Abstract
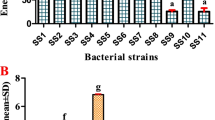
During the present research, 11 gut bacteria were isolated from the freshwater fish, Systomus sarana (General name: olive barb) and upon screening, the strains produced extracellular pectinase enzyme. Among them, the SS6 strain was found to produce a high quantity of 208.731 U/ml pectinase and through molecular characterization the SS6 strain was identified as Aeromonas guangheii. During the culture of SS6 strain, a set of parameters were optimized through the response surface methodology with a Box–Behnken design, for the production of the enzyme. The optimal conditions were found to be 2.11% of maltose, 2.20% of yeast extract, 6.5 of pH, and a temperature of 27.3 °C at 32-h incubation. Under the above conditions, the activity of pectinase production was enhanced to 371 U/ml. The purified pectinase’s molecular weight was determined to be ~ 50 kDa (by 10% 2-D PAGE). Totally, nine peptides were identified from the purified pectinase enzyme through the MALDI-TOF-MS analysis and MASCOT tool was used to get the mass spectrum of the peak at 2211 of peptide that indicated the reference pectinase protein. The referenced gene primer (pectate lyases) was PCR amplified and its nucleotide sequence was analyzed. The exo-pelA gene was cloned in pREST vector, which was found to be over expressed in Escherichia coli BL21. The ORF encoded for a mature protein comprising of 425 amino acids (1236 nucleotides) with a predicted molecular weight of ~ 48.7 kDa. The present findings underline the potential of the fish-gut microbes as a source of biotechnologically important enzymes.









Similar content being viewed by others
References
Dalvi P, Anthappan P (2007) Amylase and pectinase from single source for simultaneous desizing and scouring. Ind J Fib Text Res 32:459–465
FAO (2014) The State of World Fisheries and Aquaculture 2014. FAO, Rome. www.fao.org/publications
Hoondal G, Tiwari R, Tewari R, Dahiya N, Be Q (2002) Microbial alkaline pectinases and their industrial applications: a review. Appl Microbiol Biotechnol 59:409–418
Sasmal M, Ray RR (2015) Production of extracellular enzymes by the gut and gill microflora of Tilapia fish (Oreochromis niloticus). Asian J Multidiscip Stud 3:44–49
Mondal S, Roy T, Ray AK (2010) Characterization and identification of enzyme-producing bacteria isolated from the digestive tract of bata, Labeo bata. J World Aqua Soci 41:369–377
Debashish G, Malay S, Barindra S, Joydeep M (2005) Marine enzymes. In: Ulber R, Le Gal Y (eds) Marine biotechnology I. Advances in biochemical engineering/biotechnology. Springer, Berlin, pp 189–218
Klug-Santner BG, Schnitzhofer W, Vršanská M, Weber J, Agrawal PB, Nierstrasz VA, Guebitz GM (2006) Purification and characterization of a new bioscouring pectate lyase from Bacillus pumilus BK2. J Biotechnol 121:390–401
Mohnen D (2008) Pectin structure and biosynthesis. Cur Opin Plant Biol 11:266–277
Agüero-Chapin G, Varona-Santos J, de la Riva GA, Antunes A, González-Villa T, Uriarte E, González-Díaz H (2009) Alignment-free prediction of polygalacturonases with pseudofolding topological indices: experimental isolation from Coffea arabica and prediction of a new sequence. J Proteome Res 8:2122–2128
Khan M, Nakkeeran E, Umesh-Kumar S (2013) Potential application of pectinase in developing functional foods. Annu Rev Food Sci Technol 4:21–34
Morris GA, Kök SM, Harding SE, Adams GG (2010) Polysaccharide drug delivery systems based on pectin and chitosan. Biotechnol Gene Eng Rev 27:257–284
Sharma N, Rathore M, Sharma M (2013) Microbial pectinase: sources, characterization and applications. Rev Environ Sci Bio/Technol 12:45–60
Shevchik VE, Robert-Baudouy JANINE, Hugouvieux-Cotte-Pattat NICOLE (1997) Pectate lyase PelI of Erwinia chrysanthemi 3937 belongs to a new family. J Bacteriol 179(23):7321–7330
Hugouvieux-Cotte-Pattat N, Condemine G, Nasser W, Reverchon S (1996) Regulation of pectinolysis in Erwinia chrysanthemi. Annu Rev Microbiol 50:213–257
Jayaram KC (2010) The freshwater fishes of the Indian region, 2nd edn. Narendra Publishing House, Delhi
Hankin L, Zucker M, Sands DC (1971) Improved solid medium for the detection and enumeration of pectolytic bacteria. Appl Microbiol 22:205–209
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analyt Chem 31:426–428
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Ramesh S, Muthuvelayudham R, Rajesh Kannan R, Viruthagiri T (2013) Enhanced production of xylitol from corncob by Pachysolen tannophilus using response surface methodology. Int J Food Sci 2013:514676
Plackett RL, Burman JP (1946) The design of optimum multifactorial experiments. Biometrika 33:305–325
Kumar S, Sharma HK, Sarkar BC (2011) Effect of substrate and fermentation conditions on pectinase and cellulase production by Aspergillus niger NCIM 548 in submerged (SmF) and solid-state fermentation (SSF). Food Sci Biotechnol 20:1289
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Cakmak G, Togan I, Severcan F (2006) 17β-Estradiol induced compositionalstructural and functional changes in rainbow trout liver, revealed by FT-IR spectroscopy: a comparative study with nonylphenol. Aquat Toxic 77:53–63
Patterson SD, Aebersold R (1995) Mass spectrometric approaches for the identification of gel-separated proteins. Electrophoresis 16:1791–1814
Bordoli L, Kiefer F, Arnold K, Benkert P, Battey J, Schwede T (2009) Protein structure homology modeling using SWISS-MODEL workspace. Nat Protoc 4:1–13
Mountfort DO, Grant WD, Morgan H, Rainey FA, Stackebrandt E (1993) Isolation and characterization of an obligately anaerobic, pectinolytic, member of the genus Eubacterium from mullet gut. Arch Microbiol 159(3):289–295
Hossain TJ, Chowdhury SI, Mozumder HA, Chowdhury MN, Ali F, Rahman N, Dey S (2020) Hydrolytic exoenzymes produced by bacteria isolated and identified from the gastrointestinal tract of Bombay Duck. Front Microbiol 11:2097
Ghosh K, Sen SK, Ray AK (2002) Characterization of Bacilli isolated from the gut of rohu, Labeo rohita, fingerlings and its significance in digestion. J Appl Aquac 12:33–42
Bhowmick UD, Bhattacharjee S (2018) Bacteriological, clinical and virulence aspects of Aeromonas-associated diseases in humans. Polish J Microbiol 67:137–149
Job J, Sukumaran RK, Jayachandran K (2010) Production of a highly glucose tolerant β-glucosidase by Paecilomyces variotii MG3: optimization of fermentation conditions using Plackett–Burman and Box–Behnken experimental designs. World J Microbiol Biotechnol 26:1385–1391
Hu JN, Zhu XM, Lee KT, Zheng YN, Li W, Han LK, Sung CK (2018) Optimization of ginsenosides hydrolyzing β-glucosidase production from Aspergillus niger using response surface methodology. Biol Pharm Bull 31:1870–1874
Doğan N (2008) Production of pectinase enzyme from Aspergillus sojae batch and fed-batch systems. Master’s thesis, İzmir Institute of Technology, pp 1–84
Silva D, Martins ES, Leite RSR, Da Silva R (2007) Purification and characterization of an exo-polygalacturonase produced by Penicillium viridicatum RFC3 in solid-state fermentation. Proc Biochem 42:1237–1243
Soares MMCN, Da Silva R, Carmona EC, Gomes E (2001) Pectinolytic enzyme production by Bacillus species and their potential application on juice extraction. World J Microbiol Biotechnol 17:79–82
Van Rijsse M, Gerwig GJ, Hansen TA (1993) Isolation and characterization of an extracellular glycosylated protein complex from Clostridium thermosaccharolyticum with pectin methylesterase and polygalacturonate hydrolase activity. Appl Environ Microbiol 59:828–836
Nagel CW, Hasegawa S (1968) The isolation and characterization of a, 4 (4,5-dehydrogalacturonosyl) galacturonate hydrolase. J Food Sci 33:78–382
Kaur SJ, Gupta VK (2017) Production of pectinolytic enzymes pectinase and pectin lyase by Bacillus subtilis SAV-21 in solid state fermentation. Ann Microbiol 67:333–342
Rehman HU, Qader SAU, Aman A (2012) Polygalacturonase: production of pectin depolymerising enzyme from Bacillus licheniformis KIBGE IB-21. Carbohydr Polym 90:387–391
Rehman HU, Siddique NN, Aman A, Nawaz MA, Baloch AH, Qader SAU (2015) Morphological and molecular based identification of pectinase producing Bacillus licheniformis from rotten vegetable. J Gene Eng Biotechnol 13:139–144
Kant S, Vohra A, Gupta R (2013) Purification and physicochemical properties of polygalacturonase from Aspergillus niger MTCC 3323. Protein Expr Purif 87:11–16
Arijit D, Sourav B, Naimisha R, Rajan SS (2013) Improved production and purification of pectinase from Streptomyces sp. GHBA10 isolated from Valapattanam mangrove habitat, Kerala, India. Int Res J Biol Sci 2:16–22
Giacobbe S, Pepe O, Ventorino V, Birolo L, Vinciguerra R, Faraco V (2014) Identification and characterisation of a pectinolytic enzyme from Paenibacillus xylanolyticus. Bio Resour 9:4873–4887
Bhardwaj V, Garg N (2014) Production, purification of pectinase from Bacillus sp. MBRL576 isolate and its application in extraction of juice. Int J Sci Res 3:648–652
Zhou M, Wu J, Wang T, Gao L, Yin H, Lü X (2017) The purification and characterization of a novel alkali-stable pectate lyase produced by Bacillus subtilis PB1. World J Microbiol Biotechnol 33:190
Saladino AC, Xu Y, Tang P (2005) Homology modeling and molecular dynamics simulations of transmembrane domain structure of human neuronal nicotinic acetylcholine receptor. Biophys J 88:1009–1017
Gandreddi VS, Kappala VR, Zaveri K, Patnala K (2015) Evaluating the role of a trypsin inhibitor from soap nut (Sapindus trifoliatus L. Var. Emarginatus) seeds against larval gut proteases, its purification and characterization. BMC Biochem 16:1–16
Henrissat B, Heffron SE, Yoder MD, Lietzke SE, Jurnak F (1995) Functional implications of structure-based sequence alignment of proteins in the extracellular pectate lyase superfamily. Plant Physiol 107:963–976
Wang H, Li X, Ma Y, Song J (2014) Characterization and high-level expression of a metagenome-derived alkaline pectate lyase in recombinant Escherichia coli. Proc Biochem 49:69–76
Berensmeier S, Singh SA, Meens J, Buchholz K (2004) Cloning of the pelA gene from Bacillus licheniformis 14A and biochemical characterization of recombinant, thermostable, high-alkaline pectate lyase. Appl Microbiol Biotechnol 64(4):560–567
Soriano M, Blanco A, Díaz P, Pastor FJ (2000) An unusual pectate lyase from a Bacillus sp. with high activity on pectin: cloning and characterization. Microbiology 146:189–195
Soriano M, Diaz P, Pastor FIJ (2006) Pectate lyase C from Bacillus subtilis: a novel endo-cleaving enzyme with activity on highly methylated pectin. Microbiology 152:617–625
Jung YJ, Lee YS, Park IH, Chandra MS, Kim KK, Choi YL (2010) Molecular cloning, purification and characterization of thermostable-1, 3–1,4 glucanase from Bacillus subtilis A8–8. Ind J Biochem Biophys 47:203–210
Pandey S, Kushwah J, Tiwari R, Kumar R, Somvanshi VS, Nain L, Saxena AK (2014) Cloning and expression of β-1,4-endoglucanase gene from Bacillus subtilis isolated from soil long term irrigated with effluents of paper and pulp mill. Microbiol Res 169:693–698
Ibrahim E, Jones K, Hosseny EN (2015) Molecular cloning and expression of cellulase and polygalacturonase genes in E. coli as a promising application for biofuel production. J Petrol Environ Biotechnol 4:1–10
Bekri MA, Desair J, Keijers V, Proost P, Searle-van Leeuwen M, Vanderleyden J, Vande Broek A (1999) Azospirillum irakense produces a novel type of pectate lyase. J Bacteriol 181:2440–2447
Acknowledgements
The authors are grateful to the authorities of the Periyar University for providing the necessary facilities to carry out this research work. Thanks are due to Dr. R. Rajaram, Associate Professor, Dept. of Marine Science, Bharathidasan University, for his help in plagiarism check. PP thanks the UGC, New Delhi, for the award of BSR Faculty Fellowship (Ref. No.F.18-1/2011 (BSR), 26.06.2018).
Funding
This research work was done without funding.
Author information
Authors and Affiliations
Contributions
All authors brainstormed and came up with this study concept. NT provided the chemicals, equipment for the related microbiology tests and Response Surface Methodology (RSM-software) and related guidelines. BV provided the cloning vector, expression strain and related guidelines. PA assisted in review and editing for the manuscript. SD assisted in writing—review and editing, data curation of the manuscript. PP provided the full manuscript language correction, conceptualization, writing—review and editing—original, draft. AD provided the bacteria and carried out all the experiments, compiled and analyzed the resultant data and compiled this paper with insights from all the other authors.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical Approval
According to the Committee for the Purpose of Control and Supervision of Experiments on Animals-CPCSEA (under Ministry of Environment, Forests and Climate Change, Government of India), the edible fish which are used for laboratory experiments are exempted from obtaining Institutional Animal Ethics Committee (IAEC) approval.
Consent to Participate
The study animal that we have utilized for the present work is Systomus sarana (Hamilton, 1882) which is an edible fish, and hence the necessity did not arise to acquire the approval from IAEC. However, we have followed the OECD Guidelines for safe handling of experimental animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dhayalan, A., Thillainathan, N., Velramar, B. et al. Pectinase from a Fish Gut Bacterium, Aeromonas guangheii (SS6): Production, Cloning and Characterization. Protein J 41, 572–590 (2022). https://doi.org/10.1007/s10930-022-10077-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-022-10077-2