Abstract
Context
Ensemble of small models (ESMs) is a technique to overcome the problem of few occurrence points. Applying the ESMs in a spatially hierarchical framework could increase the accuracy of predictions and conclusions by restricting available habitat at sequentially finer spatial scales.
Objectives
Our objective was to show how applying ESMs in a hierarchical habitat selection framework could help to understand rare species’ niches at various scales. We compared the accuracy of ESMs made by committee averaging and weighted averaging methods. We also compared the predictive power of ESMs made by various modeling techniques for Virginia’s warbler (Leiothlypis virginiae) at its northeastern range limit.
Methods
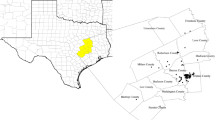
We defined biologically relevant hierarchical orders of habitat selection for Virginia’s warbler in the Black Hills, U.S.A. We modeled habitat suitabity at the broadest scale as a function of bioclimatic covariates and at finer scales as functions of landcover, soil group and landscape covariates.
Results
The performance of modeling techniques varied among scales. Using the committee averaging method led to more accurate results than weighted averaging. At the broadest order, Virginia’s warbler had a narrow climatic niche. The importance of covariates changed across finer orders, such that at broader orders many covariates were important whereas at finer orders certain covariates became more important than others.
Conclusion
We conclude that applying ESMs within a hierarchical framework can lead to detailed information about rare species’ niches, limiting factors at each habitat selection order, and potential distribution, which could help inform multiscale management.



Similar content being viewed by others
References
Allouche O, Tsoar A, Kadmon R (2006) Assessing the accuracy of species distribution models: prevalence, kappa and the true skill statistic (TSS). J Appl Ecol 43:1223–1232
Araújo MB, New M (2007) Ensemble forecasting of species distributions. Trends Ecol Evol 22:42–47
Banfield JD, Raftery AE (1993) Model-based Gaussian and non-Gaussian clustering. Biometrics 49:803–821
Bellamy C, Scott C, Altringham J (2013) Multiscale, presence-only habitat suitability models: fine-resolution maps for eight bat species. J Appl Ecol 50:892–901
Boria RA, Olson LE, Goodman SM, Anderson RP (2014) Spatial filtering to reduce sampling bias can improve the performance of ecological niche models. Ecol Modell 275:73–77
Boyce MS, Mao JS, Merrill EH, Fortin D, Turner MG, Fryxell J, Turchin P (2003) Scale and heterogeneity in habitat selection by elk in Yellowstone National Park. Écoscience 10:421–431
Breiman L (2001) Random forests. Mach Learn 45:5–32
Breiner FT, Nobis MP, Bergamini A, Guisan A (2018) Optimizing ensembles of small models for predicting the distribution of species with few occurrences. Methods Ecol Evol 9:802–808
Brotons L, Thuiller W, Araújo MB, Hirzel AH (2004) Presence-absence versus presence-only modelling methods for predicting bird habitat suitability. Ecography (Cop) 27:437–448
Bubac CM, Spellman GM (2016) How connectivity shapes genetic structure during range expansion: Insights from the Virginia’s Warbler. Auk 133:213–230
Bürkner P-C (2017) brms: an R package for Bayesian multilevel models using stan. J Stat Softw 1(1):1–8
Dasgupta A, Raftery AE (1998) Detecting features in spatial point processes with clutter via model-based clustering. J Am Stat Assoc 93:294–302
Della Rocca F, Bogliani G, Breiner FT, Milanesi P (2019) Identifying hotspots for rare species under climate change scenarios: improving saproxylic beetle conservation in Italy. Biodivers Conserv 28:433–449
Devictor V, Julliard R, Jiguet F (2008) Distribution of specialist and generalist species along spatial gradients of habitat disturbance and fragmentation. Oikos 117:507–514
Di Febbraro M, Menchetti M, Russo D et, Ancillotto L, Aloise G, Roscioni F, Preatoni DG, Loy A, Martinoli A, Bertolino S, Mori E (2019) Integrating climate and land-use change scenarios in modelling the future spread of invasive squirrels in Italy. Divers Distrib 25:644–659
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G, García Marquéz JR, Gruber B, Lafourcade B, Leitão PJ, Münkemüller T, McClean C, Osborne PE, Reineking B, Schröder B, Skidmore AK, Zurell D, Lautenbach S (2013) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography (Cop) 36:27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x
Elith J, Graham CH, Anderson RP, Dudík, Ferrier SDM, Guisan A, Hijmans RJ, Huettmann F, Leathwick JR, Lehmann A, Li J, Lohmann LG, Loiselle BA, Manion G, Moritz C, Nakamura M, Nakazawa Y, Overton McCMJ, Townsend Peterson A, Phillips JS, Richardson K, Scachetti‐Pereira R, Schapire RE, Soberón J, Williams S, Wisz MS, Zimmermann NE (2006) Novel methods improve prediction of species’ distributions from occurrence data. Ecography (Cop) 29:129–151
ESRI (2017) ArcGIS desktop: release 107. Environmental Systems Research Institute, Redlands
Ferrier S, Drielsma M, Manion G, Watson G (2002) Extended statistical approaches to modelling spatial pattern in biodiversity in northeast New South Wales II Community-level modelling. Biodivers Conserv 11:2309–2338
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315
Fielding AH, Bell JF (1997) A review of methods for the assessment of prediction errors in conservation presence/absence models. Environ Conserv 24:38–49
Fischer JM (1978) A natural history study of the Virginia’s Warbler. Northern Arizona University, Arizona
Fletcher R, Fortin M-J (2018) Space use and resource selection. Spatial ecology and conservation modeling: applications with R. Springer International Publishing, Cham, pp 271–320
Freeman EA, Moisen G (2008) Presence absence: an r package for presence absence analysis. J Stat Softw 1:11
Friedman JH (1991) Multivariate adaptive regression splines. Ann Stat 19:1–67
Gaston K, Blackburn T (2008) Pattern and process in macroecology. Blackwell Publishing Inc, Maden
Ghosh J, Li Y, Mitra R (2018) On the use of cauchy prior distributions for bayesian logistic regression. Bayesian Anal 13:359–383
Goljani Amirkhiz R, Frey JK, Cain JW, Breck SW, Bergman DL (2018) Predicting spatial factors associated with cattle depredations by the Mexican wolf (Canis lupus baileyi) with recommendations for depredation risk modeling. Biol Conserv. https://doi.org/10.1016/j.biocon.2018.06.013
Guillera-Arroita G, Lahoz-Monfort JJ, Elith J Gordon A, Kujala H, Lentini PE, McCarthy MA, Tingley R, Wintle BA (2015) Is my species distribution model fit for purpose? Matching data and models to applications. Glob Ecol Biogeogr 24:276–292
Guisan A, Thuiller W (2005) Predicting species distribution: offering more than simple habitat models. Ecol Lett 8:993–1009
Guisan A, Broennimann O, Engler R, Vust M, Yoccoz NG, Lehmann A, Zimmermann NE (2006) Using niche-based models to improve the sampling of rare species. Conserv Biol 20:501–511
Guisan A, Tingley R, Baumgartner JB et al (2013) Predicting species distributions for conservation decisions. Ecol Lett 16:1424–1435
Guisan A, Thuiller W, Zimmermann NE (2017) Habitat suitability and distribution models: with applications in R. Cambridge University Press, Cambridge
Habibzadeh N, Ludwig T (2019) Ensemble of small models for estimating potential abundance of Caucasian grouse (Lyrurus mlokosiewiczi) in Iran. Ornis Fenn 96:77–89
Hastie T, Tibshirani R (1987) Generalized additive models: some applications. J Am Stat Assoc 82:371–386
Hernandez PA, Graham CH, Master LL, Albert DL (2006) The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography (Cop) 29:773–785
Hijmans RJ, Phillips S, Leathwick J, Elith J (2017) Species distribution modeling. R package version 1.1–4
Hirzel AH, Le Lay G, Helfer V, Randin Ch, Guisan A (2006) Evaluating the ability of habitat suitability models to predict species presences. Ecol Modell 199:142–152
Hooten MB, Hobbs NT (2015) A guide to Bayesian model selection for ecologists. Ecol Monogr 85:3–28
Jackson HB, Fahrig L (2015) Are ecologists conducting research at the optimal scale? Glob Ecol Biogeogr 24:52–63
Jedlikowski J, Chibowski P, Karasek T, Brambilla M (2016) Multi-scale habitat selection in highly territorial bird species: exploring the contribution of nest, territory and landscape levels to site choice in breeding rallids (Aves: Rallidae). Acta Oecologica 73:10–20
Johnson DH (1980) The comparison of usage and availability measurements for evaluating resource preference. Ecology 61:65–71
Liu C, White M, Newell G (2013) Selecting thresholds for the prediction of species occurrence with presence-only data. J Biogeogr 40:778–789
Liu C, Newell G, White M (2016) On the selection of thresholds for predicting species occurrence with presence-only data. Ecol Evol 6:337–348
Lomba A, Pellissier L, Randin C, Vicente J, Moreira F, Honrado J, Guisan A (2010) Overcoming the rare species modelling paradox: a novel hierarchical framework applied to an Iberian endemic plant. Biol Conserv 143:2647–2657
MacFaden SW, Capen DE (2002) Avian habitat relationships at multiple scales in a New England forest. For Sci 48:243–253
Manly BF, McDonald L, Thomas DL, McDonald TL, Erickson WP (2007) Resource selection by animals: statistical design and analysis for field studies. Springer, Amsterdam
Martínez JA, Serrano D, Zuberogoitia I (2003) Predictive models of habitat preferences for the Eurasian eagle owl Bubo bubo: a multiscale approach. Ecography (Cop) 26:21–28
Mayor SJ, Schneider DC, Schaefer JA, Mahoney SP (2009) Habitat selection at multiple scales. Écoscience 16:238–247
McCune JL (2016) Species distribution models predict rare species occurrences despite significant effects of landscape context. J Appl Ecol 53:1871–1879
McGarigal K, Wan HY, Zeller KA, Timm BC, Cushman SA (2016) Multi-scale habitat selection modeling: a review and outlook. Landsc Ecol 31:1161–1175
Merow C, Smith MJ, Silander JA (2013) A practical guide to MaxEnt for modeling species’ distributions: what it does, and why inputs and settings matter. Ecography (Cop) 36:1058–1069
Meyer CB, Thuiller W (2006) Accuracy of resource selection functions across spatial scales. Divers Distrib 12:288–297
Muscarella R, Galante PJ, Soley-Guardia M, Boria RA, Kass JM, Uriarte M, Anderson RP (2014) ENMeval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5:1198–1205
O’Hara RB, Sillanpaa MJ (2009) A review of Bayesian variable selection methods: what, how and which. Bayesian Anal 4:85–117
O’Neill RV, Deangelis DL, Waide JB, Allen TFH (1986) A hierarchical concept of ecosystems. Princeton University Press, Princeton
Olson CR, Martin TE (1999) Virginia’s Warbler (Oreothlypis virginiae). In: The birds of North America. Cornell Lab of Ornithology, New York
Orr HK (1975) Watershed management in the Black Hills: the status of our knowledge. USDA Forest Service, Fort Collins CO
Pacifici K, Dorazio RM, Conroy MJ (2012) A two-phase sampling design for increasing detections of rare species in occupancy surveys. Methods Ecol Evol 3:721–730
Peterson AT, Papeş M, Soberón J (2008) Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol Modell 213:63–72
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Modell 190:231–259
Phillips SJ, Dudik M, Elith J, Graham CH, Lehmann A, Leathwick J, Ferrier S (2009) Sample selection bias and presence-only distribution models: implications for background and pseudo-absence data. Ecol Appl 19:181–197
Piironen J, Vehtari A (2017) Comparison of Bayesian predictive methods for model selection. Stat Comput 27:711–735
Razgour O, Hanmer J, Jones G (2011) Using multi-scale modelling to predict habitat suitability for species of conservation concern: the grey long-eared bat as a case study. Biol Conserv 144:2922–2930
Rettie WJ, Messier F (2000) Hierarchical habitat selection by woodland caribou: its relationship to limiting factors. Ecography (Cop) 23:466–478
Ripley BD (1996) Pattern recognition and neural networks. Cambridge University Press, Cambridge
Root T (1988) Environmental factors associated with avian distributional boundaries. J Biogeogr 15:489–505
Scherrer D, Christe P, Guisan A (2019) Modelling bat distributions and diversity in a mountain landscape using focal predictors in ensemble of small models. Divers Distrib 25:770–782
Scrucca L, Fop M, Murphy TB, Raftery AE (2016) mclust 5: clustering, classification and density estimation using gaussian finite mixture models. R J 8:289–317
Segurado P, Araújo MB (2004) An evaluation of methods for modelling species distributions. J Biogeogr 31:1555–1568
Suárez-Seoane S, Osborne PE, Alonso JC (2002) Large-scale habitat selection by agricultural steppe birds in Spain: identifying species-habitat responses using generalized additive models. J Appl Ecol 39:755–771
Swanson DL, Palmer JS, Liknes ET, Dean KL (2000) A breeding population of Virginia’s Warblers in the southwestern Black Hills of South Dakota. Southwest Nat 45:39–44
Swanson DL, Dixon MD, Palmer JS (2016) A re-assessment of the distribution of Virginia’s warbler in the Black Hills of South Dakota
Syfert MM, Smith MJ, Coomes DA (2013) The effects of sampling bias and model complexity on the predictive performance of MaxEnt species distribution models. PLoS ONE 8:e55158
Thuiller W, Lafourcade B, Engler R, Araújo MB (2009) BIOMOD – a platform for ensemble forecasting of species distributions. Ecography (Cop) 32:369–373
Van Riper IIIC, Hatten JR, Giermakowski JT, Mattson D, Holmes JA, Johnson MJ, Nowak EM, Ironside K, Peters M, Heinrich P, Cole KL, Truettner C, Schwalbe CR (2014) Projecting climate effects on birds and reptiles of the Southwestern United States. Reston, VA
VanDerWal J, Shoo LP, Johnson CN, Williams SE (2009) Abundance and the environmental niche: environmental suitability estimated from niche models predicts the upper limit of local abundance. Am Nat 174:282–291
Vehtari A, Gelman A, Gabry J (2017) Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Stat Comput 27:1413–1432
Vergara M, Cushman SA, Urra F, Ruiz-González A (2016) Shaken but not stirred: multiscale habitat suitability modeling of sympatric marten species (Martes martes and Martes foina) in the northern Iberian Peninsula. Landsc Ecol 31:1241–1260
Warren DL, Seifert SN (2011) Ecological niche modeling in Maxent: the importance of model complexity and the performance of model selection criteria. Ecol Soc Am 21:335–342
Watanabe S (2010) Asymptotic Equivalence of Bayes Cross Validation and Widely Applicable Information Criterion in Singular Learning Theory. http://arxiv.org/abs/1004.2316
Weihs C, Ligges U, Luebke K, Raabe N (2005) klaR analyzing German business cycles. In: Baier D, Decker R, Schmidt-Thieme L (eds) Data analysis and decision support. Springer-Verlag, Berlin, pp 335–343
Wilson KA, Westphal MI, Possingham HP, Elith J (2005) Sensitivity of conservation planning to different approaches to using predicted species distribution data. Biol Conserv 122:99–112
Zimmerman GS, Gutiérrez RJ, Thogmartin WE, Banerjee S (2009) Multiscale habitat selection by ruffed grouse at low population densities. Condor 111:294–304
Acknowledgements
We would like to thank J. Wesner for providing feedback regarding model construction. This work was funded by NSF OIA-1632810 and a wildlife diversity Grant (UP1500121) from the South Dakota Department of Game, Fish and Parks.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Amirkhiz, R.G., Dixon, M.D., Palmer, J.S. et al. Investigating niches and distribution of a rare species in a hierarchical framework: Virginia’s Warbler (Leiothlypis virginiae) at its northeastern range limit. Landscape Ecol 36, 1039–1054 (2021). https://doi.org/10.1007/s10980-021-01217-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-021-01217-7