Abstract
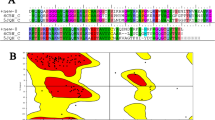
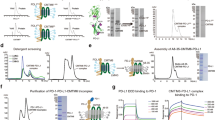
One major issue in cancer immunotherapy is the large size of mAbs (150 kDa) which usually causes limitation in tumor penetration. Camel nanobodies (< 15 kDa) could be used as an alternative for this problem. Here in, using molecular modeling we designed a novel recombinant chimeric protein containing the extra membrane loop of human CD20 which was fused to the FC region of camel with proper potential for production of camel derived nanobodies. Molecular dynamics simulation showed that the recombinant chimera can interact specifically with therapeutic antibody Rituximab Fab and the interaction site was the residue 164IYNCEPANPSE174. The recombinant chimeric protein was successfully expressed in HEK293-T cells and evaluated in vitro using western blotting analysis. In agreement with simulation results, western blotting analysis also confirmed the interactions between Rituximab and hCD20. The results of this paper show that protein engineering with the help of molecular modelling tools can be promising and efficient approach in developing new nanobodies for caner immunotherapy.



Similar content being viewed by others
References
Ahmedin J, Freddie B, Melissa C, Jacques F, Elizabeth W, David F (2011) Global cancer statistics. CA Cancer J Clin 61(2):69–90. https://doi.org/10.3322/caac.20107
Anbouhi MH, Barazandeh AF, Bouzari S et al (2012) Functional recombinant extra membrane loop of human CD20, an alternative of the full length CD20 antigen. Iran Biomed J 16(3):12–16. https://doi.org/10.6091/ibj.1082.2012
Anderson KC, Bates MP, Slaughenhoupt BL, Pinkus GS, Schlossman SF, Nadler LM (1984) Expression of human B cell-associated antigens on leukemias and lymphomas: a model of human B cell differentiation. Blood 63:1424–1433
Ansari HR, Raghava GP (2010) Identification of conformational B-cell epitopes in an antigen from its primary sequence. Immunome Res 6:1186–1186. https://doi.org/10.1186/1745-7580-6-6
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56. https://doi.org/10.1016/0010-4655(95)00042-E
Binder M, Otto F, Mertelsmann R et al (2006) The epitope recognized by rituximab. Blood 108(6):1975–1978. https://doi.org/10.1182/blood-2006-04-014639
Boye J, Elter T, Engert AL (2003) An overview of the current clinical use of the anti-CD20 monoclonal antibody rituximab. Ann Oncol 14(4):520–535. https://doi.org/10.1093/annonc/mdg175
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Broisat A, Hernot S, Toczek J et al (2012) Nanobodies targeting mouse/human VCAM1 for the nuclear imaging of atherosclerotic lesions. Circ Res 110(7):927–937. https://doi.org/10.1161/CIRCRESAHA.112.265140
Bubien JK, Zhou LJ, Bell PD, Frizzell RA, Tedder TE (1993) Transfection of the CD20 cell surface molecule into ectopic cell types generates a Ca conductance found constitutively in B lymphocytes. Cell Biol 121(5):1121–1132. https://doi.org/10.1083/jcb.121.5.1121
Choi HS, Liu W, Misra P et al (2007) Renal clearance of nanoparticles. Nat Biotechnol 25(10):1165–1170. https://doi.org/10.1038/nbt1340
Darden T, York, Pedersen L (1993) Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. https://doi.org/10.1063/1.464397
De Marco A (2009) Strategies for successful recombinant expression of disulfide bond-dependent proteins in Escherichia coli. Microb Cell Fact 8:26. https://doi.org/10.1186/1475-2859-8-26
DeLano WL (2002) The PyMOL molecular graphics system. http://pymol.org
Du J, Wang H, Zhong C et al (2007) Structural basis for recognition of CD20 by therapeutic antibody rituximab. J Biol Chem 282(20):15073–15080. https://doi.org/10.1074/jbc.M701654200
Ernst JA, Li H, Kim HS, Nakamura GR, Yansura DG, Vandlen RL (2005) Isolation and characterization of the B-cell marker CD20. Biochemistry 44(46):15150–15158. https://doi.org/10.1021/bi0511078
Eswar N, Webb B, Marti-Renom MA et al (2006) Comparative protein structure modeling using modeller. Curr Protoc Bioinform 5:1–47. https://doi.org/10.1002/0471250953.bi0506s15
Hermans J, Berendsen HJ, Van Gunsteren WF et al (1984) A consistent empirical potential for water–protein interactions. Biopolymers 23:1513–1518. https://doi.org/10.1002/bip.360230807
Hess B, Bekker H, Berendsen JC et al (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3C1463::AID-JCC4%3E3.0.CO;2-H
Holliger P, Hudson PJ (2005) Engineered antibody fragments and the rise of single domains. Nat Biotechnol 23:1126–1136. https://doi.org/10.1038/nbt1142
Hong HY, Sun YX, Guo YX et al (2000) Cloning and expression of human CD20 gene on NIH-3T3 cell membrane. Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao 32(4):430–433
Karplus M, McCammon JA (2002) Molecular dynamics simulations of biomolecules. Nat Struct Mol Biol 9(9):646–652. https://doi.org/10.1021/ar020082r
Mark J. Adler, Dimiter S, Dimitrov (2012) Therapeutic antibodies against cancer. Hematol Oncol Clin North Am 26(3):447–481. https://doi.org/10.1016/j.hoc.2012.02.013
Meng XY, Zhang HX, Mezei M, Cui M (2011) Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des 7(2):146–157
Motta G, Cea M, Moran E, Carbone F, Augusti V, Patrone F, Nencioni A (2010) Monoclonal antibodies for non-Hodgkin’s lymphoma: state of the art and perspectives. Clin Dev Immunol 20:1–14. https://doi.org/10.1155/2010/428253
Oscherwitz J, Gribbin TE, Cease KB (2010) A CD20 tandemepitope immunogen elicits antibody in mice that binds murine cell surface CD20 and depletes splenic B cells in vivo. Mol Immunol 47(7–8):1484–1491. https://doi.org/10.1016/j.molimm.2010.01.026
Parkin DM, Whelan SL, Ferlay J, Teppo L, Thomas DB (2003) Cancer Incidence in Five Continents. VIII. Lyon: International Agency for Research on Cancer
Rahbarizadeh F, Ahmadvand D, Sharifzadeh Z (2011) Nanobody; an old concept and new vehicle for immunotargeting. Immunol Invest 40(3):299–338. https://doi.org/10.3109/08820139.2010.542228
Schmid N, Eichenberger AP, Choutko A et al (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40(7):843–856. https://doi.org/10.1007/s00249-011-0700-9
Zhang XY, Sun ZW, Yu WY, Cheng JZ (2004) Cloning and expression of fusion gene of transmembrane domain of human CD20 and g3pN in Escherichia coli. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 20(4):481–483
Acknowledgements
We are very grateful to Dr Javadmanesh and Miss Marjan Azghandi for their outstanding technical assistance. Financial support of this study has been provided by Ferdowsi University of Mashhad by Grant Number 25742.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All Authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Hosseini, S.A., Tahmoorespur, M., Sekhavati, M.H. et al. Designing of a Functional Chimeric Protein for Production of Nanobodies Against Human CD20: Molecular Dynamics Simulation and In Vitro Verification. Int J Pept Res Ther 25, 1459–1465 (2019). https://doi.org/10.1007/s10989-018-9791-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-018-9791-9