Abstract
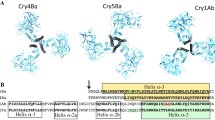
Cry5Aa is a crystal protein produced by Bacillus thuringiensis serovar. damstadiensis during its stationary phase, this δ-endotoxin is active against nematodes and has great potential for nematodes control. The theoretical model of the three-dimensional structure of Cry5Aa was predicted by homology modeling on the structures of the Cry1Aa which is specific to Lepidopteran insects. The structure of the Cry5Aa resembles previously reported Cry toxin structures but shows the following distinctions. Cry5Aa has a long insertion in α2 of domain I. Some loops in the domain II and III of Cry5Aa are exposed to the solvent. In this work we give a brief description of our model and hypothesize the residues of the Cry5Aa that could be important in receptor recognition and pore formation. This model will be helpful for the design of mutagenesis experiments aimed to the improvement of toxicity, and lead to a deep understanding of the mechanism of action of nematicidal toxins.








Similar content being viewed by others
Abbreviations
- ATP:
-
Adenosine-5′-triphosphate
- Bt:
-
Bacillus thuringiensis
- Cry:
-
Crystal
- Cyt:
-
Cytolytic
- RMSD:
-
Root mean square deviation
- PMDB:
-
Protein model data base
References
Ballester VV, Granero F, de Maagd RA, Bosch D, Mensua JL, Ferre J (1999) Appl Environ Microbiol 65:1900–1903
Bone LW, Bottjer KP, Gill SS (1987) J Parasitol 73:295–299
Boonserm P, Davis P, Ellar DJ, Li J (2005) J Mol Biol 348:363–382
Boonserm P, Mo M, Angsuthanasombat C, Lescar J (2006) J. Bacteriol 188:3391–3401
Bottjer KP, Bone LW, Gill SS (1985) Exp Parasitol 60:239–244
Burton SL, Ellar DJ, Li J, Derbyshire DJ (1999) J Mol Biol 287:1011–1022
Ciordia H, Bizzell WE (1961) J Parasitol 47:411–416
DeLano WL (2002) The PyMOL user’s manual. DeLano Scientific, San Carlos, CA
Delécluse A, Pancet S, Klier A, Rapoport G (1993) Appl Environ Microbiol 59:3922–3927
de Maagd RA, Weemen-Hendriks M, Stiekema W, Bosch D (2000) Appl Environ Microbiol 66:1559–1563
Derbyshire DJ, Ellar DJ, Li J (2001) Acta Crystallogr Sect D 57:1938–1944
Fernandez LE, Perez C, Segovia L, Rodriguez MH, Gill SS, Bravo A, Soberon M (2005) FEBS Lett 579:3508–3514
Galitsky N, Cody V, Wojtczak A, Ghosh D, Luft JR, Pangborn W, English L (2001) Acta Crystallogr Sect D 57:1101–1109
Gazit E, La Rocca P, Sansom MS, Shai Y (1998) Proc Natl Acad Sci USA 95:12289–12294
Griffitts JS, Haslam SM, Yang T, Garczynski SF, Mulloy B, Morris H, Cremer PS (2005) Science 307:922–925
Grochulski P, Masson L, Borisova S, Pusztai-Carey M, Schwartz JL, Brousseau R, Cygler M (1995) J Mol Biol 254:447–464
Guex N, Peitsch MC (1997) Electrophoresis 18:2714–2723
Gutierrez P, Alzate O, Orduz S (2001) Mem Inst Oswaldo Cruz 96:357–364
Hofmann C, Vanderbruggen H, Hofte H, Van Mellaert H (1988) Proc Natl Acad Sci USA 85:7844–7848
Hyperchem Professional 8 Evaluation (2007) Hyperchem Inc
Jurat-Fuentes JL, Adang MJ (2001) Appl Environ Microbiol 67:323–329
Knowles BH, Ellar DJ (1987) Biochem Biophys Acta 924:509–518
Kotze AC, O’Grady J, Gough JM, Pearson R, Bagnall NH, Kemp DH, Akhurst RJ (2005) Int J Parasitol 35:1013–1022
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J Appl Cryst 26:283–291
Li JD, Carroll J, Ellar DJ (1991) Nature 353:815–821
Lijnzaad P, Berendsen HJ, Argos P (1996) Proteins 25:389–397
Masson L, Tabashnik BE, Mazza A, Prefontaine G, Potvin L, Brousseau R, Schwartz JL (2002) Appl Environ Microbiol 68:194–200
Mongan NP, Baylis HA, Adcock C, Smith GR, Sansom MS, Sattelle DB (1998) Recept Channels 6:213–228
Morse RJ, Yamamoto T, Stroud RM (2001) Structure 9:409–417
Parker MW, Buckley JT, Postma JP, Tucker AD, Leonard K, Pattus F, Tsernoglou D (1994) Nature 367:292–295
Payne J, Narva KE, Fu J (1998) US patent, 5831011
Pigott CR, Ellar DJ (2007) Microbiol Mol Biol Rev 71:255–281
Roh JY, Choi JY, Li MS, Jin BR, Je YH (2007) Microbiol Biotechnol. 17:547–559
Schnepf HE, Crickmore N, van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Microbiol Mol Biol Rev 62:772–806
Schnepf HE, Schwab GE, Payne J, Narva KE, Foncerrada L (1998) US patent, 6632792
Schwartz JL, Potvin L, Chen XJ, Brousseau R, Laprade R, Dean DH (1997) Appl Environ Microbiol 63:3978–3984
Sick AJ, Schwab GE, George E, Payne JM (1994). US patent, 5281530
Tuntitippawan T, Boonserm P, Katzenmeier G, Angsuthanasombat C (2005) FEMS Microbiol Lett 242:325–332
Walters, F.S., Slatin, S.L., Kulesza, C.A., and English, L.H. (1993). 196, 921–926
Wang FX, Xia LQ, Ding XZ, Zhao XM, Shan SP, Mo XT, Zhang YM, Yu ZN (2008) Chem J Chin 29:1999–2002
Wei JZ, Hale K, Carta L, Platzer E, Wong C, Fang SC, Aroian RV (2003) Proc Natl Acad Sci 100:2760–2765
Xia LQ, Zhao XM, Ding XZ, Wang FX, Sun YJ (2008) J Mol Model 14:843–848
Zhao XM, Xia LQ, Wang FX, Ding XZ, Shan SP, Zhang YM (2008) Acta Chimi Sin 66:108–111
Acknowledgments
We thank Dr. You-min Zhang (Gene Bridge GmbH, Dresden, Germany) for a critical reading of the manuscript. This research was supported by grants from the National Natural Science Foundation of China (No. 30670052, 30570050) and 863 Program of China (2006AA02Z187, 2006AA10A212).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xin-Min, Z., Li-Qiu, X., Xue-Zhi, D. et al. The Theoretical Three-dimensional Structure of Bacillus thuringiensis Cry5Aa and Its Biological Implications. Protein J 28, 104–110 (2009). https://doi.org/10.1007/s10930-009-9169-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-009-9169-0