Abstract
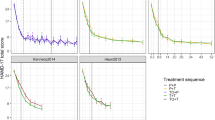
The aim of this paper is to provide a systematic methodology for modelling longitudinal data to be used in contexts of limited or even absent knowledge of the physiological mechanism underlying the disease time course. Adopting a system-theoretic paradigm, a population response model is developed where the clinical endpoint is described as a function of the patient’s health state. In particular, a continuous-time stochastic approach is proposed where the clinical score and its time-derivative summarize the patient’s health state affected by a random term accounting for exogenous unpredictable factors. The proposed approach is validated on experimental data from the placebo and drug arms of a Phase II depression trial. Since some subjects in the trial may undergo changes in their treatment dose due to the flexible dosing scheme, dose escalations are modelled as instantaneous perturbations on the state. In its simplest form—an integrated Wiener process—was able to correctly capture the individual responses in both treatment arms. However, a better description of inter-individual variability was obtained by means of a stable Markovian model. Parameter estimation has been carried out according to the empirical Bayes method.






Similar content being viewed by others
References
Mould DR, Denman NG, Duffull S (2007) Using disease progression model as a tool to detect drug effect. Clin Pharmacol Ther 82:81–86. doi:10.1038/sj.clpt.6100228
Shang EY, Gibbs MA, Landen JW, Krams M, Russell T, Denman NG, Mould DR (2009) Evaluation of structural models to describe the effect of placebo upon the time course of major depressive disorder. J Pharmacokinet Pharmacodyn 36:63–80. doi:10.1007/s10928-009-9110-3
Gomeni R, Merlo-Pich E (2006) Bayesian modelling and ROC analysis to predict placebo responders using clinical score measured in the initial weeks of treatment in depression trials. Br J Clin Pharmacol 63:595–613. doi:10.1111/j.1365-2125.2006.02815.x
Nucci G, Gomeni R, Poggesi I (2009) Model-based approaches to increase efficiency of drug development in schizophrenia: a can’t miss opportunity. Expert Opin Drug Discov 4:837–856. doi:10.1517/17460440903036073
Santen G, Danhof M, Della Pasqua O (2008) Evaluation of the treatment response in depression studies using a Bayesian parametric cure rate model. J Psychiatr Res 42:1189–1197. doi:10.1016/j.jpsychires.2007.11.009
Holford N, Li J, Benincosa L, Birath M (2002) Population disease progress models for the time course of HAMD score in depressed patients receiving placebo in anti-depressant clinical trials. In: Population Approach Group in Europe (PAGE) 11th Meeting, Abstract 311, http://www.page-meeting.org/?abstract=311
Reddy VP, Kozielska M, Johnson M, Vermeulen A, de Greef R, Liu J, Groothuis GMM, Danhof M, Proost JH (2011) Structural models describing placebo treatment effects in schizophrenia and other neuropsychiatric disorders. Clin Pharmacokinet 50:429–450
Marostica E, Russu A, Gomeni R, Zamuner S, De Nicolao G (2013) A PCA approach to population analysis: with application to a Phase II depression trial. J Pharmacokinet Pharmacodyn 40:213–227. doi:10.1007/s10928-013-9304-6
Marostica E, Russu A, Gomeni R, Zamuner S, De Nicolao G (2011). Population modelling of patient responses in antidepressant studies: a stochastic approach
Rowland M, Sheiner LB, Steimer JL (1985) Variability in drug therapy: description, estimation and control. Raven Press, New York
Morters P, Peres Y (2010) Brownian motion. Cambridge University Press, UK
Hamilton M (1960) A rating scale for depression. J Neurol Neurosur Psychiatr 23:56–62. doi:10.1136/jnnp.23.1.56
Neve M, De Nicolao G, Marchesi L (2007) Nonparametric identification of population models via Gaussian processes. Automatica 43:1134–1144. doi:10.1016/j.automatica.2006.12.024
Russu A, Poggesi I, Gomeni R, De Nicolao G (2011) Bayesian population modeling of Phase I dose escalation studies: gaussian process versus parametric approaches. IEEE Trans Biomed Engineering 58:3156–3164. doi:10.1109/TBME.2011.2164614
Robbins H (1964) The Empirical Bayes Approach to Statistical Decision Problems. Ann Math Stat 35:1–20. doi:10.1214/aoms/1177703729
R Development Core Team (2010) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, ISBN 3-900051-07-0, http://www.R-project.org
Karlsson M, Holford N (2008) A tutorial on Visual Predictive Checks. In: Population Approach Group in Europe (PAGE) 17th Meeting, Abstract 1434. http://www.page-meeting.org/default.asp?abstract=1434
Russu A, Marostica E, De Nicolao G, Hooker AC, Poggesi I, Gomeni R, Zamuner S (2012) Joint modeling of efficacy, dropout, and tolerability in flexible-dose trials: a case study in depression. Clin Pharmacol Ther 91:863–871. doi:10.1038/clpt.2011.322
Neve M, De Nicolao G, Marchesi L (2008) Nonparametric identification of population models: an MCMC approach. IEEE Trans Biomed Eng 55:41–50. doi:10.1109/TBME.2007.902240
Gomeni R, Lavergne A, Merlo-Pich E (2009) Modeling placebo response in depression trials using longitudinal model with informative dropout. Eur J Pharm Sci 36:4–10. doi:10.1016/j.ejps.2008.10.025
Hu C, Sale ME (2003) A joint model for nonlinear longitudinal data with Informative dropout. J Pharmacokinet Pharmacodyn 30:83–103. doi:10.1023/A:1023249510224
Hu C, Szapary PO, Yeilding N, Zhou H (2011) Informative dropout modeling of longitudinal ordered categorical data and model validation: application to exposure-response modeling of physician’s global assessment score for ustekinumab in patients with psoriasis. J Pharmacokinet Pharmacodyn 38:237–260. doi:10.1007/s10928-011-9191-7
Kitagawa (1977) An algorithm for solving the matrix equation X = FXF’ + S. J Control 25:745–753. doi:10.1080/00207177808922369
Poggio T, Girosi F (1990) Networks for approximation and learning. Proc IEEE 78:1481–1497. doi:10.1109/5.58326
Girosi F, Jones M, Poggio T (1995) Regularization theory and neural networks architectures. Neural Comput 7:219–269. doi:10.1162/neco.1995.7.2.219
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1: Auto-covariance function of Markov processes
In this appendix, the formulas to derive the auto-covariance functions \(\overline{R}(t, \tau )\) and \(\widetilde{R}(t, \tau )\) for the average curve and individual shift, respectively, are detailed. First, some basic properties of the auto-covariance of Markov processes are recalled. Then, we consider their application to the model used for the typical response and the individual shift. Differently from [13, 14, 19] the present derivation accounts for non-zero initial variance of the individual shift and the possible presence of exogenous factors as dose changes.
Definition of auto-covariance
A random process can be seen as a sequence of random variables over time. For example, one common random process is the white Gaussian noise where each random variable \(w(t)\) is a Gaussian random variable with zero-mean and variance \(\lambda ^2\). Given two distinct time points t and \(\tau \), the associated values of the random process, i.e. the random variables \(w(t)\) and \(w(\tau )\), are characterized by a covariance \(R(t, \tau ) = cov[w(t), w(\tau )]\) for all possible choices of \(t\) and \(\tau \). The function \(R(t, \tau )\) is commonly known as the auto-covariance function of the process \(w(t)\).
Basic properties
Given the following system of equations:
where \(x(t_0) \sim N(0, P_0), w(t) \sim WGN(\lambda ^2)\), and \(E[x(t_0)w(t)] = 0\), its solution \(x(t)\) is given by the Lagrange formula
Let \(P(t) = var[x(t)] = E[x(t)x(t)^T]\) be the variance of the state. The auto-covariance function of \(x(t)\) is given by
Accordingly, the auto-covariance function of the output \(z(t)\) is
The state variance \(P(t)\) satisfies the following Lyapunov differential equation [23]
from which
and
Auto-covariance function of the population curve
Based on the matrices \(A, B\), and \(C\) defined in (4) and recalling that \(P_0 = 0\), it can be found that
and, according to (17)
where
Moreover, based on (18), the auto-covariance function \(\overline{R}(t, \tau )\) of the population curve is
Auto-covariance function of the individual shifts: non-escalating subjects
In this case
and, based on (20),
Then, the auto-covariance function \(\widetilde{R}(t, \tau )\) is
where
where \(f_{11}(t) = f_{22}(t) = e^{at}\) and \(f_{12}(t) = te^{at}\).
Auto-covariance function of the individual shifts: escalating subjects
In order to define the auto-covariance function \(\widetilde{R}(t, \tau )\) for the escalating subjects, three scenarios must be taken into account with respect to the occurrence of the time of dose change \(t_{esc}\):
-
1.
\(t < \tau < t_{esc}\)
-
2.
\(t < t_{esc} < \tau \)
-
3.
\(t_{esc} < t < \tau \)
The auto-covariance function relative to the first two scenarios coincides with the one defined in (26) for the non-escalating subjects. For the third scenario, Eq. (25) has to be extended to account for the exogenous event:
More specifically, the elements of the matrix (20) are defined as:
with \(f_{11}(t) = f_{22}(t) = e^{at}\) and \(f_{12}(t) = te^{at}\). Therefore, the auto-covariance function \(\widetilde{R}(t, \tau )\) becomes
Appendix 2: Parameter estimation
The algorithms reported in this section are essentially the same as in [13, 14]. They are repeated here for the sake of self-consistency.
As explained in Section Methods, the initial conditions \(\overline{x}(0)\) for the average curve are characterized by infinite variance. Therefore, Eq. (2) can be reformulated as
where \(\varPhi ^T(t) = [1 \quad t]\) and \(\varPsi \sim N(0, \infty I)\).
Moreover, let \(M\) be the total number of subjects and \(n = \sum \nolimits _{i = 1}^{M} n_i\) the total number of data, Eq. (31) can be defined in vector notation:
The variance of the vectors \(\mathbf {\overline{z}}\) and \(\mathbf {\widetilde{z}}\), which are \({\mathbf {\overline{R}}}(t, \tau )\) and \({\mathbf {\widetilde{R}}}(t, \tau )\), respectively, can be defined as
where \({\mathbf {\widetilde{R}}}_i\) for the \(i\)-th subject is
Based on the definition of the auto-covariances \(\overline{R}\) and \(\widetilde{R}_i\), model (31) can be formulated as
where \(c^k_i\) represent the weights of the regularization network [24, 25] and
Let \(\alpha = [\overline{\lambda }^2 \quad \widetilde{\lambda }^2 \quad \sigma ^2_0 \quad \sigma ^2_1 \quad \sigma ^2_{\varDelta } \quad \sigma ^2]\) be the vector of the unknown hyperparameters. According to the Empirical Bayes paradigm [14, 15], two steps are performed: initially \(\alpha \) is estimated through the Maximum Likelihood (ML) approach, by maximizing the marginal likelihood \(p(Y|\alpha )\); once the ML estimates of the hyperparameters \(\alpha ^{ML}\) are available, they are plugged into the model and the Bayesian estimate is calculated through the standard conditional posterior formula for linear Gaussian models. Uninformative priors were assumed in this analysis.
In particular,
More technical details can be found in [13].
Rights and permissions
About this article
Cite this article
Marostica, E., Russu, A., Gomeni, R. et al. Continuous-time Markov modelling of flexible-dose depression trials. J Pharmacokinet Pharmacodyn 41, 625–638 (2014). https://doi.org/10.1007/s10928-014-9389-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-014-9389-6