Abstract
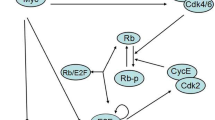
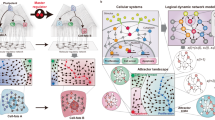
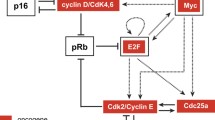
E2F/Myc genetic circuit carries out its biological functions expressed by phenotypic diversity to regulate cancer development; however, the cost involved in fluctuating environments is unclear. Here, considering the regulation of miR-17-92 in an E2F/Myc genetic circuit, we propose a toy model with coloured noise to focus on the effects of the noise/correlation strength (NS/CS) and the auto/cross-correlation time (AT/CT) on cell phenotype transition and energy cost. The results indicate that increasing AT/CT always slows down the bimodal regulation of NS/CS while extending AT of multiplicative noise amplifies this effect to shift the bimodal regime; the changing trend of the mean first passage time (MFPT) also confirms the regulatory function of noises, i.e., CT attenuates cancer spread induced by increasing CS. Moreover, by reconstructing the effective topology network, we can validate that there is an optimal switching path existed by regulating NS/CS and AT/CT according to the principle of minimum energy consumption, which is nearly independent of CT in the phase plane of CS to CT. The overall analysis indicates that E2F/Myc genetic circuit would regulate NS/CS and AT/CT of noises to achieve phenotype diversity.











Similar content being viewed by others
Data Availability
Data sharing does not apply to this article as no new data were created or analyzed in this study.
References
Balázsi, G., Oudenaarden, A., Collins, J.J.: Cellular decision making and biological noise: from microbes to mammals. Cell 144, 910–925 (2011)
Arias, A.M., Brickman, J.M.: Gene expression heterogeneities in embryonic stem cell populations: origin and function. Curr. Opin. Cell Biol. 23, 650–656 (2011)
Garcia-Ojalvo, J., Arias, A.M.: Towards a statistical mechanics of cell fate decisions. Curr. Opin. Genet. Dev. 22, 619–626 (2012)
Bose, T., Trimper, S.: A stochastic model for tumour growth with immunization. Phys. Rev. E 79, 051903 (2009)
Elowitz, M.B.: Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Sanchez, A., Golding, I.: Genetic determinants and cellular constraints in noisy gene expression. Science 342, 1188–1193 (2013)
Sui, H., Eichler, G., Bar-Yam, Y., Ingber, D.E.: Cell fates as high-dimensional attractor states of a complex gene regulatory network. Phys. Rev. Lett. 94, 128701 (2005)
Li, Q., Wennborg, A., Erik, A.: Dynamics inside the cancer cell attractor reveal cell heterogeneity, limits of stability, and escape. Proc. Natl. Acad. Sci. USA 113, 2672–2677 (2016)
Aguda, B.D., Kim, Y., Piper-Hunter, M.G., Friedman, A., Marsh, C.B.: MicroRNA regulation of a cancer network: consequences of the feedback loops involving miR-17-92, E2F, and Myc. Proc. Natl. Acad. Sci. USA 105, 19678–19683 (2008)
Grillari, J., Hackl, M., Grillari-Voglauer, R.: miR-17-92 cluster: ups and downs in cancer and aging. Biogerontology 11, 501–506 (2010)
Li, Y., Li, Y., Zhang, H., Chen, Y.: MicroRNA-mediated positive feedback loop and optimized bistable switch in a cancer network Involving miR-17-92. PLoS ONE 6, e26302 (2011)
Jia, C., Wang, L.Y., Yin, G.G., Zhang, M.Q.: Single-cell stochastic gene expression kinetics with coupled positive-plus-negative feedback. Phys. Rev. E 100, 52406 (2019)
Karpova, T.S., Kim, M.J., Spriet, C., Nalley, K., Stasevich, T.J., Kherrouche, Z., McNally, J.G.: Concurrent fast and slow cycling of a transcriptional activator at an endogenous promoter. Science 319, 466–469 (2008)
Pedraza, J.M., Paulsson, J.: Effects of molecular memory and bursting on fluctuations in gene expression. Science 319, 339–343 (2008)
Gutschner, T., Hammerle, M., Eissmann, M., Hung, G., Revenko, A., Stentrup, M., Gross, M., Zornig, M.: The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 73, 1180–1189 (2013)
Hanggi, P., Mroczkowski, T.J., Moss, F., Mcclintock, P.V.: Bistability driven by coloured noise: theory and experiment. Phys. Rev. A 32, 695–698 (1985)
Ge, H., Qian, H., Xie, X.S.: Stochastic phenotype transition of a single cell in an intermediate region of gene state switching. Phys. Rev. Lett. 114, 78101–78101 (2015)
Berry, S., Dean, C., Howard, M.: Slow chromatin dynamics allow polycomb target genes to filter fluctuations in transcription factor activity. Cell Syst. 4, 445–457 (2017)
Li, C., Wang, E., Wang, J.: Landscape and flux decomposition for exploring global natures of non-equilibrium dynamical systems under intrinsic statistical fluctuations. Chem. Phys. Lett. 505, 75–80 (2011)
Li, C., Wang, J.: Landscape and flux reveal a new global view and physical quantification of mammalian cell cycle. Proc. Natl. Acad. Sci. USA 111, 14130–14135 (2014)
Qian, H.: The mathematical theory of molecular motor movement and chemomechanical energy transduction. J. Math. Chem. 27, 219–234 (2000)
Wang, J., Xu, L., Wang, E.: Potential landscape and flux framework of nonequilibrium networks: robustness, dissipation, and coherence of biochemical oscillations. Proc. Natl. Acad. Sci. USA 105, 12271–12276 (2008)
Be Rut, A., Arakelyan, A., Petrosyan, A., CiliBeRto, S., Dillenschneider, R., Lutz, E.: Experimental verification of Landauer’s principle linking information and thermodynamics. Nature 483, 187–189 (2012)
Ge, H., Qian, H.: Thermodynamic limit of a nonequilibrium steady-state: maxwell-type construction for a bistable biochemical system. Phys. Rev. Lett. 103, 148103–148103 (2009)
Lebowitz, J.L., Spohn, H.: A gallavotti–cohen-type symmetry in the large deviation functional for stochastic dynamics. J. Stat. Phys. 95, 333–365 (1999)
Mehta, P., David, J.S.: Energetic costs of cellular computation. Proc. Natl. Acad. Sci. USA 109, 17978–17982 (2012)
Vainstein, M.H., Rub, J.M.: Gaussian noise and time-reversal symmetry in nonequilibrium Langevin models. Phys. Rev. E 75, 31106–31106 (2007)
Ao, P.: Department, emerging of stochastic dynamical equalities and steady state thermodynamics from darwinian dynamics. Commun. Theor. Phys. 5, 5–22 (2008)
Jin, S., MacLean, A.L., Peng, T., Nie, Q.: scEpath: energy landscape-based inference of transition probabilities and cellular trajectories from single-cell transcriptomic data. Bioinformatics 15, 2077–2086 (2018)
Huang, L., Yuan, Z., Yu, J., Zhou, T.: Fundamental principles of energy consumption for gene expression. Chaos 25, 123101 (2015)
Campisi, J.: Cancer and ageing: rival demons? Nat. Rev. Cancer 3, 339–349 (2003)
Raser, J.M., O’Shea, E.K.: Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Elf, J.: Fast evaluation of fluctuations in biochemical networks with the linear noise approximation. Genome Res. 13, 2475–2484 (2003)
Friedman, N., Cai, L., Xie, X.S.: Linking stochastic dynamics to population distribution: an analytical framework of gene expression. Phys. Rev. Lett. 97, 168302 (2006)
Qian, H., Shi, P.Z., Xing, J.: Stochastic bifurcation, slow fluctuations, and bistability as an origin of biochemical complexity. Phys. Chem. Chem. Phys. 11, 4861–4870 (2009)
Thomas, P., Popovi, N., Grima, R.: Phenotypic switching in gene regulatory networks. Proc. Natl. Acad. Sci. USA 111, 6994–6999 (2014)
Bennett, M.R., Volfson, D., Tsimring, L., Hasty, J.: Transient dynamics of genetic regulatory networks. Biophys. J. 92, 3501–3512 (2007)
Rodrigo, G., Poyatos, J.F.: Genetic redundancies enhance information transfer in noisy regulatory circuits. PLoS Comput. Biol. 12, e1005156 (2016)
Sneppen, K., Ringrose, L.: Theoretical analysis of polycomb-trithorax systems predicts that poised chromatin is bistable and not bivalent. Nat. Commun. 10, 2133 (2019)
Norman, T.M., Lord, N.D., Paulsson, J., Losick, R.: Memory and modularity in cell-fate decision making. Nature 503, 481–486 (2013)
Zhang, J., Zhou, T.: Markovian approaches to modeling intracellular reaction processes with molecular memory. Proc. Natl. Acad. USA 116, 201913926 (2019)
Hermsen, R., Erickson, D.W., Hwa, T.: Speed, sensitivity, and bistability in auto-activating signaling circuits. PLoS Comput. Biol. 7, e1002265 (2011)
Wolf, L., Silander, O.K., Nimwegen, E.V.: Expression noise facilitates the evolution of gene regulation. Elife (2015). https://doi.org/10.7554/eLife.05856
Zambrano, S., Bianchi, M.E., Agresti, A., Molina, N.: Interplay between stochasticity and negative feedback leads to pulsed dynamics and distinct gene activity patterns. Phys. Rev. E 92, 022711 (2015)
Ronald, F., Fox: Numerical simulations of stochastic differential equations. J. Stat. Phys. 54, 1353–1366 (1989)
Frank, T.D.: Delay Fokker–Planck equations, Novikov’s theorem, and Boltzmann distributions as small delay approximations. Phys. Rev. E 72, 11112 (2005)
Zhu, P.: Associated relaxation time and intensity correlation function of a bistable system driven by cross-correlation additive and multiplicative coloured noise sources. Eur. Phys. J. B 55, 447–452 (2007)
Tang, Y., Yuan, R., Wang, G., Zhu, X., Ao, P.: Potential landscape of high dimensional nonlinear stochastic dynamics with large noise. Sci. Rep. 7, 15762 (2016)
Houchmandzadeh, B., Vallade, M.: Exact results for a noise-induced bistable system. Phys. Rev. E 91, 022115 (2015)
Fang, X., Liu, Q., Bohrer, C., Hensel, Z., Han, W., Wang, J., Xiao, J.: Cell fate potentials and switching kinetics uncovered in a classic bistable genetic switch. Nat. Commun. 9, 2787 (2018)
Jafarpour, F., Biancalani, T., Goldenfeld, N.: Noise-induced mechanism for biological homochirality of early life self-replicators. Phys. Rev. Lett. 115, 158101 (2015)
Lenstra, T.L., Rodriguez, J., Chen, H., Larson, D.R.: Transcription dynamics in living cells. Annu. Rev. Biophys. 45, 25 (2016)
Jin, Y., Wei, X.: Mean first-passage time of a bistable kinetic model driven by two different kinds of coloured noises. Chaos Soliton Fract. 23, 275–280 (2005)
Hanggi, P., Bartussek, R., Talkner, P., Uczka, J.: Noise-induced transport in symmetric periodic potentials: White shot noise versus deterministic noise. EPL 35, 315–317 (1996)
Ronald, F., Fox: Laser-noise analysis by first-passage-time techniques. Phys. Rev. A 34, 3405 (1986)
Zhang, X.J., Qian, H., Qian, M.: Stochastic theory of nonequilibrium steady states and its applications. Part I. Phys. Rep. 510, 1–86 (2012)
Shahrezaei, V., Swain, P.S.: Analytical distributions for stochastic gene expression. Proc. Natl. Acad. Sci. USA 105, 17256–17261 (2008)
Kumar, N., Platini, T., Kulkarni, R.V.: Exact distributions for stochastic gene expression models with bursting and feedback. Phys. Rev. Lett. 113, 268105–268105 (2014)
Cao, Z., Grima, R.: Linear mapping approximation of gene regulatory networks with stochastic dynamics. Nat. Commun. 9, 3305–3305 (2018)
Xu, C.: Phenomenological bifurcation in a stochastic logistic model with correlated coloured noises. Appl. Math. Lett. 101, 106064–106064 (2019)
Cole, J.A., Luthey-Schulten, Z.: Careful accounting of extrinsic noise in protein expression reveals correlations among its sources. Phys. Rev. E 95, 062418 (2017)
Singh, A., Bokes, P.: Consequences of mRNA transport on stochastic variability in protein levels. Biophys. J. 103, 1087–1096 (2012)
Baudrimont, A., Jaquet, V., Wallerich, S., Voegeli, S., Becskei, A.: Contribution of RNA degradation to intrinsic and extrinsic noise in gene expression. Cell Rep. 26, 3752–3761 (2019)
Acknowledgements
The author would like to thank three anonymous reviewers for valuable comments, and also thank Elsevier Language Editing again (https://webshop.elsevier.com/) for English language editing.
Funding
The work was supported by the Hainan Province Science and Technology Special Fund (Grant No. ZDYF2021SHFZ231), the National Natural Science Foundation of China (Grant Nos. 12261028, 11961018, 11761025), Natural Science Foundation of Hainan Province (Grant Nos. 120RC451, 2019RC168), Hainan Province Innovative Scientific Research Project for Graduate Students (Grant Nos. Qhys2021-208, Qhys2022-182, Qhys2022-183), the financial support from Academician Shi Jianming Station of Hainan Province.
Author information
Authors and Affiliations
Contributions
HHW and LLC proposed and designed this study, did all numerical simulations described in the paper, interpreted the results, and wrote the paper. ZGW, YW and HHW performed an analytical treatment of a stochastic differential equation. Both authors contributed to the discussions.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that there is no conflict of interest.
Additional information
Communicated by Lei-Han Tang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, L., Wang, Y., Wang, Z. et al. Coloured Noises Induced Regime Shift Yet Energy-Consuming in an E2F/Myc Genetic Circuit Involving miR-17-92. J Stat Phys 190, 84 (2023). https://doi.org/10.1007/s10955-023-03095-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10955-023-03095-6