Abstract
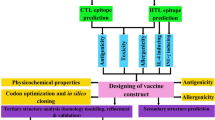
Breast cancer (BC) is the most common type of women’s cancer with a prevalence of about 25%, although it is rare in men (< 1%). Until now, BC therapy approaches have not been successful for various reasons. The purpose of this study was to design a candidate vaccine against BC using bioinformatics tools. Herein, in silico design of a fusion vaccine candidate containing extracellular domain of HER-2/neu protein and N-terminal sequence of GP96 protein (NTGP96) were performed. Different bioinformatics and immunoinformatics softwares were used to study the important characteristics of the designed vaccine, including physicochemical properties, secondary, and 3D structures, as well as antigenicity and allergenicity. Structural and immunological computational results indicated the potential of our designed construct for proper stimulation of cellular and humoral immune responses against BC. According to the obtained data, the candidate vaccine could be introduced for BC therapy, after the efficacy of it was confirmed by in vivo and in vitro immunological assays.







Similar content being viewed by others
References
Akhoon BA, Gupta SK, Verma V, Dhaliwal G, Srivastava M, Gupta SK, Ahmad RF (2010) In silico designing and optimization of anti-breast cancer antibody mimetic oligopeptide targeting HER-2 in women. J Mol Graph Model 28(7):664–669
Andreatta M, Nielsen M (2015) Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 32:511–517
Ansari HR, Raghava GP (2010) Identification of conformational B-cell epitopes in an antigen from its primary sequence. Immun Res 6:6
Argos P (1990) An investigation of oligopeptides linking domains in protein tertiary structures and possible candidates for general gene fusion. J Mol Biol 211:943–958
Ashokan K, Pillai M (2008) In silico characterization of silk fibroin protein using computational tools and servers. Asian J Exp Sci 22265–22274
Atapour A, Mokarram P, Mostafavi-Pour Z, Ramezani A (2017) Molecular cloning, expression, and purification of a recombinant fusion protein (rNT-gp96-NT300). BioPharm Int 30:38–44
Atapour A, Mokarram P, MostafaviPour Z, Hosseini SY, Ghasemi Y, Mohammadi S, Nezafat N (2018) Designing a fusion protein vaccine against HCV: an in silico approach. Int J Pept Res Ther. https://doi.org/10.1007/s10989-018-9735-4
Baloria U, Akhoon BA, Gupta SK, Sharma S, Verma V (2012) In silico proteomic characterization of human epidermal growth factor receptor 2 (HER-2) for the mapping of high affinity antigenic determinants against breast cancer. Amino Acids 42(4):1349–1360
Bhasin M, Raghava G (2004) Prediction of CTL epitopes using QM, SVM and ANN techniques. Vaccine 22:3195–3204
Billod J-M, Lacetera A, Guzmán-Caldentey J, Martín-Santamaría S (2016) Computational approaches to toll-like receptor 4 modulation. Molecules 21:994
Bjellqvist B et al (1993) The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. Electrophoresis 14:1023–1031
Björklund ÅK, Soeria-Atmadja D, Zorzet A, Hammerling U, Gustafsson MG (2004) Supervised identification of allergen-representative peptides for in silico detection of potentially allergenic proteins. Bioinformatics 21:39–50
Buchan DW, Minneci F, Nugent TC, Bryson K, Jones DT (2013) Scalable web services for the PSIPRED protein analysis work bench nucleic acids research 41:W349–W357
Caballero MT et al (2015) TLR4 genotype and environmental LPS mediate RSV bronchiolitis through Th2 polarization. J Clin Investig 125:571–582
Calderwood SK, Gong J, Murshid A (2016) Extracellular HSPs: the complicated roles of extracellular HSPs in immunity. Front Immunol 7:159
Chauhan N, Tiwari S, Iype T, Jain U (2017) An overview of adjuvants utilized in prophylactic vaccine formulation as immunomodulators. Exp Rev Vaccines 16:491–502
Chen X, Zaro JL, Shen W-C (2013) Fusion protein linkers: property, design and functionality. Adv Drug Deliv Rev 65:1357–1369
Childers NK, Miller KL, Tong G, Llarena JC, Greenway T, Ulrich JT, Michalek SM (2000) Adjuvant activity of monophosphoryl lipid A for nasal and oral immunization with soluble or liposome-associated antigen. Infect Immun 68:5509–5516
Cluff CW (2009) Monophosphoryl lipid A (MPL) as an adjuvant for anti-cancer vaccines: clinical results. In: Lipid A in cancer therapy. Springer, New York, pp 111–123
Colovos C, Yeates T (1993) ERRAT: an empirical atom-based method for validating protein structures. Protein Sci 2:1511–1519
Conniot J et al (2014) Cancer immunotherapy: nanodelivery approaches for immune cell targeting and tracking. Front Chem 2:105
Dhanda SK, Vir P, Raghava GP (2013) Designing of interferon-gamma inducing MHC class-II binders. Biol Direct 8:30
Didierlaurent AM et al (2009) AS04, an aluminum salt-and TLR-4 agonist-based adjuvant system, induces a transient localized innate immune response leading to enhanced adaptive immunity. J Immunol 183:6186–6197
Dimitrov I, Bangov I, Flower DR, Doytchinova I (2014) AllerTOP v. 2—a server for in silico prediction of allergens. J Mol Model 20:2278
Doytchinova IA, Flower DR (2007) VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform 8:4
Doytchinova IA, Flower DR (2018) In silico prediction of cancer immunogens: current state of the art. BMC Immunol 19:11
EL-Manzalawy Y, Dobbs D, Honavar V (2008) Predicting linear B-cell epitopes using string kernels. J Mol Recogn Interdiscip 21:243–255
Emens LA (2008) Cancer vaccines: on the threshold of success. Exp Opin Emerg Drugs 13:295–308
Fooladi AAI, Hosseini HM, Amani J (2015) An in silico chimeric vaccine targeting breast cancer containing inherent adjuvant. Iran J Cancer Prev 8:3
Gasteiger E, Hoogland C, Gattiker A, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: The proteomics protocols handbook. Springer, New York, pp 571–607
Grote A, Hiller K, Scheer M, Münch R, Nörtemann B, Hempel DC, Jahn D (2005) JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res 33:W526–W531
Hajizadeh MR, Mokarram P (2013) Recombinant nonstructural 3 protein, rNS3, of hepatitis C virus along with recombinant GP96 induce IL-12, TNFα and α5integrin expression in antigen presenting cells. Hepat Mon 13:6
Haste Andersen P, Nielsen M, Lund O (2006) Prediction of residues in discontinuous B-cell epitopes using protein 3D structures. Protein Sci 15:2558–2567
Hosseinzadeh S, Daemi A (2012) Heat shock proteins as the efficient vehicle in cancer. J Solid Tumors 2:47
Hu Z, Ott PA, Wu CJ (2018) Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat Rev Immunol 18:168
Ikai A (1980) Thermostability and aliphatic index of globular proteins. J Biochem 88:1895–1898
Imani-Fooladi AA, Yousefi F, Mousavi SF, Amani J (2014) In silico design and analysis of TGFαl3-seb fusion protein as “a new antitumor agent” candidate by ligand-targeted superantigens technique. Iran J Cancer Prev 7:152
Jahangiri A, Amani J, Halabian R (2018) In silico analyses of staphylococcal enterotoxin B as a DNA vaccine for cancer therapy. Int J Pept Res Ther 24:131–142
Jensen KK et al (2018) Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology 154:394–406
Khatoon N, Ojha R, Mishra A, Prajapati VK (2018) Examination of antigenic proteins of Trypanosoma cruzi to fabricate an epitope-based subunit vaccine by exploiting epitope mapping mechanism. Vaccine 36:6290–6300
Li SG, Li L (2013) Targeted therapy in HER2-positive breast cancer. Biomed Rep 1:499–505
Li Z, Menoret A, Srivastava P (2002) Roles of heat-shock proteins in antigen presentation and cross-presentation. Curr Opin Immunol 14:45–51
Liu W et al (2016) Interaction of toll-like receptors with the molecular chaperone Gp96 is essential for its activation of cytotoxic T lymphocyte response. PLoS ONE 11:e0155202
Lovell SC et al (2003) Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins Struct Funct Bioinform 50:437–450
Magnan CN, Randall A, Baldi P (2009) SOLpro: accurate sequence-based prediction of protein solubility. Bioinformatics 25:2200–2207
Magnan CN, Zeller M, Kayala MA, Vigil A, Randall A, Felgner PL, Baldi P (2010) High-throughput prediction of protein antigenicity using protein microarray data. Bioinformatics 26:2936–2943
Mahdavi M, Mohabatkar H, Keyhanfar M, Dehkordi AJ, Rabbani M (2012) Linear and conformational B cell epitope prediction of the HER 2 ECD-subdomain III by in silico methods. Asian Pac J Cancer Prev 13:3053–3059
Mahdavi M, Keyhanfar M, Jafarian A, Mohabatkar H, Rabbani M (2015) Production and characterization of new anti-HER2 monoclonal antibodies. Monoclon Antib Immunodiagn Immunother 34:213–221
Mohammadi S et al (2018) In silico analysis of different signal peptides for the excretory production of recombinant NS3-GP96 fusion protein in Escherichia coli. Int J Pept Res Ther. https://doi.org/10.1007/s10989-018-9775-9
Mohit E, Bolhassani A, Zahedifard F, Taslimi Y, Rafati S (2012) The contribution of NT-gp96 as an adjuvant for increasing HPV16 E7-specific immunity in C57BL/6 mouse model. Scand J Immunol 75:27–37
Nezafat N, Ghasemi Y, Javadi G, Khoshnoud MJ, Omidinia E (2014) A novel multi-epitope peptide vaccine against cancer: an in silico approach. J Theor Biol 349:121–134
Nezafat N, Eslami M, Negahdaripour M, Rahbar MR, Ghasemi Y (2017) Designing an efficient multi-epitope oral vaccine against Helicobacter pylori using immunoinformatics and structural vaccinology approaches. Mol BioSyst 13:699–713
Nielsen M, Lundegaard C, Lund O (2007) Prediction of MHC class II binding affinity using SMM-align, a novel stabilization matrix alignment method. BMC Bioinform 8(1):238
Nold-Petry CA et al (2017) gp96 peptide antagonist gp96-ii confers therapeutic effects in murine intestinal inflammation. Front Immunol 8:1531
Ozturk OG (2012) HLA alleles in breast cancer. Pathol Oncol Res 18:1099–1099
Palladini A et al (2018) Virus-like particle display of HER2 induces potent anti-cancer responses. Oncoimmunology 7:e1408749
Pandey RK, Bhatt TK, Prajapati VK (2018) Novel immunoinformatics approaches to design multi-epitope subunit vaccine for malaria by investigating anopheles salivary protein. Sci Rep 8:1125
Pihlasalo S, Auranen L, Hänninen P, Härmä H (2012) Method for estimation of protein isoelectric point. Anal Chem 84:8253–8258
Ponomarenko J, Bui HH, Li W, Fusseder N, Bourne PE, Sette A, Peters B (2008 Dec) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinform 9(1):514
Quevedo-Diaz MA, Song C, Xiong Y, Chen H, Wahl LM, Radulovic S, Medvedev AE (2010) Involvement of TLR2 and TLR4 in cell responses to Rickettsia akari. J Leukoc Biol 88:675–685
Rapp UK, Kaufmann SH (2004) DNA vaccination with gp96-peptide fusion proteins induces protection against an intracellular bacterial pathogen. Int Immunol 16:597–605
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725
Saha S, Raghava G (2006) AlgPred: prediction of allergenic proteins and mapping of IgE epitopes. Nucleic Acids Res 34:W202–W209
Sen Gupta P, Mandal B, Bandyopadhyay A (2013) In silico characterization of human cyclooxygenase using computational tools and servers. Int J Inst Pharm Life Sci 3:111
Shah P, Mistry J, Reche PA, Gatherer D, Flower DR (2018) In silico design of Mycobacterium tuberculosis epitope ensemble vaccines. Mol Immunol 97:56–62
Shevtsov M, Multhoff G (2016) Heat shock protein-peptide and HSP-based immunotherapies for the treatment of cancer. Front Immunol 7:171
Shin W-H, Lee GR, Heo L, Lee H, Seok C (2014) Prediction of protein structure and interaction by GALAXY protein modeling programs. Bio Des 2:1–11
Shirota H, Tross D, Klinman D (2015) CpG oligonucleotides as cancer vaccine adjuvants. Vaccines 3:390–407
Sivakumar K, Balaji S (2007) In silico characterization of antifreeze proteins using computational tools and servers. J Chem Sci 119:571–579
Skwarczynski M, Toth I (2016) Peptide-based synthetic vaccines. Chem Sci 7:842–854
Soria-Guerra RE, Nieto-Gomez R, Govea-Alonso DO, Rosales-Mendoza S (2015) An overview of bioinformatics tools for epitope prediction: implications on vaccine development. J Biomed Inform 53:405–414
Steinhagen F, Kinjo T, Bode C, Klinman DM (2011) TLR-based immune adjuvants. Vaccine 29:3341–3355
Tai W, Mahato R, Cheng K (2010) The role of HER2 in cancer therapy and targeted drug delivery. J Controll Release 146:264–275
Torchala M, Bates PA (2014) Predicting the structure of protein–protein complexes using the SwarmDock web server. In: Protein structure prediction. Springer, New York, pp 181–197
Waterhouse A et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410
Zhao W, Sher X (2018) Systematically benchmarking peptide-MHC binding predictors: from synthetic to naturally processed epitopes. PLoS Comput Biol 14:e1006457
Zheng J et al (2017) In silico analysis of epitope-based vaccine candidates against hepatitis B virus polymerase protein. Viruses 9:112
Zininga T, Ramatsui L, Shonhai A (2018) Heat shock proteins as immunomodulants. Molecules 23:2846
Acknowledgements
The authors wish to thank Shiraz University of Medical Sciences for supporting the conduct of this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have none conflict of interest.
Research Involving Human and Animal Participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Atapour, A., Negahdaripour, M., Ghasemi, Y. et al. In Silico Designing a Candidate Vaccine Against Breast Cancer. Int J Pept Res Ther 26, 369–380 (2020). https://doi.org/10.1007/s10989-019-09843-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-019-09843-1