Abstract
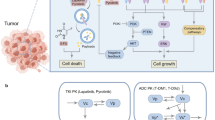
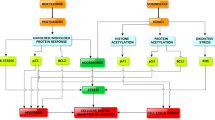
HER2-positive breast cancer (BC) is a rapidly growing and aggressive BC subtype that predominantly affects younger women. Despite improvements in patient outcomes with anti-HER2 therapy, primary and/or acquired resistance remain a major clinical challenge. Here, we sought to use a quantitative systems pharmacological (QSP) approach to evaluate the efficacy of lapatinib (LAP), abemaciclib (ABE) and 5-fluorouracil (5-FU) mono- and combination therapies in JIMT-1 cells, a HER2+ BC cell line exhibiting intrinsic resistance to trastuzumab. Concentration–response relationships and temporal profiles of cellular viability were assessed upon exposure to single agents and their combinations. To quantify the nature and intensity of drug-drug interactions, pharmacodynamic cellular response models were generated, to characterize single agent and combination time course data. Temporal changes in cell-cycle phase distributions, intracellular protein signaling, and JIMT-1 cellular viability were quantified, and a systems-based protein signaling network model was developed, integrating protein dynamics to drive the observed changes in cell viability. Global sensitivity analyses for each treatment arm were performed, to identify the most influential parameters governing cellular responses. Our QSP model was able to adequately characterize protein dynamic and cellular viability trends following single and combination drug exposure. Moreover, the model and subsequent sensitivity analyses suggest that the activation of the stress pathway, through pJNK, has the greatest impact over the observed declines of JIMT-1 cell viability in vitro. These findings suggest that dual HER2 and CDK 4/6 inhibition may be a promising novel treatment strategy for refractory HER2+ BC, however, proof-of-concept in vivo studies are needed to further evaluate the combined use of these therapies.









Similar content being viewed by others
References
Loibl S, Gianni L (2017) HER2-positive breast cancer. Lancet 389(10087):2415–2429. https://doi.org/10.1016/S0140-6736(16)32417-5
Park JW, Neve RM, Szollosi J, Benz CC (2008) Unraveling the biologic and clinical complexities of HER2. Clin Breast Cancer 8(5):392–401. https://doi.org/10.3816/CBC.2008.n.047
Arteaga CL, Sliwkowski MX, Osborne CK, Perez EA, Puglisi F, Gianni L (2011) Treatment of HER2-positive breast cancer: current status and future perspectives. Nat Rev Clin Oncol 9(1):16–32. https://doi.org/10.1038/nrclinonc.2011.177
Zaczek A, Brandt B, Bielawski KP (2005) The diverse signaling network of EGFR, HER2, HER3 and HER4 tyrosine kinase receptors and the consequences for therapeutic approaches. Histol Histopathol 20(3):1005–1015
Giordano SH, Temin S, Chandarlapaty S, Crews JR, Esteva FJ, Kirshner JJ, Krop IE, Levinson J, Lin NU, Modi S, Patt DA, Perlmutter J, Ramakrishna N, Winer EP, Davidson NE (2018) Systemic therapy for patients with advanced human epidermal growth factor receptor 2-positive breast cancer: ASCO clinical practice guideline update. J Clin Oncol 36(26):2736–2740. https://doi.org/10.1200/JCO.2018.79.2697
Giordano SH, Temin S, Kirshner JJ, Chandarlapaty S, Crews JR, Davidson NE, Esteva FJ, Gonzalez-Angulo AM, Krop I, Levinson J, Lin NU, Modi S, Patt DA, Perez EA, Perlmutter J, Ramakrishna N, Winer EP, American Society of Clinical Oncology (2014) Systemic therapy for patients with advanced human epidermal growth factor receptor 2-positive breast cancer: American Society of Clinical Oncology clinical practice guideline. J Clin Oncol 32(19):2078–2099. https://doi.org/10.1200/JCO.2013.54.0948
Koninki K, Barok M, Tanner M, Staff S, Pitkanen J, Hemmila P, Ilvesaro J, Isola J (2010) Multiple molecular mechanisms underlying trastuzumab and lapatinib resistance in JIMT-1 breast cancer cells. Cancer Lett 294(2):211–219. https://doi.org/10.1016/j.canlet.2010.02.002
Nagy P, Friedlander E, Tanner M, Kapanen AI, Carraway KL, Isola J, Jovin TM (2005) Decreased accessibility and lack of activation of ErbB2 in JIMT-1, a herceptin-resistant, MUC4-expressing breast cancer cell line. Can Res 65(2):473–482
Oliveras-Ferraros C, Vazquez-Martin A, Cufi S, Torres-Garcia VZ, Sauri-Nadal T, Barco SD, Lopez-Bonet E, Brunet J, Martin-Castillo B, Menendez JA (2011) Inhibitor of Apoptosis (IAP) survivin is indispensable for survival of HER2 gene-amplified breast cancer cells with primary resistance to HER1/2-targeted therapies. Biochem Biophys Res Commun 407(2):412–419. https://doi.org/10.1016/j.bbrc.2011.03.039
Sun Z, Shi Y, Shen Y, Cao L, Zhang W, Guan X (2015) Analysis of different HER-2 mutations in breast cancer progression and drug resistance. J Cell Mol Med 19(12):2691–2701. https://doi.org/10.1111/jcmm.12662
Carrick S, Parker S, Thornton CE, Ghersi D, Simes J, Wilcken N (2009) Single agent versus combination chemotherapy for metastatic breast cancer. Coch Database Syst Rev. https://doi.org/10.1002/14651858.CD003372.pub3
Petrelli F, Ghidini M, Lonati V, Tomasello G, Borgonovo K, Ghilardi M, Cabiddu M, Barni S (2017) The efficacy of lapatinib and capecitabine in HER-2 positive breast cancer with brain metastases: a systematic review and pooled analysis. Eur J Cancer 84:141–148. https://doi.org/10.1016/j.ejca.2017.07.024
Geyer CE, Forster J, Lindquist D, Chan S, Romieu CG, Pienkowski T, Jagiello-Gruszfeld A, Crown J, Chan A, Kaufman B, Skarlos D, Campone M, Davidson N, Berger M, Oliva C, Rubin SD, Stein S, Cameron D (2006) Lapatinib plus capecitabine for HER2-positive advanced breast cancer. N Engl J Med 355(26):2733–2743. https://doi.org/10.1056/NEJMoa064320
Cameron D, Casey M, Oliva C, Newstat B, Imwalle B, Geyer CE (2010) Lapatinib plus capecitabine in women with HER-2-positive advanced breast cancer: final survival analysis of a phase III randomized trial. Oncologist 15(9):924–934. https://doi.org/10.1634/theoncologist.2009-0181
Landis MW, Pawlyk BS, Li T, Sicinski P, Hinds PW (2006) Cyclin D1-dependent kinase activity in murine development and mammary tumorigenesis. Cancer Cell 9(1):13–22. https://doi.org/10.1016/j.ccr.2005.12.019
Yu Q, Geng Y, Sicinski P (2001) Specific protection against breast cancers by cyclin D1 ablation. Nature 411(6841):1017–1021. https://doi.org/10.1038/35082500
Yu Q, Sicinska E, Geng Y, Ahnstrom M, Zagozdzon A, Kong Y, Gardner H, Kiyokawa H, Harris LN, Stal O, Sicinski P (2006) Requirement for CDK4 kinase function in breast cancer. Cancer Cell 9(1):23–32. https://doi.org/10.1016/j.ccr.2005.12.012
Witkiewicz AK, Cox D, Knudsen ES (2014) CDK4/6 inhibition provides a potent adjunct to Her2-targeted therapies in preclinical breast cancer models. Genes Cancer 5(7–8):261–272
Goel S, Wang Q, Watt AC, Tolaney SM, Dillon DA, Li W, Ramm S, Palmer AC, Yuzugullu H, Varadan V, Tuck D, Harris LN, Wong KK, Liu XS, Sicinski P, Winer EP, Krop IE, Zhao JJ (2016) Overcoming therapeutic resistance in HER2-positive breast cancers with CDK4/6 inhibitors. Cancer Cell 29(3):255–269. https://doi.org/10.1016/j.ccell.2016.02.006
Tanner M, Kapanen AI, Junttila T, Raheem O, Grenman S, Elo J, Elenius K, Isola J (2004) Characterization of a novel cell line established from a patient with Herceptin-resistant breast cancer. Mol Cancer Ther 3(12):1585–1592
Holford NH, Sheiner LB (1982) Kinetics of pharmacologic response. Pharmacol Ther 16(2):143–166. https://doi.org/10.1016/0163-7258(82)90051-1
Lobo ED, Balthasar JP (2002) Pharmacodynamic modeling of chemotherapeutic effects: application of a transit compartment model to characterize methotrexate effects in vitro. AAPS Pharmsci 4(4):E42. https://doi.org/10.1208/ps040442
Hamed SS, Straubinger RM, Jusko WJ (2013) Pharmacodynamic modeling of cell cycle and apoptotic effects of gemcitabine on pancreatic adenocarcinoma cells. Cancer Chemother Pharmacol 72(3):553–563. https://doi.org/10.1007/s00280-013-2226-6
Miao X, Koch G, Ait-Oudhia S, Straubinger RM, Jusko WJ (2016) Pharmacodynamic modeling of cell cycle effects for gemcitabine and trabectedin combinations in pancreatic cancer cells. Front Pharmacol 7:421. https://doi.org/10.3389/fphar.2016.00421
Vaidya TR, Ande A, Ait-Oudhia S (2019) Combining multiscale experimental and computational systems pharmacological approaches to overcome resistance to HER2-targeted therapy in breast cancer. J Pharmacol Exp Ther 369(3):531–545. https://doi.org/10.1124/jpet.118.255752
Valabrega G, Capellero S, Cavalloni G, Zaccarello G, Petrelli A, Migliardi G, Milani A, Peraldo-Neia C, Gammaitoni L, Sapino A, Pecchioni C, Moggio A, Giordano S, Aglietta M, Montemurro F (2011) HER2-positive breast cancer cells resistant to trastuzumab and lapatinib lose reliance upon HER2 and are sensitive to the multitargeted kinase inhibitor sorafenib. Breast Cancer Res Treat 130(1):29–40. https://doi.org/10.1007/s10549-010-1281-5
Carpenter RL, Lo HW (2013) Regulation of apoptosis by HER2 in breast cancer. J Carcinogen Mutagen. https://doi.org/10.4172/2157-2518.S7-003
Dayneka NL, Garg V, Jusko WJ (1993) Comparison of four basic models of indirect pharmacodynamic responses. J Pharmacokinet Biopharm 21(4):457–478
Mager DE, Jusko WJ (2001) Pharmacodynamic modeling of time-dependent transduction systems. Clin Pharmacol Ther 70(3):210–216. https://doi.org/10.1067/mcp.2001.118244
Sharma A, Ebling WF, Jusko WJ (1998) Precursor-dependent indirect pharmacodynamic response model for tolerance and rebound phenomena. J Pharm Sci 87(12):1577–1584. https://doi.org/10.1021/js980171q
Sun YN, Jusko WJ (1998) Transit compartments versus gamma distribution function to model signal transduction processes in pharmacodynamics. J Pharm Sci 87(6):732–737. https://doi.org/10.1021/js970414z
Sobol IM (2001) Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Math Comput Simul 55(1–3):271–280
Herman JD, Usher W (2017) SALib: An open-source python library for sensitivity analysis. J Open Source Software 2(9):97
Ramakrishnan V, Mager DE (2019) Pharmacodynamic models of differential bortezomib signaling across several cell lines of multiple myeloma. CPT 8(3):146–157. https://doi.org/10.1002/psp4.12358
Saltelli A (2002) Making best use of model evaluations to compute sensitivity indices. Comput Phys Commun 145(2):280–297
Saltelli A, Annoni P, Azzini I, Campolongo F, Ratto M, Tarantola S (2010) Variance based sensitivity analysis of model output. Design and estimator for the total sensitivity index. Comput Phys Commun 181(2):259–270
Zhang XY, Trame MN, Lesko LJ, Schmidt S (2015) Sobol sensitivity analysis: a tool to guide the development and evaluation of systems pharmacology models. CPT 4(2):69–79. https://doi.org/10.1002/psp4.6
Molins EAG, Jusko WJ (2018) Assessment of three-drug combination pharmacodynamic interactions in pancreatic cancer cells. AAPS J 20(5):80. https://doi.org/10.1208/s12248-018-0235-4
Earp J, Krzyzanski W, Chakraborty A, Zamacona MK, Jusko WJ (2004) Assessment of drug interactions relevant to pharmacodynamic indirect response models. J Pharmacokinet Pharmacodyn 31(5):345–380. https://doi.org/10.1007/s10928-004-8319-4
Hafner M, Mills CE, Subramanian K, Chen C, Chung M, Boswell SA, Everley RA, Liu C, Walmsley CS, Juric D, Sorger PK (2019) Multiomics profiling establishes the polypharmacology of FDA-approved CDK4/6 inhibitors and the potential for differential clinical activity. Cell Chem Biol 26(8):1067–1080. https://doi.org/10.1016/j.chembiol.2019.05.005
Nahta R, Yuan LX, Du Y, Esteva FJ (2007) Lapatinib induces apoptosis in trastuzumab-resistant breast cancer cells: effects on insulin-like growth factor I signaling. Mol Cancer Ther 6(2):667–674. https://doi.org/10.1158/1535-7163.MCT-06-0423
Wainberg ZA, Anghel A, Desai AJ, Ayala R, Luo T, Safran B, Fejzo MS, Hecht JR, Slamon DJ, Finn RS (2010) Lapatinib, a dual EGFR and HER2 kinase inhibitor, selectively inhibits HER2-amplified human gastric cancer cells and is synergistic with trastuzumab in vitro and in vivo. Clin Cancer Res 16(5):1509–1519. https://doi.org/10.1158/1078-0432.CCR-09-1112
Yoshikawa R, Kusunoki M, Yanagi H, Noda M, Furuyama JI, Yamamura T, Hashimoto-Tamaoki T (2001) Dual antitumor effects of 5-fluorouracil on the cell cycle in colorectal carcinoma cells: a novel target mechanism concept for pharmacokinetic modulating chemotherapy. Can Res 61(3):1029–1037
Marchal JA, Boulaiz H, Suarez I, Saniger E, Campos J, Carrillo E, Prados J, Gallo MA, Espinosa A, Aranega A (2004) Growth inhibition, G(1)-arrest, and apoptosis in MCF-7 human breast cancer cells by novel highly lipophilic 5-fluorouracil derivatives. Invest New Drugs 22(4):379–389. https://doi.org/10.1023/B:DRUG.0000036680.52016.5f
Manning BD, Toker A (2017) AKT/PKB signaling: navigating the network. Cell 169(3):381–405. https://doi.org/10.1016/j.cell.2017.04.001
Xia W, Petricoin EF 3rd, Zhao S, Liu L, Osada T, Cheng Q, Wulfkuhle JD, Gwin WR, Yang X, Gallagher RI, Bacus S, Lyerly HK, Spector NL (2013) An heregulin-EGFR-HER3 autocrine signaling axis can mediate acquired lapatinib resistance in HER2+ breast cancer models. Breast Cancer Res 15(5):R85. https://doi.org/10.1186/bcr3480
Franco J, Balaji U, Freinkman E, Witkiewicz AK, Knudsen ES (2016) Metabolic reprogramming of pancreatic cancer mediated by CDK4/6 inhibition elicits unique vulnerabilities. Cell Rep 14(5):979–990. https://doi.org/10.1016/j.celrep.2015.12.094
Zacharek SJ, Xiong Y, Shumway SD (2005) Negative regulation of TSC1-TSC2 by mammalian D-type cyclins. Can Res 65(24):11354–11360. https://doi.org/10.1158/0008-5472.CAN-05-2236
Bubici C, Papa S (2014) JNK signalling in cancer: in need of new, smarter therapeutic targets. Br J Pharmacol 171(1):24–37. https://doi.org/10.1111/bph.12432
Davis RJ (2000) Signal transduction by the JNK group of MAP kinases. Cell 103(2):239–252. https://doi.org/10.1016/s0092-8674(00)00116-1
Dhanasekaran DN, Reddy EP (2008) JNK signaling in apoptosis. Oncogene 27(48):6245–6251. https://doi.org/10.1038/onc.2008.301
Spector NL, Robertson FC, Bacus S, Blackwell K, Smith DA, Glenn K, Cartee L, Harris J, Kimbrough CL, Gittelman M, Avisar E, Beitsch P, Koch KM (2015) Lapatinib plasma and tumor concentrations and effects on HER receptor phosphorylation in tumor. PLoS ONE 10(11):e0142845. https://doi.org/10.1371/journal.pone.0142845
Acknowledgements
The authors would like to thank Alexander Franco Anusha Ande and Brett Fleisher for technical assistance.
Author information
Authors and Affiliations
Contributions
Participated in research design: YLF, VR, TRV and SAO. Conducted experiments: YLF and LP. Contributed new reagents or analytic tools: YLF, VR, TRV, HM and SAO. Performed data analysis: YLF, VR, TRV, HM and SAO. Wrote or contributed to the writing of the manuscript: YLF, VR, TRV, HM and SAO.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10928_2020_9732_MOESM1_ESM.tif
Supplementary file1 (TIF 44 KB) Supplementary Figure 1: Observations vs. individual prediction plot for estimation of 72-h time course interaction parameter (Ψ). The solid line represents the identity line, while the symbols represent observed data
10928_2020_9732_MOESM2_ESM.tif
Supplementary file2 (TIF 598 KB) Supplementary Figure 2: Concentration-effect surface plots for abemaciclib and 5-fluorouracil (A), abemaciclib and lapatinib (B), and lapatinib and 5-fluorouracil (C). The surface represents predicted cell viability assuming a psi of 1. Blue points below the simulated surface represent concentrations exhibiting additive to synergistic effects, while red points above the surface represent concentrations exhibiting antagonistic effects
Rights and permissions
About this article
Cite this article
Franco, Y.L., Ramakrishnan, V., Vaidya, T.R. et al. A quantitative systems pharmacological approach identified activation of JNK signaling pathway as a promising treatment strategy for refractory HER2 positive breast cancer. J Pharmacokinet Pharmacodyn 48, 273–293 (2021). https://doi.org/10.1007/s10928-020-09732-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-020-09732-x