Abstract
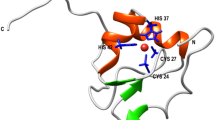
Arginine kinase (AK) is a key metabolic enzyme for keeping energy balance in invertebrates. Therefore, regulation of the enzymatic activity and the folding studies of AK from the various invertebrates have been the focus of investigation. We studied the effects of helical structures by using hexafluoroisopropanol (HFIP) on AK folding. Folding kinetic studies showed that the folding rates of the urea-denatured AKs were significantly decelerated after being induced in various concentrations of HFIP. AK lost its activity completely at concentrations greater than 60%. The results indicated that the HFIP-induced helical structures in the denatured state play a negative role in protein folding, and the helical structures induced in 5% (v/v) HFIP act as the most effective barrier against AK taking its native structure. The computational docking simulations (binding energies for −2.19 kcal/mol for AutoDock4.2 and −20.47 kcal/mol for Dock6.3) suggested that HFIP interacts with the several important residues that are predicted by both programs. The excessively pre-organized helical structures not only hampered the folding process, but also ultimately brought about changes in the three-dimensional conformation and biological function of AK.








Similar content being viewed by others
Abbreviations
- AK:
-
Arginine kinase
- HFIP:
-
1,1,1,3,3,3-Hexafluoroisopropanol
- TFE:
-
2,2,2-Trifluoroethanol
- CK:
-
Creatine kinase
- ANS:
-
8-Anilino-1-naphthalenesulfonic acid
- CD:
-
Circular dichroism
- UV:
-
Ultraviolet
- SDS:
-
Sodium dodecyl sulfate
- PAGE:
-
Polyacrylamide gel electrophoresis
References
Abkevich VI, Gutin AM, Shakhnovich EI (1995) J Mol Biol 252:460–471
Ahmad B, Haq SK, Varshney A, Moosavi-Movahedi AA, Khan RH (2010) Biochemistry (Mosc) 75:486–530
Ahmad B, Islam Z, Varshney A, Khan RH (2010) Protein Pept Lett 17:660–666
Anderson VL, Ramlall TF, Rospigliosi CC, Webb WW, Eliezer D (2010) Proc Natl Acad Sci USA 107:18850–18855
Arnold K, Bordoli L, Kopp J, Schwede T (2006) Bioinformatics 22:195–201
Baldwin RL, Rose GD (1999) Trends Biochem Sci 24:26–33
Benkert P, Künzli M, Schwede T (2009) Nucleic Acids Res 37:W510–W514
Brockwell DJ, Smith DA, Radford SE (2000) Curr Opin Struct Biol 10:16–25
Chi CN, Bach A, Engström A, Wang H, Strømgaard K, Gianni S, Jemth P (2009) Biochemistry 48:7089–7097
Dadlez M (1999) Acta Biochim Pol 46:487–508
Daggett V, Fersht A (2003) Natl Rev Mol Cell Biol 4:497–502
Glyakina AV, Galzitskaya OV (2010) Biochemistry (Mosc) 75:995–1005
Havens J, Castellani M, Kleinschroth T, Ludwig B, Durham B, Millett F (2011) Biochemistry 50:10462–10472
Huang K, Park YD, Cao ZF, Zhou HM (2001) Biochim Biophys Acta 1545:305–313
Huey R, Morris GM, Olson AJ, Goodsell DS (2007) J Comput Chem 28:1145–1152
Lewandowska A, Ołdziej S, Liwo A, Scheraga HA (2010) Biophys Chem 151:1–9
Li C, Sun S, Park D, Jeong HO, Chung HY, Liu XX, Zhou HM (2011) Int J Biol Macromol 49:910–916
Liu N, Wang JS, Wang WD, Pan JC (2011) Int J Biol Macromol 49:98–102
López-Hernández E, Cronet P, Serrano L, Muñoz V (1997) J Mol Biol 266:610–620
Lü ZR, Oh SH, Zhou SS, Zou HC, Park D, Park SJ, Zhou HW, Bhak J, Park YD, Zou F (2010) Appl Biochem Biotechnol 160:831–842
Lü ZR, Wang YJ, Lee DY, Park YD, Zou HC, Zou F (2009) J Biomol Struct Dyn 26:567–574
Miyawaki O, Tatsuno M (2011) J Biosci Bioeng 111:198–203
Mohan PM, Hosur RV (2009) J Biosci 34:465–479
Mu H, Lü ZR, Park D, Kim BC, Bhak J, Zou F, Yang JM, Li S, Park YD, Zou HC, Zhou HM (2010) Appl Biochem Biotechnol 160:1309–1320
Naseem F, Khan RH (2005) Biochim Biophys Acta 1723:192–200
Pan JC, Yu Z, Su XY, Sun YQ, Rao XM, Zhou HM (2004) Protein Sci 13:1892–1901
Radford SE (2000) Trends Biochem Sci 25:611–618
Roccatano D, Fioroni M, Zacharias M, Colombo G (2005) Protein Sci 14:2582–2589
Scharnagl C, Reif M, Friedrich J (2005) Biochim Biophys Acta 1749:187–213
Sen P, Ahmad B, Rabbani G, Khan RH (2010) Int J Biol Macromol 46:250–254
Shi L, Xia Y, Zhang M, Yin SJ, Si YX, Qian GY, Lü ZR, Zhou HM, Park D, Chung HY, Zou F, Park YD (2011) Protein Pept Lett 18:726–732
Suzuki T, Kawasaki Y, Furukohri T (1997) Biochem J 328:301–306
Teixeira J (2009) Gen Physiol Biophys 28:168–173
Uversky VN (2009) Protein J 28:305–325
Weinkam P, Zimmermann J, Romesberg FE, Wolynes PG (2010) Acc Chem Res 43:652–660
Wong KB, Clarke J, Bond CJ, Neira JL, Freund SM, Fersht AR, Daggett V (2000) J Mol Biol 296:1257–1282
Yousef MS, Fabiola F, Gattis JL, Somasundaram T, Chapman MS (2002) Acta Crystallogr D Biol Crystallogr 58:2009–2017
Yu Z, Pan J, Zhou HM (2002) Protein Pept Lett 9:545–552
Zou HC, Yu ZH, Wang YJ, Yang JM, Zhou HM, Meng FG, Park YD (2007) J Biomol Struct Dyn 24:359–368
Acknowledgments
This study was supported by the grants from the Science and Technology Bureau of Jiaxing, Zhejiang (No. 2008AY2032) and the Science and Technology Planning Project of Zhejiang Province (No. 2010C33139). Dr. Wei-Jiang Hu was supported by a grant from China Postdoctoral Science Foundation (No. 20060400467). Dr. Hae Young Chung was supported by National Research Foundation of Korea (NRF) grant funded by the Korea government (MOST) (No. 20090083538) and thanks Aging Tissue Bank for providing research information. Dr. Jun-Mo Yang was supported by the grant of the Korea Health 21 R&D Project (Ministry of Health, Welfare and Family Affairs, Republic of Korea, 01-PJ3-PG6-01GN12-0001) and by a grant (Grant No. C-A9-220-1) from Samsung Biomedical Research Institute.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Li, HL., Zhou, SM., Park, D. et al. Deceleration of Arginine Kinase Refolding by Induced Helical Structures. Protein J 31, 267–274 (2012). https://doi.org/10.1007/s10930-012-9397-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-012-9397-6