitive in terms of performance and runtime, they are designed for fully labeled graphs. The challenge of comparing large, partially labeled graphs—which can easily be considered by propagation kernels introduced in the present paper—remains to a large extent unsolved. A straightforward way to compute graph kernels between partially labeled graphs is to mark unlabeled nodes with a unique symbol or their degree as suggested in Shervashidze et al. (2011) for the case of unlabeled graphs. However, both solutions neglect any notion of uncertainty in the labels. Another option is to propagate labels across the graph and then run a graph kernel on the imputed labels (Neumann et al. 2012). Unfortunately, this also ignores the uncertainty induced by the inference procedure, as hard labels have to be assigned after convergence. A key observation motivating propagation kernels is that intermediate label distributions induced will, before convergence, carry information about the structure of the graph. Propagation kernels interleave label inference and kernel computation steps, avoiding the requirement of running inference to termination prior to the kernel computation.
2.2 Kernels on graphs and within-network relational learning
Measuring the structural similarity of local node neighborhoods has recently become popular for inference in networked data (Kondor and Lafferty 2002; Desrosiers and Karypis 2009; Neumann et al. 2013) where this idea has been used for designing kernels on graphs (kernels between the nodes of a graph) and for within-network relational learning approaches. An example of the former are coinciding walk kernels (Neumann et al. 2013) which are defined in terms of the probability that the labels encountered during parallel random walks starting from the respective nodes of a graph coincide. Desrosiers and Karypis (2009) use a similarity measure based on parallel random walks with constant termination probability in a relaxation-labeling algorithm. Another approach exploiting random walks and the structure of subnetworks for node-label prediction is heterogeneous label propagation (Hwang and Kuang 2010). Random walks with restart are used as proximity weights for so-called “ghost edges” in Gallagher et al. (2008), but then the features considered by a bag of logistic regression classifiers are only based on a one-step neighborhood. These approaches, as well as propagation kernels, use random walks to measure structure similarity. Therefore, propagation kernels establish an important connection of graph-based machine learning for inference about node- and graph-level properties.
2.3 Kernels between probability distributions and kernels between sets
Finally, propagation kernels mark another connection, namely between graph kernels and kernels between probability distributions (Jaakkola and Haussler 1998; Lafferty and Lebanon 2002; Moreno et al. 2003; Jebara et al. 2004) and between sets (Kondor and Jebara 2003; Shi et al. 2009). However, whereas the former essentially build kernels based on the outcome of probabilistic inference after convergence, propagation kernels intuitively count common sub-distributions induced after each iteration of running inference in two graphs.
Kernels between sets and more specifically between structured sets, also called hash kernels (Shi et al. 2009), have been successfully applied to strings, data streams, and unlabeled graphs. While propagation kernels hash probability distributions and derive count features from them, hash kernels directly approximate the kernel values \(k(x,x')\), where x and \(x'\) are (structured) sets. Propagation kernels iteratively approximate node kernels k(u, v) comparing nodes u in graph \(G^{(i)}\) with nodes v in graph \(G^{(j)}\). Counts summarizing these approximations are then fed into a base kernel that is computed exactly. Before we give a detailed definition of propagation kernels, we introduce the basic concept of information propagation on graphs, and exemplify important propagation schemes and concepts when utilizing random walks for learning with graphs.
3 Information propagation on graphs
Information propagation or diffusion on a graph is most commonly modeled via Markov random walks (rws). Propagation kernels measure the similarity of two graphs by comparing node label or attribute distributions after each step of an appropriate random walk. In the following, we review label diffusion and label propagation via rws—two techniques commonly used for learning on the node level (Zhu et al. 2003; Szummer and Jaakkola 2001). Based on these ideas, we will then develop propagation kernels in the subsequent sections.
3.1 Basic notation
Throughout, we consider graphs whose nodes are endowed with (possibly partially observed) label and/or attribute information. That is, a graph \(G=(V,E,\ell , a)\) is represented by a set of \(|V|=n\) vertices, a set of edges E specified by a weighted adjacency matrix \(A \in \mathbb {R}^{n \times n}\), a label function \(\ell :V \rightarrow [k]\), where k is the number of available node labels, and an attribute function with \(a:V \rightarrow \mathbb {R}^D\), where D is the dimension of the continuous attributes. Given \(V = \{v_1,v_2,...,v_n\}\), node labels \(\ell (v_i)\) are represented by nominal values and attributes \(a(v_i)=\mathbf {x}_i \in \mathbb {R}^D\) are represented by continuous vectors.
3.2 Markov random walks
Consider a graph \(G = (V,E)\). A random walk on G is a Markov process \(X = \{X_t : t\ge 0 \}\) with a given initial state \(X_0 = v_i\). We will also write \(X_{t\mid i}\) to indicate that the walk began at \(v_i\). The transition probability \(T_{ij} = P(X_{t+1}=v_j \mid X_{t}=v_i)\) only depends on the current state \(X_t=v_i\) and the one-step transition probabilities for all nodes in V can be easily represented by the row-normalized adjacency or transition matrix \(T = D^{-1}A\), where \(D = {{\mathrm{diag}}}(\sum _j A_{ij})\).
3.3 Information propagation via random walks
For now, we consider (partially) labeled graphs without attributes, where \(V = V_L \cup V_U\) is the union of labeled and unlabeled nodes and \(\ell :V \rightarrow [k]\) is a label function with known values for the nodes in \(V_L\). A common mechanism for providing labels for the nodes of an unlabeled graph is to define the label function by \(\ell (v_i) = \sum _j A_{ij} = \text {degree}(v_i)\). Hence for fully labeled and unlabeled graphs we have \(V_U = \emptyset \). We will monitor the distribution of labels encountered by random walks leaving each node in the graph to endow each node with a k-dimensional feature vector. Let the matrix \(P_0 \in \mathbb {R} ^{n \times k}\) give the prior label distributions of all nodes in V, where the ith row \((P_0)_{i} = p_{0,v_i}\) corresponds to node \(v_i\). If node \(v_i \in V_L\) is observed with label \(\ell (v_i)\), then \(p_{0,v_i}\) can be conveniently set to a Kronecker delta distribution concentrating at \(\ell (v_i)\); i.e., \(p_{0,v_i} = \delta _{\ell (v_i)}\). Thus, on graphs with \(V_U = \emptyset \) the simplest rw-based information propagation is the label diffusion process or simply diffusion process
where \(p_{t,v_i}\) gives the distribution over \(\ell (X_{t \mid i})\) at iteration t.
Let \(S \subseteq V\) be a set of nodes in G. Given T and S, we define an absorbing random walk to have the modified transition probabilities \(\hat{T}\), defined by:
Nodes in S are “absorbing” in that a walk never leaves a node in S after it is encountered. The ith row of \(P_0\) now gives the probability distribution for the first label encountered, \(\ell (X_{0\mid i})\), for an absorbing rw starting at \(v_i\). It is easy to see by induction that by iterating the map
\(p_{t,v_i}\) similarly gives the distribution over \(\ell (X_{t \mid i})\).
In the case of partially labeled graphs we can now initialize the label distributions for the unlabeled nodes \(V_U\) with some prior, for example a uniform distribution.Footnote 1 If we define the absorbing states to be the labeled nodes, \(S = V_L\), then the label propagation algorithm introduced in Zhu et al. (2003) can be cast in terms of simulating label-absorbing rws with transition probabilities given in Eq. (2) until convergence, then assigning the most probable absorbing label to the nodes in \(V_U\).
The schemes discussed so far are two extreme cases of absorbing rws: one with no absorbing states, the diffusion process, and one which absorbs at all labeled nodes, label propagation. One useful extension of absorbing rws is to soften the definition of absorbing states. This can be naturally achieved by employing partially absorbing random walks (Wu et al. 2012). As the propagation kernel framework does not require a specific propagation scheme, we are free to choose any rw-based information propagation scheme suitable for the given graph types. Based on the basic techniques introduced in this section, we will suggest specific propagation schemes for (un-)labeled, partially labeled, directed, and attributed graphs as well as for graphs with regular structure in Sect. 6.
3.4 Steady-state distributions versus early stopping
Assuming non-bipartite graphs, all rws, absorbing or not, converge to a steady-state distribution \(P_\infty \) (Lovász 1996; Wu et al. 2012). Most existing rw-based approaches only analyze the walks’ steady-state distributions to make node-label predictions (Kondor and Lafferty 2002; Zhu et al. 2003; Wu et al. 2012). However, rws without absorbing states converge to a constant steady-state distribution, which is clearly uninteresting. To address this, the idea of early stopping was successfully introduced into power-iteration methods for node clustering (Lin and Cohen 2010), node-label prediction (Szummer and Jaakkola 2001), as well as for the construction of a kernel for node-label prediction (Neumann et al. 2013). The insight here is that the intermediate distributions obtained by the rws during the convergence process provide useful insights about their structure. In this paper, we adopt this idea for the construction of graph kernels. That is, we use the entire evolution of distributions encountered during rws up to a given length to represent graph structure. This is accomplished by summing contributions computed from the intermediate distributions of each iteration, rather then only using the limiting distribution.
4 Propagation kernel framework
In this section, we introduce the general family of propagation kernels (pks).
4.1 General definition
Here we will define a kernel \(K:\mathcal {X} \times \mathcal {X} \rightarrow \mathbb {R}\) among graph instances \(G^{(i)} \in \mathcal {X} \). The input space \(\mathcal {X}\) comprises graphs \(G^{(i)} = (V^{(i)},E^{(i)},\ell ,a)\), where \(V^{(i)}\) is the set of \(|V^{(i)}| = n_i\) nodes and \(E^{(i)}\) is the set of edges in graph \(G^{(i)}\). Edge weights are represented by weighted adjacency matrices \(A^{(i)} \in \mathbb {R}^{n_i \times n_i}\) and the label and attribute functions \(\ell \) and a endow nodes with label and attribute informationFootnote 2 as defined in the previous section.
A simple way to compare two graphs \(G^{(i)}\) and \(G^{(j)}\) is to compare all pairs of nodes in the two graphs:
where k(u, v) is an arbitrary node kernel determined by node labels and, if present, node attributes. This simple graph kernel, however, does not account for graph structure given by the arrangement of node labels and attributes in the graphs. Hence, we consider a sequence of graphs \(G^{(i)}_t\) with evolving node information based on information propagation, as introduced for node labels in the previous section. We define the kernel contribution of iteration t by
An important feature of propagation kernels is that the node kernel k(u, v) is defined in terms of the nodes’ corresponding probability distributions \(p_{t,u}\) and \(p_{t,v}\), which we update and maintain throughout the process of information propagation. For propagation kernels between labeled and attributed graphs we define
where \(k_\ell (u,v)\) is a kernel corresponding to label information and \(k_a(u,v)\) is a kernel corresponding to attribute information. If no attributes are present, then \(k(u,v) = k_\ell (u,v)\). \(k_\ell \) and \(k_a\) will be defined in more detail later. For now assume they are given, then the \(t_{\textsc {max}}\)-iteration propagation kernel is given by
Lemma 1
Given that \(k_\ell (u,v)\) and \(k_a(u,v)\) are positive semidefinite node kernels, the propagation kernel \(K_{t_{\textsc {max}}}\) is a positive semidefinite kernel.
Proof
As \(k_\ell (u,v)\) and \(k_a(u,v)\) are assumed to be valid node kernels, k(u, v) is a valid node kernel as the product of positive semidefinite kernels is again positive semidefinite. As for a given graph \(G^{(i)}\) the number of nodes is finite, \(K(G^{(i)}_t, G^{(j)}_t)\) is a convolution kernel (Haussler 1999). As sums of positive semidefinite matrices are again positive semidefinite, the propagation kernel as defined in Eq. (6) is positive semidefinite. \(\square \)

Let \(|V^{(i)}| = n_i\) and \(|V^{(j)}| = n_j\). Assuming that all node information is given, the complexity of computing each contribution between two graphs, Eq. (4), is \(\mathcal {O}(n_i \, n_j)\). Even for medium-sized graphs this can be prohibitively expensive considering that the computation has to be performed for every pair of graphs in a possibly large graph database. However, if we have a node kernel of the form
where \( condition \) is an equality condition on the information of nodes u and v, we can compute K efficiently by binning the node information, counting the respective bin strengths for all graphs, and computing a base kernel among these counts. That is, we compute count features \(\phi (G^{(i)}_t)\) for each graph and plug them into a base kernel \(\langle \cdot ,\cdot \rangle \):
In the simple case of a linear base kernel, the last step is just an outer product of count vectors \(\varPhi _t \varPhi _t^\top \), where the ith row of \(\varPhi _t, \left( \varPhi _t \right) _{i\, \cdot } = \phi (G^{(i)}_t)\). Now, for two graphs, binning and counting can be done in \(\mathcal {O}(n_i+n_j)\) and the computation of the linear base kernel value is \(\mathcal {O}(|\text {bins}|)\). This is one of the main insights for efficient graph-kernel computation and it has already been exploited for labeled graphs in previous work (Shervashidze et al. 2011; Neumann et al. 2012).
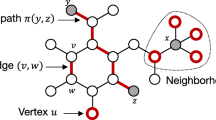
Figure 1 illustrates the propagation kernel computation for \(t=0\) and \(t=1\) for two example graphs and Algorithm 1 summarizes the kernel computation for a graph database \(\{G^{(i)}\}_i\). From this general algorithm and Eqs. (4) and (6), we see that the two main components to design a propagation kernel are
-
the node kernel k(u, v) comparing propagated information, and
-
the propagation scheme \(P^{(i)}_{t+1} \leftarrow P^{(i)}_{t}\) propagating the information within the graphs.
The propagation scheme depends on the input graphs and we will give specific suggestions for different graph types in Sect. 6. Before defining the node kernels depending on the available node information in Sect. 5, we briefly discuss the general runtime complexity of propagation kernels.
Propagation kernel computation. Distributions, bins, count features, and kernel contributions for two graphs \(G^{(i)}\) and \(G^{(j)}\) with binary node labels and one iteration of label propagation, cf. Eq. (3), as the propagation scheme. Node-label distributions are decoded by color: white means \(p_{0,u} = [1,0]\), dark red stands for \(p_{0,u} = [0,1]\), and the initial distributions for unlabeled nodes (light red) are \(p_{0,u} = [{1}/{2},{1}/{2}]\). a Initial label distributions \((t=0)\), b updated label distributions (\(t=1\)) (Color figure online)
4.2 Complexity analysis
The total runtime complexity of propagation kernels for a set of n graphs with a total number of N nodes and M edges is \(\mathcal {O}\bigl ((t_{\textsc {max}}-1) M + t_{\textsc {max}} \,n^2\,n^{\star }\bigr )\), where \(n^{\star } := \max _i(n_i)\). For a pair of graphs the runtime complexity of computing the count features, that is, binning the node information and counting the bin strengths is \(\mathcal {O}(n_i + n_j)\). Computing and adding the kernel contribution is \(\mathcal {O}(|\text {bins}|)\), where \(|\text {bins}|\) is bounded by \(n_i + n_j\). So, one iteration of the kernel computation for all graphs is \(\mathcal {O}(n^2\,n^{\star })\). Note that in practice \(|\text {bins}| \ll 2 n^{\star }\) as we aim to bin together similar nodes to derive a meaningful feature representation.
Feature computation basically depends on propagating node information along the edges of all graphs. This operation depends on the number of edges and the information propagated, so it is \(\mathcal {O}((k+D)M) = \mathcal {O}(M)\), where k is the number of node labels and D is the attribute dimensionality. This operation has to be performed \(t_{\textsc {max}}-1\) times. Note that the number of edges is usually much lower than \(N^2\).
5 Propagation kernel component 1: node kernel
In this section, we define node kernels comparing propagated information appropriate for the use in propagation kernels. Moreover, we introduce locality sensitive hashing, which is used to discretize the distributions arsing from rw-based information propagation as well as the continuous attributes directly.
5.1 Definitions
Above, we saw that one way to allow for efficient computation of propagation kernels is to restrict the range of the node kernels to \(\lbrace 0, 1 \rbrace \). Let us now define the two components of the node kernel [Eq. (5)] in this form. The label kernel can be represented as
where \(p_{t,u}\) is the node-label distribution of node u at iteration t and \(h_\ell (\cdot )\) is a quantization function (Gersho and Gray 1991), more precisely a locality sensitive hash (lsh) function (Datar and Indyk 2004), which will be introduced in more detail in the next section. Note that \(p_{t,u}\) denotes a row in the label distribution matrix \(P^{(i)}_t\), namely the row corresponding to node u of graph \(G^{(i)}\).
Propagation kernels can be computed for various kinds of attribute kernels as long as they have the form of Eq. (7). The most rudimentary attribute kernel is
where \(x_u\) is the one-dimensional continuous attribute of node u and \(h_{a}(\cdot )\) is again an lsh function. Figure 2 contrasts this simple attribute kernel for a one-dimensional attribute to a thresholded Gaussian function and the Gaussian kernel commonly used to compare node attributes in graph kernels. To deal with higher-dimensional attributes, we can choose the attribute kernel to be the product of kernels on each attribute dimension:
where \(x_{u,d}\) is the respective dimension of the attribute \(\mathbf {x}_u\) of node u. Note that each dimension now has its own lsh function \(h_{a_d}(\cdot )\). However, analogous to the label kernel, we can also define an attribute kernel based on propagated attribute distributions
where \(q_{t,u}\) is the attribute distribution of node u at iteration t. Next we explain the locality sensitive hashing approach used to discretize distributions and continuous attributes directly. In Sect. 6.3, we will then derive an efficient way to propagate and hash continuous attribute distributions.
5.2 Locality sensitive hashing
We now describe our quantization approach for implementing propagation kernels for graphs with node-label distributions and continuous attributes. The idea is inspired by locality sensitive hashing (Datar and Indyk 2004) which seeks quantization functions on metric spaces where points “close enough” to each other in that space are “probably” assigned to the same bin. In the case of distributions, we will consider each node-label vector as being an element of the space of discrete probability distributions on k items equipped with an appropriate probability metric. If we want to hash attributes directly, we simply consider metrics for continuous values.
Definition 1
(Locality Sensitive Hash (lsh)) Let \(\mathcal {X}\) be a metric space with metric \(d:\mathcal {X} \times \mathcal {X} \rightarrow {\mathbb {R}}\), and let \(\mathcal {Y} = \lbrace 1, 2, \cdots , k' \rbrace \). Let \(\theta > 0\) be a threshold, \(c > 1\) be an approximation factor, and \(p_1, p_2 \in (0, 1)\) be the given success probabilities. A set of functions \(\mathcal {H}\) from \(\mathcal {X}\) to \(\mathcal {Y}\) is called a \((\theta , c\theta , p_1, p_2)\)-locality sensitive hash if for any function \(h \in \mathcal {H}\) chosen uniformly at random, and for any two points \(x, x' \in \mathcal {X}\), it holds that
-
if \(d(x, x') < \theta \), then \(\Pr (h(x) = h(x')) > p_1\), and
-
if \(d(x, x') > c\theta \), then \(\Pr (h(x) = h(x')) < p_2\).
It is known that we can construct lsh families for \(\ell ^p\) spaces with \(p \in (0, 2]\) (Datar and Indyk 2004). Let V be a real-valued random variable. V is called p-stable if for any \(\lbrace x_1, x_2,\, \cdots , x_d \rbrace , x_i \in {\mathbb {R}}\) and independently sampled \(v_1, v_2, \cdots , v_d\), we have \(\sum x_i v_i \sim ||\mathbf {x} ||_p V\).
Explicit p-stable distributions are known for some p; for example, the standard Cauchy distribution is 1-stable, and the standard normal distribution is 2-stable. Given the ability to sample from a p-stable distribution V, we may define an lsh \(\mathcal {H}\) on \({\mathbb {R}}^d\) with the \(\ell ^p\) metric (Datar and Indyk 2004). An element h of \(\mathcal {H}\) is specified by three parameters: a width \(w \in {\mathbb {R}}^+\), a d-dimensional vector \(\mathbf {v}\) whose entries are independent samples of V, and b drawn from \(\mathcal {U}[0, w]\). Given these, h is then defined as
We may now consider \(h(\cdot )\) to be a function mapping our distributions or attribute values to integer-valued bins, where similar distributions end up in the same bin. Hence, we obtain node kernels as defined in Eqs. (9) and (12) in the case of distributions, as well as simple attribute kernels as defined in Eqs. (10) and (11). To decrease the probability of collision, it is common to choose more than one random vector \(\mathbf {v}\). For propagation kernels, however, we only use one hyperplane, as we effectively have \(t_{\textsc {max}}\) hyperplanes for the whole kernel computation and the probability of a hash conflict is reduced over the iterations.

The intuition behind the expression in Eq. (13) is that p-stability implies that two vectors that are close under the \(\ell ^p\) norm will be close after taking the dot product with \(\mathbf {v}\); specifically, \((\mathbf {v}^\top \mathbf {x} - \mathbf {v}^\top \mathbf {x'})\) is distributed as \(||\mathbf {x} - \mathbf {x}' ||_p V\). So, in the case where we want to construct a hashing for D-dimensional continuous node attributes to preserve \(\ell ^1\) (l1) or \(\ell ^2\) (l2) distance
we directly apply Eq. (13). In the case of distributions, we are concerned with the space of discrete probability distributions on k elements, endowed with a probability metric d. Here we specifically consider the total variation (tv) and Hellinger (h) distances:
The total variation distance is simply half the \(\ell ^1\) metric, and the Hellinger distance is a scaled version of the \(\ell ^2\) metric after applying the map \(p \mapsto \sqrt{p}\). We may therefore create a locality-sensitive hash family for \(d_{{\textsc {tv}}}\) by direct application of Eq. (13) and create a locality-sensitive hash family for \(d_{{\textsc {h}}}\) by using Eq. (13) after applying the square root map to our label distributions. The lsh computation for a matrix \(X \in \mathbb {R}^{N \times D}\), where \(\mathbf {x}_u\) is the row in X corresponding to node u, is summarized in Algorithm 2.
6 Propagation kernel component 2: propagation scheme
As pointed out in the introduction, the input graphs for graph kernels may vary considerably. One key to design efficient and powerful propagation kernels is the choice of a propagation scheme appropriate for the graph dataset at hand. By utilizing random walks (rws) we are able to use efficient off-the-shelf algorithms, such as label diffusion or label propagation (Szummer and Jaakkola 2001; Zhu et al. 2003; Wu et al. 2012), to implement information propagation within the input graphs. In this section, we explicitly define propagation kernels for fully labeled, unlabeled, partially labeled, directed, and attributed graphs as well as for graphs with a regular grid structure using appropriate rws. In each particular algorithm, the specific parts changing compared to the general propagation kernel computation (Algorithm 1) will be marked in color.
6.1 Labeled and unlabeled graphs
For fully labeled graphs we suggest the use of the label diffusion process from Eq. (1) as the propagation scheme. Given a database of fully labeled graphs \(\{G^{(i)}\}_{i=1,\dots ,n}\) with a total number of \(N=\sum _i n_i\) nodes, label diffusion on all graphs can be efficiently implemented by multiplying a sparse block-diagonal transition matrix \(T \in \mathbb {R}^{N\times N}\), where the blocks are the transition matrices \(T^{(i)}\) of the respective graphs, with the label distribution matrix \(P_t = \left[ P^{(1)}_t,\dots ,P^{(n)}_t \right] ^\top \in \mathbb {R}^{N\times k}\). This can be done efficiently due to the sparsity of T. The propagation kernel computation for labeled graphs is summarized in Algorithm 3. The specific parts compared to the general propagation kernel computation (Algorithm 1) for fully labeled graphs are marked in green (input) and blue (computation). For unlabeled graphs we suggest to set the label function to be the node degree \(\ell (u) = \text {degree}(u)\) and then apply the same pk computation as for fully labeled graphs.

6.2 Partially labeled and directed graphs
For partially labeled graphs, where some of the node labels are unknown, we suggest label propagation as an appropriate propagation scheme. Label propagation differs from label diffusion in the fact that before each iteration of the information propagation, the labels of the originally labeled nodes are pushed back (Zhu et al. 2003). Let \(P_0 = \left[ P_{0,[\text {labeled}]}, P_{0,[\text {unlabeled}]}\right] ^\top \) represent the prior label distributions for the nodes of all graphs in the graph database, where the distributions in \(P_{0,[\text {labeled}]}\) represent observed labels and \(P_{0,[\text {unlabeled}]}\) are initialized uniformly. Then label propagation is defined by
Note that this propagation scheme is equivalent to the one defined in Eq. (3) using fully absorbing rws. Other similar update schemes, such as “label spreading” (Zhou et al. 2003), could be used in a propagation kernel as well. Thus, the propagation kernel computation for partially labeled graphs is essentially the same as Algorithm 3, where the initialization for the unlabeled nodes has to be adapted, and the (partial) label push back has to be added before the node information is propagated. The relevant parts are the ones marked in blue. Note that for graphs with large fractions of labeled nodes it might be preferable to use label diffusion even though they are partially labeled.
To implement propagation kernels between directed graphs, we can proceed as above after simply deriving transition matrices computed from the potentially non-symmetric adjacency matrices. That is, for the propagation kernel computation only the input changes (marked in green in Algorithm 3). The same idea allows weighted edges to be accommodated; again, only the transition matrix has to be adapted. Obviously, we can also combine partially labeled graphs with directed or weighted edges by changing both the blue and green marked parts accordingly.
6.3 Graphs with continuous node attributes
Nowadays, learning tasks often involve graphs whose nodes are attributed with continuous information. Chemical compounds can be annotated with the length of the secondary structure elements (the nodes) or measurements for various properties, such as hydrophobicity or polarity. 3d point clouds can be enriched with curvature information, and images are inherently composed of 3-channel color information. All this information can be modeled by continuous node attributes. In Eq. (10) we introduced a simple way to deal with attributes. The resulting propagation kernel essentially counts similar label arrangements only if the corresponding node attributes are similar as well. Note that for higher-dimensional attributes it can be advantageous to compute separate lshs per dimension, leading to the node kernel introduced in Eq. (11). This has the advantage that if we standardize the attributes, we can use the same bin-width parameter \(w_a\) for all dimensions. In all our experiments we normalize each attribute to have unit standard deviation and will set \(w_a = 1\). The disadvantage of this method, however, is that the arrangement of attributes in the graphs is ignored.

In the following, we derive p2k, a variant of propagation kernels for attributed graphs based on the idea of propagating both attributes and labels. That is, we model graph similarity by comparing the arrangement of labels and the arrangement of attributes in the graph. The attribute kernel for p2k is defined as in Eq. (12); now the question is how to efficiently propagate the continuous attributes and how to efficiently model and hash the distributions of (multivariate) continuous variables. Let \(X \in \mathbb {R}^{N \times D}\) be the design matrix, where a row \(\mathbf {x}_u\) represents the attribute vector of node u. We will associate with each node of each graph a probability distribution defined on the attribute space, \(q_u\), and will update these as attribute information is propagated across graph edges as before. One challenge in doing so is ensuring that these distributions can be represented with a finite description. The discrete label distributions from before have naturally a finite number of dimensions and could be compactly represented and updated via the \(P_t\) matrices. We seek a similar representation for attributes. Our proposal is to define the node-attribute distributions to be mixtures of D-dimensional multivariate Gaussians, one centered on each attribute vector in X:
where the sum ranges over all nodes \(v, W_{u, \cdot }\) is a vector of mixture weights, and \(\varSigma \) is a shared \(D \times D\) covariance matrix for each component of the mixture. In particular, here we set \(\varSigma \) to be the sample covariance matrix calculated from the N vectors in X. Now the \(N \times N\) row-normalized W matrix can be used to compactly represent the entire set of attribute distributions. As before, we will use the graph structure to iteratively spread attribute information, updating these W matrices, deriving a sequence of attribute distributions for each node to use as inputs to node attribute kernels in a propagation kernel scheme.
We begin by defining the initial weight matrix \(W_0\) to be the identity matrix; this is equivalent to beginning with each node attribute distribution being a single Gaussian centered on the corresponding attribute vector:
Now, in each propagation step the attribute distributions are updated by the distribution of their neighboring nodes \(Q_{t+1} \leftarrow Q_t\). We accomplish this by propagating the mixture weights W across the edges of the graph according to a row-normalized transition matrix T, derived as in Sect. 3:
We have described how attribute distributions are associated with each node and how they are updated via propagating their weights across the edges of the graph. However, the weight vectors contained in W are not themselves directly suitable for comparing in an attribute kernel \(k_a\), because any information about the similarity of the mean vectors is ignored. For example, imagine that two nodes u and v had exactly the same attribute vector, \(\mathbf {x}_u = \mathbf {x}_v\). Then mass on the u component of the Gaussian mixture is interchangeable with mass on the v component; a simple kernel among weight vectors cannot capture this. For this reason, we use a vector more appropriate for kernel comparison. Namely, we select a fixed set of sample points in attribute space (in our case, chosen uniformly from the node attribute vectors in X), evaluate the pdfs of the Gaussian mixtures associated with each node at these points, and use this vector to summarize the node information. This handles the exchangeability issue from above and also allows a more compact representation for hash inputs; in our experiments, we used 100 sample points and achieved good performance. As before, these vectors can then be hashed jointly or individually for each sample point. Note that the bin width \(w_a\) has to be adapted accordingly. In our experiments, we will use the latter option and set \(w_a = 1\) for all datasets.
The computational details of p2k are given in Algorithm 4, where the additional parts compared to Algorithm 1 are marked in blue (computation) and green (input). An extension to Algorithm 4 would be to refit the gms after a couple of propagation iterations. We did not consider refitting in our experiments as the number of kernel iterations \(t_{\textsc {max}}\) was set to 10 or 15 for all datasets—following the descriptions in existing work on iterative graph kernels (Shervashidze et al. 2011; Neumann et al. 2012).
6.4 Grid graphs
One of our goals in this paper is to compute propagation kernels for pixel grid graphs. A graph kernel between grid graphs can be defined such that two grids should have a high kernel value if they have similarly arranged node information. This can be naturally captured by propagation kernels as they monitor information spread on the grids. Naïvely, one could think that we can simply apply Algorithm 3 to achieve this goal. However, given that the space complexity of this algorithm scales with the number of edges and even medium sized images such as texture patches will easily contain thousands of nodes, this is not feasible. For example considering \(100 \times 100\)-pixel image patches with an 8-neighborhood graph structure, the space complexity required would be 2.4 million unitsFootnote 3 (floating point numbers) per graph. Fortunately, we can exploit the flexibility of propagation kernels by exchanging the propagation scheme. Rather than label diffusion as used earlier, we employ discrete convolution; this idea was introduced for efficient clustering on discrete lattices (Bauckhage and Kersting 2013). In fact, isotropic diffusion for denoising or sharpening is a highly developed technique in image processing (Jähne 2005). In each iteration, the diffused image is derived as the convolution of the previous image and an isotropic (linear and space-invariant) filter. In the following, we derive a space- and time-efficient way of computing propagation kernels for grid graphs by means of convolutions.
6.4.1 Basic Definitions
Given that the neighborhood of a node is the subgraph induced by all its adjacent vertices, we define a d-dimensional grid graph as a lattice graph whose node embedding in \(\mathbb {R}^d\) forms a regular square tiling and the neighborhoods \(\mathcal {N}\) of each non-border node are isomorphic (ignoring the node information). Figure 3 illustrates a regular square tiling and several isomorphic neighborhoods of a 2-dimensional grid graph. If we ignore boundary nodes, a grid graph is a regular graph; i.e., each non-border node has the same degree. Note that the size of the border depends on the radius of the neighborhood. In order to be able to neglect the special treatment for border nodes, it is common to view the actual grid graph as a finite section of an actually infinite graph.
A grid graph whose node embedding in \(\mathbb {R}^d\) forms a regular square tiling can be derived from the graph Cartesian product of line graphs. So, a two-dimensional grid is defined as
where \(L_{m_{i,1}}\) is a line graph with \(m_{i,1}\) nodes. \(G^{(i)}\) consists of \(n_i = m_{i,1} \, m_{i,2}\) nodes, where non-border nodes have the same number of neighbors. Note that the grid graph \(G^{(i)}\) only specifies the node layout in the graph but not the edge structure. The edges are given by the neighborhood \(\mathcal {N}\) which can be defined by any arbitrary matrix B encoding the weighted adjacency of its center node. The nodes, being for instance image pixels, can carry discrete or continuous vector-valued information. Thus, in the most-general setting the database of grid graphs is given by \(\mathbf {G} = \{G^{(i)}\}_{i=1,\ldots ,n}\) with \(G^{(i)} = (V^{(i)}, \mathcal {N}, \ell )\), where \(\ell : V^{(i)} \rightarrow \mathcal {L}\) with \(\mathcal {L} = ([k], \mathbb {R}^D)\). Commonly used neighborhoods \(\mathcal {N}\) are the 4-neighborhood and the 8-neighborhood illustrated in Fig. 3b, c.
6.4.2 Discrete Convolution
The general convolution operation on two functions f and g is defined as
That is, the convolution operation produces a modified, or filtered, version of the original function f. The function g is called a filter. For two-dimensional grid graphs interpreted as discrete functions of two variables x and y, e.g., the pixel location, we consider the discrete spatial convolution defined by:
where the computation is in fact done for finite intervals. As convolution is a well-studied operation in low-level signal processing and discrete convolution is a standard operation in digital image processing, we can resort to highly developed algorithms for its computation; see for example Chapter 2 in Jähne (2005). Convolutions can be computed efficiently via the fast Fourier transformation in \(\mathcal {O}(n_i\,\log \,n_i)\) per graph.

6.4.3 Efficient Propagation Kernel Computation
Now let \(\mathbf {G} = \{G^{(i)}\}_i\) be a database of grid graphs. To simplify notation, however without loss of generality, we assume two-dimensional grids \(G^{(i)} = L_{m_{i,1}} \times L_{m_{i,2}}\). Unlike in the case of general graphs, each graph now has a natural two-dimensional structure, so we will update our notation to reflect this structure. Instead of representing the label probability distributions of each node as rows in a two-dimensional matrix, we now represent them in the third dimension of a three-dimensional tensor \(P^{(i)}_t \in \mathbb {R}^{m_{i,1}\times m_{i,2} \times k}\). Modifying the structure makes both the exposition more clear and also enables efficient computation. Now, we can simply consider discrete convolution on k matrices of label probabilities \(P^{(i,j)}\) per grid graph \(G^{(i)}\), where \(P^{(i,j)} \in \mathbb {R}^{m_{i,1}\times m_{i,2}}\) contains the probabilities of all nodes in \(G^{(i)}\) of being label j and \(j \in \{1, \dots , k\}\). For observed labels, \(P^{(i)}_0\) is again initialized with a Kronecker delta distribution across the third dimension and, in each propagation step, we perform a discrete convolution of each matrix \(P^{(i,j)}\) per graph. Thus, we can create various propagation schemes efficiently by applying appropriate filters, which are represented by matrices B in our discrete case. We use circular symmetric neighbor sets \(N_{r,p}\) as introduced in Ojala et al. (2002), where each pixel has p neighbors which are equally spaced pixels on a circle of radius r. We use the following approximated filter matrices in our experiments:
The propagation kernel computation for grid graphs is summarized in Algorithm 5, where the specific parts compared to the general propagation kernel computation (Algorithm 1) are highlighted in green (input) and blue (computation). Using fast Fourier transformation, the time complexity of Algorithm 5 is \(\mathcal {O}\left( (t_{\textsc {max}}-1) N \log N + t_{\textsc {max}}\,n^2\,n^{\star }\right) \). Note that for the purpose of efficient computation, calculate-lsh has to be adapted to take the label distributions \(\{P^{(i)}_t\}_i\) as a set of 3-dimensional tensors. By virtue of the invariance of the convolutions used, propagation kernels for grid graphs are translation invariant, and when using the circular symmetric neighbor sets they are also 90-degree rotation invariant. These properties make them attractive for image-based texture classification. The use of other filters implementing for instance anisotropic diffusion depending on the local node information is a straightforward extension.
7 Experimental evaluation
Our intent here is to investigate the power of propagation kernels (pks) for graph classification. Specifically, we ask:
- (Q1) :
-
How sensitive are propagation kernels with respect to their parameters, and how should propagation kernels be used for graph classification?
- (Q2) :
-
How sensitive are propagation kernels to missing and noisy information?
- (Q3) :
-
Are propagation kernels more flexible than state-of-the-art graph kernels?
- (Q4) :
-
Can propagation kernels be computed faster than state-of-the-art graph kernels while achieving comparable classification performance?
Towards answering these questions, we consider several evaluation scenarios on diverse graph datasets including chemical compounds, semantic image scenes, pixel texture images, and 3d point clouds to illustrate the flexibility of pks.
7.1 Datasets
The datasets used for evaluating propagation kernels come from a variety of different domains and thus have diverse properties. We distinguish graph databases of labeled and attributed graphs, where attributed graphs usually also have label information on the nodes. Also, we separate image datasets where we use the pixel grid graphs from general graphs, which have varying node degrees. Table 1 summarizes the properties of all datasets used in our experiments.Footnote 4
7.1.1 Labeled Graphs
For labeled graphs, we consider the following benchmark datasets from bioinformatics: mutag, nci1, nci109, and d&d. mutag contains 188 sets of mutagenic aromatic and heteroaromatic nitro compounds, and the label refers to their mutagenic effect on the Gram-negative bacterium Salmonella typhimurium (Debnath et al. 1991). nci1 and nci109 are anti-cancer screens, in particular for cell lung cancer and ovarian cancer cell lines, respectively (Wale and Karypis 2006). d&d consists of 1178 protein structures (Dobson and Doig 2003), where the nodes in each graph represent amino acids and two nodes are connected by an edge if they are less than 6 Ångstroms apart. The graph classes are enzymes and non-enzymes.
7.1.2 Partially Labeled Graphs
The two real-world image datasets msrc 9-class and msrc 21-classFootnote 5 are state-of-the-art datasets in semantic image processing originally introduced in Winn et al. (2005). Each image is represented by a conditional Markov random field graph, as illustrated in Fig. 4a, b. The nodes of each graph are derived by oversegmenting the images using the quick shift algorithm,Footnote 6 resulting in one graph among the superpixels of each image. Nodes are connected if the superpixels are adjacent, and each node can further be annotated with a semantic label. Imagining an image retrieval system, where users provide images with semantic information, it is realistic to assume that this information is only available for parts of the images, as it is easier for a human annotator to label a small number of image regions rather than the full image. As the images in the msrc datasets are fully annotated, we can derive semantic (ground-truth) node labels by taking the mode ground-truth label of all pixels in the corresponding superpixel. Semantic labels are, for example, building, grass, tree, cow, sky, sheep, boat, face, car, bicycle, and a label void to handle objects that do not fall into one of these classes. We removed images consisting of solely one semantic label, leading to a classification task among eight classes for msrc9 and 20 classes for msrc21.
Semantic scene and point cloud graphs. The rgb image in (a) is represented by a graph of superpixels (b) with semantic labels b \(=\) building, c \(=\) car, v \(=\) void, and ? \(=\) unlabeled. c Point clouds of household objects represented by labeled 4-nn graphs with part labels top (yellow), middle (blue), bottom (red), usable-area (cyan), and handle (green). Edge colors are derived from the adjacent nodes. a rgb image, b superpixel graph, c point cloud graphs (Color figure online)
7.1.3 Attributed Graphs
To evaluate the ability of pks to incorporate continuous node attributes, we consider the attributed graphs used in Feragen et al. (2013), Kriege and Mutzel (2012). Apart from one synthetic dataset (synthetic), the graphs are all chemical compounds (enzymes, proteins, pro-full, bzr, cox2, and dhfr). synthetic comprises 300 graphs with 100 nodes, each endowed with a one-dimensional normally distributed attribute and 196 edges each. Each graph class, A and B, has 150 examples, where in A, 10 node attributes were flipped randomly and in B, 5 were flipped randomly. Further, noise drawn from \(\mathcal {N}(0,0.45^2)\) was added to the attributes in B. proteins is a dataset of chemical compounds with two classes (enzyme and non-enzyme) introduced in Dobson and Doig (2003). enzymes is a dataset of protein tertiary structures belonging to 600 enzymes from the brenda database (Schomburg et al. 2004). The graph classes are their ec (enzyme commission) numbers which are based on the chemical reactions they catalyze. In both datasets, nodes are secondary structure elements (sse), which are connected whenever they are neighbors either in the amino acid sequence or in 3d space. Node attributes contain physical and chemical measurements including length of the sse in Ångstrom, its hydrophobicity, its van der Waals volume, its polarity, and its polarizability. For bzr, cox2, and dhfr—originally used in Mahé and Vert (2009)—we use the 3d coordinates of the structures as attributes.
7.1.4 Point Cloud Graphs
In addition, we consider the object database db,Footnote 7 introduced in Neumann et al. (2013). db is a collection of 41 simulated 3d point clouds of household objects. Each object is represented by a labeled graph where nodes represent points, labels are semantic parts (top, middle, bottom, handle, and usable-area), and the graph structure is given by a k-nearest neighbor (k-nn) graph w.r.t. Euclidean distance of the points in 3d space, cf. Fig. 4c. We further endowed each node with a continuous curvature attribute approximated by its derivative, that is, by the tangent plane orientations of its incident nodes. The attribute of node u is given by \(x_{u} = \sum _{v \in \mathcal {N}(u)} 1-|\mathbf {n}_u \cdot \mathbf {n}_v|\), where \(\mathbf {n}_u\) is the normal of point u and \(\mathcal {N}(u)\) are the neighbors of node u. The classification task here is to predict the category of each object. Examples of the 11 categories are glass, cup, pot, pan, bottle, knife, hammer, and screwdriver.
7.1.5 Grid Graphs
We consider a classical benchmark dataset for texture classification (brodatz) and a dataset for plant disease classification (plants). All graphs in these datasets are grid graphs derived from pixel images. That is, the nodes are image pixels connected according to circular symmetric neighbor sets \(N_{r,p}\) as exemplified in Eq. (16). Node labels are computed from the rgb color values by quantization.
brodatz,Footnote 8 introduced in Valkealahti and Oja (1998), covers 32 textures from the Brodatz album with 64 images per class comprising the following subsets of images: 16 “original” images (o), 16 rotated versions (r), 16 scaled versions (s), and 16 rotated and scaled versions (rs) of the “original” images. Figure 5a, b show example images with their corresponding quantized versions (e) and (f). For parameter learning, we used a random subset of 20 % of the original images and their rotated versions, and for evaluation we use test suites similar to the ones provided with the dataset.Footnote 9 All train/test splits are created such that whenever an original image (o) occurs in one split, their modified versions (r,s,rs) are also included in the same split.
The images in plants, introduced in Neumann et al. (2014), are regions showing disease symptoms extracted from a database of 495 rgb images of beet leaves. The dataset has six classes: five disease symptoms cercospora, ramularia, pseudomonas, rust, and phoma, and one class for extracted regions not showing a disease symptom. Figure 5c, d illustrates two regions and their quantized versions (g) and (h). We follow the experimental protocol in Neumann et al. (2014) and use 10 % of the full data covering a balanced number of classes (296 regions) for parameter learning and the full dataset for evaluation. Note that this dataset is highly imbalanced, with two infrequent classes accounting for only 2 % of the examples and two frequent classes covering 35 % of the examples.
7.2 Experimental protocol
We implemented propagation kernels in MatlabFootnote 10 and classification performance on all datasets except for db is evaluated by running c-svm classifications using libSVM.Footnote 11 For the parameter analysis (Sect. 7.3), the cost parameter c was learned on the full dataset (\(c \in \{10^{-3}, 10^{-1}, 10^{1}, 10^{3}\}\) for normalized kernels and \(c \in \{10^{-3}, 10^{-2}, 10^{-1},10^{0}\}\) for unnormalized kernels), for the sensitivity analysis (Sect. 7.4), it was set to its default value of 1 for all datasets, and for the experimental comparison with existing graph kernels (Sect. 7.5), we learned it via 5-fold cross-validation on the training set for all methods (\(c \in \{10^{-7}, 10^{-5}, \dots , 10^{5},10^{7} \}\) for normalized kernels and \(c \in \{10^{-7}, 10^{-5}, 10^{-3},10^{-1} \}\) for unnormalized kernels). The number of kernel iterations \(t_{\textsc {max}}\) was learned on the training splits (\(t_{\textsc {max}} \in \{0,1,\dots , 10\}\) unless stated otherwise). Reported accuracies are an average of 10 reruns of a stratified 10-fold cross-validation.
For db, we follow the protocol introduced in Neumann et al. (2013). We perform a leave-one-out (loo) cross validation on the 41 objects in db, where the kernel parameter \(t_{\textsc {max}}\) is learned on each training set again via loo. We further enhanced the nodes by a standardized continuous curvature attribute, which was only encoded in the edge weights in previous work (Neumann et al. 2013).
For all pks, the lsh bin-width parameters were set to \(w_l = 10^{-5}\) for labels and to \(w_a = 1\) for the normalized attributes, and as lsh metrics we chose \(\textsc {m}_l = \textsc {tv}\) and \(\textsc {m}_a = \textsc {l1}\) in all experiments. Before we evaluate classification performance and runtimes of the proposed propagation kernels, we analyze their sensitivity towards the choice of kernel parameters and with respect to missing and noisy observations.
7.3 Parameter analysis
To analyze parameter sensitivity with respect to the kernel parameters w (lsh bin width) and \(t_{\textsc {max}}\) (number of kernel iterations), we computed average accuracies over 10 randomly generated test sets for all combinations of w and \(t_{\textsc {max}}\), where \(w \in \{10^{-8},10^{-7}, \dots , 10^{-1} \}\) and \(t_{\textsc {max}} \in \{0,1,\dots ,14\}\) on mutag, enzymes, nci1, and db. The propagation kernel computation is as described in Algorithm 3, that is, we used the label information on the nodes and the label diffusion process as propagation scheme. To assess classification performance, we performed a 10-fold cross validation (cv). Further, we repeated each of these experiments with the normalized kernel, where normalization means dividing each kernel value by the square root of the product of the respective diagonal entries. Note that for normalized kernels we test for larger svm cost values. Figure 6 shows heatmaps of the results.
Parameter sensitivity of pk. The plots show heatmaps of average accuracies (tenfold cv) of pk (labels only) w.r.t. the bin widths parameter w and the number of kernel iterations \(t_{\textsc {max}}\) for four datasets mutag, enzymes, nci1, and db. In panels (a, c, e, g) we used the kernel matrix directly, in panels (b, d, f, h) we normalized the kernel matrix. The svm cost parameter is learned for each combination of w and \(t_{\textsc {max}}\) on the full dataset. \(\varvec{\times }\) marks the highest accuracy. a mutag; b mutag, normalized; c enzymes; d enzymes, normalized; e nci1; f nci1, normalized; g db and h db, normalized
In general, we see that the pk performance is relatively smooth, especially if \(w < 10^{-3}\) and \(t_{\textsc {max}} > 4\). Specifically, the number of iterations leading to the best results are in the range from \(\{4,\dots ,10\}\) meaning that we do not have to use a larger number of iterations in the pk computations, helping to keep a low computation time. This is especially important for parameter learning. Comparing the heatmaps of the normalized pk to the unnormalized pk leads to the conclusion that normalizing the kernel matrix can actually hurt performance. This seems to be the case for the molecular datasets mutag and nci1. For mutag, Fig. 6a, b, the performance drops from 88.2 to 82.9 %, indicating that for this dataset the size of the graphs, or more specifically the amount of labels from the different kind of node classes, are a strong class indicator for the graph label. Nevertheless, incorporating the graph structure, i.e., comparing \(t_{\textsc {max}}=0\) to \(t_{\textsc {max}}=10\), can still improve classification performance by 1.5 %. For other prediction scenarios such as the object category prediction on the db dataset, Fig. 6g, h, we actually want to normalize the kernel matrix to make the prediction independent of the object scale. That is, a cup scanned from a larger distance being represented by a smaller graph is still a cup and should be similar to a larger cup scanned from a closer view. So, for our experiments on object category prediction we will use normalized graph kernels whereas for the chemical compounds we will use unnormalized kernels unless stated otherwise.
Recall that our propagation kernel schemes are randomized algorithms, as there is randomization inherent in the choice of hyperplanes used during the lsh computation. We ran a simple experiment to test the sensitivity of the resulting graph kernels with respect to the hyperplane used. We computed the pk between all graphs in the datasets mutag, enzymes, msrc9, and msrc21 with \(t_{\textsc {max}} = 10\) 100 times, differing only in the random selection of the lsh hyperplanes. To make comparisons easier, we normalized each of these kernel matrices. We then measured the standard deviation of each kernel entry across these repetitions to gain insight into the stability of the pk to changes in the lsh hyperplanes. The median standard deviations were: mutag: \(5.5 \times 10^{-5}\), enzymes: \(1.1 \times 10^{-3}\), msrc9: \(2.2 \times 10^{-4}\), and msrc21: \(1.1 \times 10^{-4}\). The maximum standard deviations over all pairs of graphs were: mutag: \(6.7 \times 10^{-3}\), enzymes: \(1.4 \times 10^{-2}\), msrc9: \(1.4 \times 10^{-2}\), and msrc21: \(1.1 \times 10^{-2}\). Clearly the pk values are not overly sensitive to random variation due to differing random lsh hyperplanes.
In summary, we can answer (Q1) by concluding that pks are not overly sensitive to the random selection of the hyperplane as well as to the choice of parameters and we propose to learn \(t_{\textsc {max}} \in \{0,1,\dots ,10\}\) and fix \(w \le 10^{-3}\). Further, we recommend to decide on using the normalized version of pks only when graph size invariance is deemed important for the classification task.
7.4 Sensitivity to missing and noisy information
This section analyzes the performance of propagation kernels in the presence of missing and noisy information.
To asses how sensitive propagation kernels are to missing information, we randomly selected x % of the nodes in all graphs of db and removed their labels (labels) or attributes (attr), where \(x \in \{0,10,\ldots ,90, 95,98,99,99.5,100\}\). To study the performance when both label and attribute information is missing, we selected (independently) x % of the nodes to remove their label information and \(x\%\) of the nodes to remove their attribute information (labels & attr). Figure 7 shows the average accuracy of 10 reruns. While we see that the accuracy decreases with more missing information, the performance remains stable in the case when attribute information is missing. This suggests that the label information is more important for the problem of object category prediction. Further, the standard error is increasing with more missing information, which corresponds to the intuition that fewer available information results in a higher variance in the predictions.
Missing information versus accuracy on db. We plot missing information (%) versus avg. accuracy (%) \(\pm \) standard error of p2k on db. For labels only label information is missing, for attr only attribute information is missing and for labels & attr both is missing. The reported accuracies are averaged over ten reruns. random indicates the result of a random predictor
We also compare the predictive performance of propagation kernels when only some graphs, as for instance graphs at prediction time, have missing labels. Therefore, we divided the graphs of the following datasets, mutag, enzymes, msrc9, and msrc21, into two groups. For one group (fully labeled) we consider all nodes to be labeled, and for the other group (missing labels) we remove x % of the labels at random, where \(x \in \{10,20,\dots ,90,91,\dots ,99\}\). Figure 8 shows average accuracies over 10 reruns for each dataset. Whereas for mutag we do not observe a significant difference of the two groups, for enzymes the graphs with missing labels could only be predicted with lower accuracy, even when only 20 % of the labels were missing. For both msrc datasets, we observe that we can still predict the graphs with full label information quite accurately; however, the classification accuracy for the graphs with missing information decreases significantly with the amount of missing labels. For all datasets removing even 99 % of the labels still leads to better classification results than a random predictor. This result may indicate that the size of the graphs itself bears some predictive information. This observation confirms the results from Sect. 7.3.
Missing node labels. We plot missing labels (%) versus avg. accuracy (%) \(\pm \) standard error for mutag, enzymes, msrc9, and msrc21. We divided the graphs randomly in two equally sized groups (fully labeled and missing labels). The reported accuracies are averaged over 10 reruns. random indicates the result of a predictor randomly choosing the class label. a mutag, b enzymes, c msrc9, d msrc21
The next experiment analyzes the performance of propagation kernels when label information is encoded as attributes in a one-hot encoding. We also examine how sensitive they are in the presence of label noise. We corrupted the label encoding by an increasing amount of noise. A noisy label distribution vector \(\mathbf {n}_u\) was generated by sampling \(n_{u,i} \sim \mathcal {U}(0, 1)\) and normalizing so that \(\sum n_{u,i} = 1\). Given a noise level \(\alpha \), we used the following values encoded as attributes
Figure 9 shows average accuracies over 10 reruns for msrc9, msrc21, mutag, and enzymes. First, we see that using the attribute encoding of the label information in a p2k variant only propagating attributes achieves similar performances to propagating the labels directly in pk. This confirms that the Gaussian mixture approximation of the attribute distributions is a reasonable choice. Moreover, we can observe that the performance on msrc9 and mutag is stable across the tested noise levels. For msrc21 the performance drops for noise levels larger than 0.3. Whereas the same happens for enzymes, adding a small amount of noise (\(\alpha =0.1\)) actually increases performance. This could be due to a regularization effect caused by the noise and should be investigated in future work.
Noisy node labels. Noise level \(\alpha \) is plotted versus avg. accuracy \(\pm \) standard error of 10 reruns on msrc9, msrc21, mutag, and enzymes when encoding the labels as k-dimensional attributes using a one-hot representation and attribute propagation as in Algorithm 4. The dashed lines indicate the performance when using the usual label encoding without noise and Algorithm 3
Finally, we performed an experiment to test the sensitivity of pks with respect to noise in edge weights. For this experiment, we used the datasets bzr, cox2, and dhfr, and defined edge weights between connected nodes according to the distance between the corresponding structure elements in 3d space. Namely, the edge weight (before row normalization) was taken to be the inverse Euclidean distance between the incident nodes. Given a noise-level \(\sigma \), we corrupted each edge weight by multiplying by random log-normally distributed noise:
where \(\varepsilon \sim \mathcal {N}(0, \sigma ^2)\). Figure 10 shows the average test accuracy across ten repetitions of 10-fold cross-validation for this experiment. The bzr and cox2 datasets tolerated a large amount of edge-weight noise without a large effect on predictive performance, whereas dhfr was somewhat more sensitive to larger noise levels.
Summing up these experimental results we answer (Q2) by concluding that propagation kernels behave well in the presence of missing and noisy information.
7.5 Comparison to existing graph kernels
We compare classification accuracy and runtime of propagation kernels (pk) with the following state-of-the-art graph kernels: the Weisfeiler–Lehman subtree kernel (wl) (Shervashidze et al. 2011), the shortest path kernel (sp) (Borgwardt and Kriegel 2005), the graph hopper kernel (gh) (Feragen et al. 2013), and the common subgraph matching kernel (csm) (Kriege and Mutzel 2012). Table 2 lists all graph kernels and the types of information they are intended for. For all wl computations, we used the fast implementationFootnote 12 introduced in (Kersting et al. 2014). In sp, gh, and csm, we used a Dirac kernel to compare node labels and a Gaussian kernel \(k_a(u,v) = \exp (-\gamma \Vert x_u - x_v\Vert ^2)\) with \(\gamma = {1}/{D}\) for attribute information, if feasible. csm for the bigger datasets (enzymes, proteins, synthetic) was computed using a Gaussian truncated for inputs with \(\Vert x_u - x_v\Vert > 1\). We made this decision to encourage sparsity in the generated (node) kernel matrices, reducing the size of the induced product graphs and speeding up computation. Note that this is technically not a valid kernel between nodes; nonetheless, the resulting graph kernels were always positive definite. For pk and wl the number of kernel iterations (\(t_{\textsc {max}}\) or \(h_{\textsc {max}}\)) and for csm the maximum size of subgraphs (k) was learned on the training splits via 10-fold cross validation. For all runtime experiments all kernels were computed for the largest value of \(t_{\textsc {max}}, h_{\textsc {max}}\), or k, respectively. We used a linear base kernel for all kernels involving count features, and attributes, if present, were standardized. Further, we considered two baselines that do not take the graph structure into account. labels, corresponding to a pk with \(t_{\textsc {max}}=0\), only compares the label proportions in the graphs and a takes the mean of a Gaussian node kernel among all pairs of nodes in the respective graphs.
7.5.1 Graph classification on benchmark data
In this section, we consider graph classification for fully labeled, partially labeled, and attributed graphs.
Fully labeled graphs The experimental results for labeled graphs are shown in Table 3. On mutag, the baseline using label information only (labels) already gives the best performance indicating that for this dataset the actual graph structure is not adding any predictive information. On nci1 and nci109, wl performs best; however, propagation kernels come in second while being computed over one minute faster. Although sp can be computed quickly, it performs significantly worse than pk and wl. This is also the case for gh, whose computation time is significantly higher. In general, the results on labeled graphs show that propagation kernels can be computed faster than state-of-the-art graph kernels but achieve comparable classification performance, thus question (Q4) can be answered affirmatively.
Partially labeled graphs To assess the predictive performance of propagation kernels on partially labeled graphs, we ran the following experiments 10 times. We randomly removed 20–80 % of the node labels in all graphs in msrc9 and msrc21 and computed cross-validation accuracies and standard errors. Because the wl-subtree kernel was not designed for partially labeled graphs, we compare pk to two variants: one where we treat unlabeled nodes as an additional label “u” (wl) and another where we use hard labels derived from running label propagation (lp) until convergence (lp \(+\) wl). For this experiment we did not learn the number of kernel iterations, but selected the best performing \(t_{\textsc {max}}\) resp. \(h_{\textsc {max}}\).
The results are shown in Table 4. For larger fractions of missing labels, pk obviously outperforms the baseline methods, and, surprisingly, running label propagation until convergence and then computing wl gives slightly worse results than wl. However, label propagation might be beneficial for larger amounts of missing labels. The runtimes of the different methods on msrc21 are shown in Fig. 12 in the “Appendix 1”. wl computed via the string-based implementation suggested in (Shervashidze et al. 2011) is over 36 times slower than pk. These results again confirm that propagation kernels have attractive scalability properties for large datasets. The lp \(+\) wl approach wastes computation time while running lp to convergence before it can even begin calculating the kernel. The intermediate label distributions obtained during the convergence process are already extremely powerful for classification. These results clearly show that propagation kernels can successfully deal with partially labeled graphs and suggest an affirmative answer to questions (Q3) and (Q4).
Attributed graphs The experimental results for various datasets with attributed graphs are illustrated in Fig. 11. The plots show runtime versus average accuracy, where the error bars reflect standard deviation of the accuracies. As we are interested in good predictive performance while achieving fast kernel computation, methods in the upper-left corners provide the best performance with respect to both quality and speed. For pk, sp, gh, and csm we compare three variants: one where we use the labels only, one where we use the attribute information only, and one where both labels and attributes are used. wl is computed with label information only. For synthetic, cf. Fig. 11a, we used the node degree as label information. Further, we compare the performance of p2k, which propagates labels and attributes as described in Sect. 6.3. Detailed results on synthetic and all bioinformatics datasets are provided in Table 8 (average accuracies) and Table 7 (runtimes) in the “Appendix 2”. From Fig. 11 we clearly see that propagation kernels tend to appear in the upper-left corner, that is, they are achieving good predictive performance while being fast, leading to a positive answer of question (Q4). Note that the runtimes are shown on a log scale. We can also see that p2k, propagating both labels and attributes, (blue star) usually outperforms the the simple pk implementation not considering attribute arrangements (blue diamond). However, this comes at the cost of being slower. So, we can use the flexibility of propagation kernels to trade predictive quality against speed or vice versa according to the requirements of the application at hand. This supports a positive answer to question (Q3).
Attributed graphs: log runtime versus accuracy. The plots show log(runtimes) in seconds plotted against average accuracy (\(\pm \) standard deviation). Methods are encoded by color and the used information (labels, attributes or both) is encoded by shape. Note that p2k also uses both, labels and attributes, however in contrast to pk both are propagated. For pro-full the csm kernel computation exceeded 32 GB of memory. On synthetic csm using the attribute information only could not be computed within 72 h. a synthetic, b enzymes, c bzr, d pro-full (Color figure online)
7.5.2 Graph classification on novel applications
The flexibility of propagation kernels arising from easily interchangeable propagation schemes and their efficient computation via lsh allows us to apply graph kernels to novel domains. First, we are able to compare larger graphs with reasonable time expended, opening up the use of graph kernels for object category prediction of 3d point clouds in the context of robotic grasping (Neumann et al. 2013). Depending on their size and the perception distance, point clouds of household objects can easily consist of several thousands of nodes. Traditional graph kernels suffer from enormous computation times or memory problems even on datasets like db, which can still be regarded medium sized. These issues aggravate even more when considering image data. So far, graph kernels have been used for image classification on the scene level where the nodes comprise segments of similar pixels and one image is then represented by less than 100 so-called superpixels. Utilizing off-the-shelf techniques for efficient diffusion on grid graphs allows the use of propagation kernels to analyze images on the pixel level and thus opens up a whole area of interesting problems in the intersection of graph-based machine learning and computer vision. As a first step, we apply graph kernels, more precisely propagation kernels, to texture classification, where we consider datasets with thousands of graphs containing a total of several millions of nodes.
3 d object category prediction In this set of experiments, we follow the protocol introduced in Neumann et al. (2013), where the graph kernel values are used to derive a prior distribution on the object category for a given query object. The experimental results for the 3d-object classification are summarized in Table 5. We observe that propagation kernels easily deal with the point-cloud graphs. From the set of baseline graph kernels considered, only wl was feasible to compute, however with poor performance. Propagation kernels clearly benefit form their flexibility as we can improve the classification accuracy from 75.4 to 80.7 % when considering the object curvature attribute. These results are extremely promising given that we tackle a classification problem with 11 classes having only 40 training examples for each query object.
Grid graphs For brodatz and plants we follow the experimental protocol in Neumann et al. (2014). The pk parameter \(t_{\textsc {max}}\) was learned on a training subset of the full dataset (\(t_{\textsc {max}} \in \{0,3,5,8,10,15,20\}\)). For plants this training dataset consists of 10 % of the full data; for brodatz we used 20 % of the brodatz-o-r data as the number of classes in this dataset is much larger (32 textures). We also learned the quantization values (\( col \in \{3,5,8,10,15\}\)) and neighborhoods (\(B \in \{N_{1,4}, N_{1,8}, N_{2,16}\}\), cf. Eq. (16)). For brodatz the best performance on the training data was achieved with 3 colors and a 8-neighborhood, whereas for plants 5 colors and the 4-neighborhood was learned. We compare pk to the simple baseline labels using label counts only and to a powerful second-order statistical feature based on the gray-level co-occurrence matrix (Haralick et al. 1973) comparing intensities (glcm-gray) resp. quantized labels (glcm-quant) of neighboring pixels. The experimental results for grid graphs are shown in Table 6. While not outperforming sophisticated and complex state-of-the-art computer vision approaches to texture classification, we find that it is feasible to compute pks on huge image datasets achieving respectable performance out of the box. This is—compared to the immense tuning of features and methods commonly done in computer vision—a great success. On plants pk achieves an average accuracy of 82.5 %, where the best reported result so far is 83.7 %, which was only achieved after tailoring a complex feature ensemble (Neumann et al. 2014). In conclusion, propagation kernels are an extremely promising approach in the intersection of machine learning, graph mining, and computer vision.
Summarizing all experimental results, the capabilities claimed in Table 2 are supported. Propagation kernels have proven extremely flexible and efficient and thus question (Q3) can ultimately be answered affirmatively.
8 Conclusion
Random walk-based models provide a principled way of spreading information and even handling missing and uncertain information within graphs. Known labels are, for example, propagated through the graph in order to label all unlabeled nodes. In this paper, we showed how to use random walks to discover structural similarities shared between graphs for the construction of a graph kernel, namely the propagation kernel. Intuitively, propagation kernels count common sub-distributions induced in each iteration of running inference in two graphs leading to the insight that graph kernels are much closer to graph-based learning than assumed before.
As our experimental results demonstrate, propagation kernels are competitive in terms of accuracy with state-of-the-art kernels on several classification benchmark datasets of labeled and attributed graphs. In terms of runtime, propagation kernels outperform all recently developed efficient and scalable graph kernels. Moreover, being tied to the propagation scheme, propagation kernels can be easily adapted to novel applications not having been tractable for graph kernels before.
Propagation kernels provide several interesting avenues for future work. For handling continuous attributes using mixture of different covariance matrices could improve performance. Also, the effect that adding noise to the label encoding actually improves the predictive performance should be investigated in more details. While we have used classification to guide the development of propagation kernels, the results are directly applicable to regression, clustering, and ranking, among other tasks. Employing message-based probabilistic inference schemes such as (loopy) belief propagation directly paves the way to deriving novel graph kernels from graphical models. Exploiting that graph kernels and graph-based learning (learning on the node level) are closely related, hence, a natural extension to this work is the derivation of a unifying propagation-based framework for structure representation independent of the learning task being on the graph or node level.
Notes
This prior could also be the output of an external classifier built on available node attributes.
Note that not both label and attribute information have to be present and both could also be partially observed.
Using a coordinate list sparse representation, the memory usage per pixel grid graph for Algorithm 3 is \(\mathcal {O}(3m_1m_2p)\), where \(m_1 \times m_2\) are the grid dimensions and p is the size of the pixel neighborhood.
All datasets are publicly available for download from http://tiny.cc/PK_MLJ_data.
The test suites provided with the data are incorrect; we use a corrected version.
References
Bauckhage, C., & Kersting, K. (2013). Efficient information theoretic clustering on discrete lattices (2013). arxiv:1310.7114.
Borgwardt, K., & Kriegel, H. P. (2005). Shortest-path kernels on graphs. In Proceedings of international conference on data mining (ICDM-05), pp. 74–81.
Datar, M., & Indyk, P. (2004). Locality-sensitive hashing scheme based on p-stable distributions. In Proceedings of the 20th annual symposium on computational geometry (SCG-2004), pp. 253–262.
Debnath, A., de Compadre, R. L., Debnath, G., Schusterman, A., & Hansch, C. (1991). Structure-activity relationship of mutagenic aromatic and heteroaromatic nitro compounds. Correlation with molecular orbital energies and hydrophobicity. Journal of Medicinal Chemistry, 34(2), 786–797.
Desrosiers, C., & Karypis, G. (2009). Within-network classification using local structure similarity. In Proceedings of the European conference on machine learning and knowledge discovery in databases (ECML/PKDD-09), pp. 260–275.
Dobson, P. D., & Doig, A. J. (2003). Distinguishing enzyme structures from non-enzymes without alignments. Journal of Molecular Biology, 330(4), 771–783.
Feragen, A., Kasenburg, N., Petersen, J., de Bruijne, M., & Borgwardt, K. M. (2013). Scalable kernels for graphs with continuous attributes. Advances in Neural Information Processing Systems, 26(NIPS–13), 216–224.
Gallagher, B., Tong, H., Eliassi-Rad, T., & Faloutsos, C. (2008). Using ghost edges for classification in sparsely labeled networks. In Proceedings of the 14th ACM SIGKDD international conference on knowledge discovery and data mining (KDD-08), pp. 256–264.
Gärtner, T., Flach, P. A., & Wrobel, S. (2003). On graph kernels: Hardness results and efficient alternatives. In Proceedings of computational learning theory and kernel machines (COLT-03), pp. 129–143.
Gersho, A., & Gray, R. (1991). Vector quantization and signal compression. Norwell, MA: Kluwer Academic Publishers.
Haralick, R. M., Shanmugam, K., & Dinstein, I. H. (1973). Textural features for image classification. IEEE Transactions on Systems, Man and Cybernetics, SMC–3(6), 610–621.
Harchaoui, Z., & Bach, F. (2007). Image classification with segmentation graph kernels. In CVPR. IEEE Computer Society.
Haussler, D. (1999). Convolution kernels on discrete structures. Tech. rep., Department of Computer Science, University of California, Santa Cruz.
Hido, S., & Kashima, H. (2009). A linear-time graph kernel. In Proceedings of the 9th IEEE international conference on data mining (ICDM-09), pp. 179–188.
Horváth, T., Gärtner, T., & Wrobel, S. (2004). Cyclic pattern kernels for predictive graph mining. In Proceedings of knowledge discovery in databases (KDD-04), pp. 158–167.
Hwang, T., & Kuang, R. (2010). A heterogeneous label propagation algorithm for disease gene discovery. In Proceedings of the SIAM international conference on data mining (SDM-10), pp. 583–594.
Jaakkola, T., & Haussler, D. (1998). Exploiting generative models in discriminative classifiers. Advances in Neural Information Processing Systems, 11(NIPS–98), 487–493.
Jähne, B. (2005). Digital Image Processing (6th ed.). Berlin: Springer.
Jebara, T., Kondor, R., & Howard, A. (2004). Probability product kernels. Journal of Machine Learning Research, 5, 819–844.
Kashima, H., Tsuda, K., & Inokuchi, A. (2003). Marginalized kernels between labeled graphs. In Proceedings of the 20th international conference on machine learning (ICML-03), pp. 321–328.
Kersting, K., Mladenov, M., Garnett, R., & Grohe, M. (2014). Power iterated color refinement. In Proceedings of the 28th AAAI conference on artificial intelligence (AAAI-14), pp. 1904–1910.
Kondor, R., & Jebara, T. (2003). A kernel between sets of vectors. In Proceedings of the twentieth international conference on machine learning (ICML-03), pp. 361–368.
Kondor, R. I., & Lafferty, J. D. (2002). Diffusion kernels on graphs and other discrete input spaces. In Proceedings of the nineteenth international conference on machine learning (ICML-02), pp. 315–322.
Kriege, N., & Mutzel, P. (2012). Subgraph matching kernels for attributed graphs. In Proceedings of the 29th international conference on machine learning (ICML-12).
Lafferty, J., & Lebanon, G. (2002). Information diffusion kernels. Advances in Neural Information Processing Systems, 22(NIPS–02), 375–382.
Lin, F., & Cohen, W. W. (2010). Power iteration clustering. In Proceedings of the 27th international conference on machine learning (ICML-10), pp. 655–662.
Lovász, L. (1996). Random walks on graphs: A survey. In D. Miklós, V. T. Sós, & T. Szőnyi (Eds.), Combinatorics, Paul Erdős is Eighty (Vol. 2, pp. 353–398). Budapest: János Bolyai Mathematical Society.
Mahé, P., & Vert, J. P. (2009). Graph kernels based on tree patterns for molecules. Machine Learning, 75(1), 3–35.
Moreno, P., Ho, P., & Vasconcelos, N. (2003). A Kullback–Leibler divergence based kernel for SVM classification in multimedia applications. Advances in Neural Information Processing Systems, it 23(NIPS-03), 1385–1392.
Neumann, M., Garnett, R., & Kersting, K. (2013). Coinciding walk kernels: Parallel absorbing random walks for learning with graphs and few labels. In Asian conference on machine learning (ACML-13), pp. 357–372.
Neumann, M., Hallau, L., Klatt, B., Kersting, K., & Bauckhage, C. (2014). Erosion band features for cell phone image based plant disease classification. In Proceedings of the 22nd international conference on pattern recognition (ICPR-14), pp. 3315–3320.
Neumann, M., Moreno, P., Antanas, L., Garnett, R., & Kersting, K. (2013). Graph kernels for object category prediction in task-dependent robot grasping. In Eleventh workshop on mining and learning with graphs (MLG-13), Chicago, Illinois.
Neumann, M., Patricia, N., Garnett, R., & Kersting, K. (2012). Efficient graph kernels by randomization. In European conference on machine learning and knowledge discovery in databases (ECML/PKDD-12), pp. 378–393.
Ojala, T., Pietikäinen, M., & Mäenpää, T. (2002). Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Transactions on Pattern Analysis and Machine Intelligence, 24(7), 971–987.
Ramon, J., & Gärtner, T. (2003). Expressivity versus efficiency of graph kernels. In Proceedings of the 1st international workshop on mining graphs, trees and sequences, pp. 65–74.
Schomburg, I., Chang, A., Ebeling, C., Gremse, M., Heldt, C., Huhn, G., et al. (2004). Brenda, the enzyme database: Updates and major new developments. Nucleic Acids Research, 32(Database–Issue), 431–433.
Shervashidze, N., Schweitzer, P., van Leeuwen, E., Mehlhorn, K., & Borgwardt, K. (2011). Weisfeiler–Lehman graph kernels. Journal of Machine Learning Research, 12, 2539–2561.
Shervashidze, N., Vishwanathan, S., Petri, T., Mehlhorn, K., & Borgwardt, K. (2009). Efficient graphlet kernels for large graph comparison. Journal of Machine Learning Research—Proceedings Track, 5, 488–495.
Shi, Q., Petterson, J., Dror, G., Langford, J., Smola, A. J., & Vishwanathan, S. V. N. (2009). Hash kernels for structured data. Journal of Machine Learning Research, 10, 2615–2637.
Szummer, M., & Jaakkola, T. (2001). Partially labeled classification with Markov random walks. Advances in Neural Information Processing Systems, 15(NIPS–01), 945–952.
Valkealahti, K., & Oja, E. (1998). Reduced multidimensional co-occurrence histograms in texture classification. IEEE Transactions on Pattern Analysis and Machine Intelligence, 20(1), 90–94.
Vishwanathan, S., Schraudolph, N., Kondor, R., & Borgwardt, K. (2010). Graph kernels. Journal of Machine Learning Research, 11, 1201–1242.
Wale, N., & Karypis, G. (2006). Comparison of descriptor spaces for chemical compound retrieval and classification. In Proceedings of the international conference on data mining (ICDM-06) (pp. 678–689). Silver Spring, MD: IEEE Computer Society.
Winn, J. M., Criminisi, A., & Minka, T. P. (2005). Object categorization by learned universal visual dictionary. In 10th IEEE international conference on computer vision (ICCV-05), pp. 1800–1807.
Wu, X. M., Li, Z., So, A. M. C., Wright, J., & Chang, S. F. (2012). Learning with partially absorbing random walks. Advances in Neural Information Processing Systems, 26(NIPS–12), 3086–3094.
Zhou, D., Bousquet, O., Lal, T. N., Weston, J., & Schölkopf, B. (2003). Learning with local and global consistency. Advances in Neural Information Processing Systems, 17(NIPS–03), 321–328.
Zhu, X., Ghahramani, Z., & Lafferty, J. D. (2003). Semi-supervised learning using Gaussian fields and harmonic functions. In Proceedings of the twentieth international conference on machine learning (ICML-03), pp. 912–919.
Acknowledgments
Part of this work grew out of the dfg Collaborative Research Center sfb 876 “Providing Information by Resource-Constrained Analysis” and discussions with Petra Mutzel and Christian Sohler. We thank Nino Shervashidze, Nils Kriege, Aasa Feragen, and Karsten Borgwardt for our discussions and for sharing their kernel implementations and datasets. Part of this work was supported by the German Science Foundation (dfg) under the reference number ‘GA 1615/1-1’.
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: Karsten Borgwardt.
Appendices
Appendix 1: Runtimes for Partially Labeled Graphs
See Fig. 12.
Runtime for partially labeled msrc21. Avg. time in seconds over 10 instances of the msrc21 dataset with 50 % labeled nodes for kernel iterations from 0 to 10. We compare pk to wl with unlabeled nodes treated as additional label and with hard labels derived from converged label propagation (\(\textsc {lp}+\textsc {wl}\))
Appendix 2: Detailed Results on Attributed Graphs
Rights and permissions
About this article
Cite this article
Neumann, M., Garnett, R., Bauckhage, C. et al. Propagation kernels: efficient graph kernels from propagated information. Mach Learn 102, 209–245 (2016). https://doi.org/10.1007/s10994-015-5517-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10994-015-5517-9