Abstract
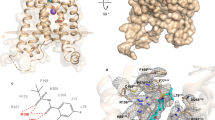
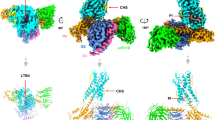
Leukotriene B4 (LTB4) exerts its biological effects through stimulation of specific G protein-coupled receptors (GPCRs)—namely BLT1 and BLT2. Due to the absence of human BLT1 and BLT2 crystal structures, the current study was set to predict the 3D structures of these two receptors for structure-based anti-inflammatory drug discovery. Homology modeling of the BLT1 receptor was first constructed, based on various X-ray and NMR GPCR templates, followed by molecular dynamics (MD) refinement. Using a single-template approach, nine well-established alignment methods and ten secondary structure prediction methods during the backbone generation were implemented and assessed. The binding sites of the BLT1 receptor were then mapped using fifteen chemical probes with the help of FTMAP and AutoDock Vina 4.2 software. Model validation was performed through the docking of eight specific antagonists that have experimental inhibition constants (ki) towards BLT1. The antagonists-BLT1 docked structures were then subjected to AMBER-based molecular mechanical minimization and the corresponding binding energies were calculated using molecular mechanics–generalized Born surface area (MM/GBSA) approach. According to the results, the most energetically stable models were constructed using SAlign method for the alignment process and PSIPRED for secondary structure prediction. In comparison, the refined BLT1 model built on 2KS9 as an NMR template has the lowest DOPE energy compared to those built on 4EA3 and 4XT1 as X-ray templates. According to the mapping results, two main binding sites were identified: one was among TMs II, III and VII and the other was among TMs III, IV and V. For the antagonists, correlation between binding energies and experimental data was in a good agreement, with a correlation coefficient (R2 value) of 0.91. Due to the great amino acid sequence similarity between BLT1 and BLT2 receptors (calculated as 45.2%), BLT2 model was constructed based on the predicted BLT1 model.









Similar content being viewed by others
References
Michino M, Abola E, Participant GPCRD 2008, Brooks CL, Dixon JS, Moult J, Stevens RC (2009) Community-wide assessment of GPCR structure modelling and ligand docking: GPCR Dock 2008. Nat Rev Drug Discov 8:455–463
Vyas VK, Ukawala RD, Ghate M, Chintha C (2012) Homology modeling a fast tool for drug discovery: current perspectives. Indian J Pharm Sci 74:1–17
Latek D, Pasznik P, Carlomagno T, Filipek S (2013) Towards improved quality of GPCR models by usage of multiple templates and profile-profile comparison. PLoS ONE 8:e56742–e56751
Yarnitzky T, Levit A, Niv MY (2010) Homology modeling of G-protein-coupled receptors with X-ray structures on the rise. Curr Opin Drug Discov Dev 13:317–325
Cavasotto CN, Phatak SS (2009) Homology modeling in drug discovery: current trends and applications. Drug Discov Today 14:676–683
Kufareva I, Rueda M, Katritch V, Stevens RC, Abagyan R (2011) Status of GPCR modeling and docking as reflected by community-wide GPCR dock 2010 assessment. Structure 19:1108–1126
Tager AM, Luster AD (2003) BLT1 and BLT2: the leukotriene B4 receptors. Prostaglandins Leukot Essent Fat Acids 69:123–134
Liu M, Yokomizo T (2015) The role of leukotrienes in allergic diseases. Allergol Int 64:17–26
Mathis SP, Jala VR, Lee DM, Haribabu B (2010) Nonredundant roles for leukotriene receptors BLT1 and BLT2 in inflammatory arthritis. J Immunol 185:3049–3056
Hori T, Okuno T, Hirata K, Yamashita K, Kawano Y, Yamamoto M, Hato M, Nakamura M, Shimizu T, Yokomizo T, Miyano M, Yokoyama S (2018) Na+-mimicking ligands stabilize the inactive state of leukotriene B4 receptor BLT1. Nat Chem Biol 14:262–269
Okuno T, Ishitani T, Yokomizo T (2015) Biochemical characterization of three BLT receptors in Zebrafish. PLoS ONE 10:1–19
Okuno T, Yokomizo T, Hori T, Miyano M, Shimizu T (2005) Leukotriene B4 receptor and the function of its helix 8. J Biol Chem 280:32049–32052
Lavigne R-P, Stanková J, Chen MZ, Gouill CL, Gaudreau PR, Beaulieu M-E (2004) Structural determinants regulating expression of the high affinity leukotriene B4 receptor. J Biol Chem 279:10338–10345
Martin A, Damian M, Laguerre M, Parello J, Pucci B, Serre L, Mary S, Marie J, Banères J-L (2009) Engineering a G protein-coupled receptor for structural studies: stabilization of the BLT1 receptor ground state. Protein Sci 18:727–734
Webb B, Sali A (2014) Comparative protein structure modeling using MODELLER. Curr Protoc Bioinform 54:5.6.1–5.6.37
Martí-Renom MA, Stuart AC, Fiser A, Sánchez R, Melo F, Šali A (2000) Comparative protein structure modeling of genes and genomes. Annu Rev Biophys Biomol Struct 29:291–325
Katoh K, Misawa K, Kuma K-I, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Rice P, Longden I, Bleasby A (2000) EMBOSS: the european molecular biology open software suite. Trends Genet 16:276–277
Chang J-M, Di Tommaso P, Taly J-F, Notredame C (2012) Accurate multiple sequence alignment of transmembrane proteins with PSI-Coffee. BMC Bioinform 13:1–7
Floden EW, Tommaso PD, Chatzou M, Magis C, Notredame C, Chang J-M (2016) PSI/TM-Coffee: a web server for fast and accurate multiple sequence alignments of regular and transmembrane proteins using homology extension on reduced databases. Nucleic Acids Res 44:W339–W343
Braberg H, Webb BM, Tjioe E, Pieper U, Sali A, Madhusudhan MS (2012) SALIGN: a web server for alignment of multiple protein sequences and structures. Bioinformatics 28:2072–2073
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539–539
Simossis VA, Heringa J (2005) PRALINE: a multiple sequence alignment toolbox that integrates homology-extended and secondary structure information. Nucleic Acids Res 33:W289–W294
Joyeux L, Penczek PA (2002) Efficiency of 2D alignment methods. Ultramicroscopy 92:33–46
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Sen TZ, Jernigan RL, Garnier J, Kloczkowski A (2005) GOR V server for protein secondary structure prediction. Bioinformatics 21:2787–2788
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16:404–405
Tusnády GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics 17:849–850
Käll L, Krogh A, Sonnhammer ELL (2004) A combined transmembrane topology and signal peptide prediction method. J Mol Biol 338:1027–1036
Pollastri G, McLysaght A (2005) Porter: a new, accurate server for protein secondary structure prediction. Bioinformatics 21:1719–1720
Hofmann K, WS (1993) TMBASE—a database of membrane spanning protein segments. Biol Chem Hoppe-Seyler 374:166
Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Lin K, Simossis VA, Taylor WR, Heringa J (2005) A simple and fast secondary structure prediction method using hidden neural networks. Bioinformatics 21:152–159
Rost B, Yachdav G, Liu J (2004) The predictprotein server. Nucleic Acids Res 32:W321–W326
Bernhofer M, Kloppmann E, Reeb J, Rost B (2016) TMSEG: novel prediction of transmembrane helices. Proteins Struct Funct Bioinform 84:1706–1716
Miao Z, Cao Y (2016) Quantifying side-chain conformational variations in protein structure. Sci Rep 6:1–10
Miao Z, Cao Y, Jiang T (2011) RASP: rapid modeling of protein side chain conformations. Bioinformatics 27:3117–3122
Wang Q, Canutescu AA, Dunbrack RL (2008) SCWRL and MolIDE: computer programs for side-chain conformation prediction and homology modeling. Nat Protoc 3:1832–1847
Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham TE III, Darden TA, Duke RE, Gohlke H, Goetz AW, Gusarov S, Homeyer N, Janowski P, Kaus J, Kolossváry I, Kovalenko A, Lee TS, LeGrand S, Luchko T, Luo R, Madej B, Merz KM, Paesani F, Roe DR, Roitberg A, Sagui C, Salomon-Ferrer R, Seabra G, Simmerling CL, Smith W, Swails J, Walker RC, Wang J, Wolf RM, Wu X, Kollman PA (2014) Amber 14. University of California, San Francisco
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C (2015) ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J Chem Theory Comput 11:3696–3713
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Seeliger D, de Groot BL (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput-Aided Mol Des 24:417–422
Ngan CH, Bohnuud T, Mottarella SE, Beglov D, Villar EA, Hall DR, Kozakov D, Vajda S (2012) FTMAP: extended protein mapping with user-selected probe molecules. Nucleic Acids Res 40:W271–W275
Kozakov D, Grove LE, Hall DR, Bohnuud T, Mottarella SE, Luo L, Xia B, Beglov D, Vajda S (2015) The FTMap family of web servers for determining and characterizing ligand-binding hot spots of proteins. Nat Protoc 10:733–755
Forli S, Huey R, Pique ME, Sanner M, Goodsell DS, Olson AJ (2016) Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat Protoc 11:905–919
SZYBKI 1.9.0.3 (2016) OpenEye Scientific Software, Santa Fe
Massova I, Kollman P (2000) Combined molecular mechanical and continuum solvent approach (MM-PBSA/GBSA) to predict ligand binding. Perspect Drug Discov Des 18:113–135
Morales JL, Nocedal J (2000) Automatic preconditioning by limited memory Quasi-Newton updating. SIAM J Optim 10:1079–1096
Im W, Feig M, Brooks CL (2003) An implicit membrane generalized Born theory for the study of structure, stability, and interactions of membrane proteins. Biophys J 85:2900–2918
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25:1157–1174
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas O, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09. Revision E01 edn. Gaussian Inc., Wallingford
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Humphrey W, Dalke A, Schulten K (1996) VMD—visual molecular dynamics. J Mol Graph 14:33–38
Costanzi S (2012) Homology modeling of class A G protein-coupled receptors. Methods Mol Biol 857:259–279
Dalton JAR, Jackson RM (2007) An evaluation of automated homology modelling methods at low target–template sequence similarity. Bioinformatics 23:1901–1908
Ji Y-Y, Li Y-Q (2010) The role of secondary structure in protein structure selection. Eur Phys J E 32:103–107
King RD, Sternberg MJ (1996) Identification and application of the concepts important for accurate and reliable protein secondary structure prediction. Protein Sci 5:2298–2310
Singh M, Sandhu PS, Kaur RK (2008) Protein secondary structure prediction. World Acad Sci Eng Technol 42:458–461
Rossi KA, Weigelt CA, Nayeem A, Krystek SR (2007) Loopholes and missing links in protein modeling. Protein Sci 16:1999–2012
Nair PC, Miners JO (2014) Molecular dynamics simulations: from structure function relationships to drug discovery. Silico Pharmacol 2:1–4
Durrant JD, McCammon JA (2011) Molecular dynamics simulations and drug discovery. BMC Biol 9:1–9
Liu G, Li Z, Chiang Y, Acton T, Montelione GT, Murray D, Szyperski T (2005) High-quality homology models derived from NMR and X-ray structures of E. coli proteins YgdK and Suf E suggest that all members of the YgdK/Suf E protein family are enhancers of cysteine desulfurases. Protein Sci 14:1597–1608
Basu S, Jala VR, Mathis S, Rajagopal ST, Del Prete A, Maturu P, Trent JO, Haribabu B (2007) Critical role for polar residues in coupling leukotriene B4 binding to signal transduction in BLT1. J Biol Chem 282:10005–10017
Marder P, Sawyer JS, Froelich LL, Mann LL, Spaethe SM (1995) Blockade of human neutrophil activation by 2-[2-propyl-3-[3-[2-ethyl-4-(4-fluorophenyl)-5-hydroxyphenoxy]propoxy]phenoxy]benzoic acid (LY293111), a novel leukotriene B4 receptor antagonist. Biochem Pharmacol 49:1683–1690
Showell HJ, Breslow R, Conklyn MJ, Hingorani GP, Koch K (1996) Characterization of the pharmacological profile of the potent LTB4 antagonist CP-105,696 on murine LTB4 receptors in vitro. Br J Pharmacol 117:1127–1132
Jackson WT, Froelich LL, Boyd RJ, Schrementi JP, Saussy DL, Schultz RM, Sawyer JS, Sofia MJ, Herron DK, Goodson T, Snyder DW, Pechous PA, Spaethe SM, Roman CR, Fleisch JH (1999) Pharmacologic actions of the second-generation leukotriene B4 receptor antagonist LY293111: in vitro studies. J Pharmacol Exp Ther 288:286–294
Hansel TT, Barnes PJ (2001) New drugs for asthma, allergy and COPD, vol 31, 1st edn. S. Karger, Basel
Daines RA, Chambers PA, Foley JJ, Griswold DE, Kingsbury WD, Martin LD, Schmidt DB, Sham KKC, Sarau HM (1996) (E)-3-[6-[[(2,6-Dichlorophenyl)thio]methyl]-3-(2-phenylethoxy)-2-pyridinyl]-2- propenoic acid: a high-affinity leukotriene B4 receptor antagonist with oral antiinflammatory activity. J Med Chem 39:3837–3841
Cohen N, Bizzarro FT, May WP, Toth K, Lee FK, Heslin PH, Holland GW, Kwoh SC, Franco LS, Simko BA, Yagaloff KA (1994) Benzenepropanoic acids containing chromanone or naphthalenone moieties are potent and orally active leukotriene B4 antagonists. Bioorg Med Chem Lett 4:2883–2888
Psychoyos S, Uziel-Fusi S, Morrissey MM, Ganu V, Smith CW (1991) Thick filter neutrophil chemotaxis performed in the absence of albumin. J Immunol Methods 137:37–46
Taylor BM, Crittenden NJ, Bruden MN, Wishka DG, Morris J, Richards IM, Sun FF (1991) Biological activity of leukotriene B4 analogs: inhibition of guinea pig eosinophil migration in vitro by the 2,6-disubstituted pyridine analogs U-75,302 and U-75,485. Prostaglandins 42:211–224
Yokomizo T, Kato K, Terawaki K, Izumi T, Shimizu T (2000) A second leukotriene B(4) receptor, BLT2: a new therapeutic target in inflammation and immunological disorders. J Exp Med 192:421–432
Funding
This work was funded by the Science and Technology Development Fund, STDF, Egypt (Grant No. 7972).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ibrahim, M.A.A., Hassan, A.M.A. Comparative Modeling and Evaluation of Leukotriene B4 Receptors for Selective Drug Discovery Towards the Treatment of Inflammatory Diseases. Protein J 37, 518–530 (2018). https://doi.org/10.1007/s10930-018-9797-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-018-9797-3