Abstract
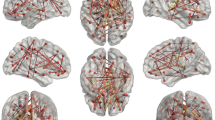
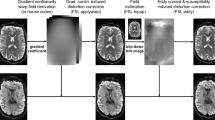
High Angular Resolution Diffusion Imaging (HARDI) is a type of brain imaging that collects a very large amount of data, and if many subjects are considered then it amounts to a big data framework (e.g., the human connectome project has 20 Terabytes of data). HARDI is also becoming increasingly relevant for clinical settings (e.g., detecting early cerebral ischemic changes in acute stroke, and in pre-clinical assessment of white matter-WM anatomy using tractography). Thus, this method is becoming a routine assessment in clinical settings. In such settings, the computation time is critical, and finding forms of reducing the processing time in high computation processes such as Diffusion Spectrum Imaging (DSI), a form of HARDI data, is very relevant to increase data-processing speed. Here we analyze a method for reducing the computation time of the dMRI-based axonal orientation distribution function h by using Monte Carlo sampling-based methods for voxel selection. Results evidenced a robust reduction in required data sampling of about 50 % without losing signal’s quality. Moreover, we show that the convergence to the correct value in this type of Monte Carlo HARDI/DSI data-processing has a linear improvement in data-processing speed of the ODF determination. Although further improvements are needed, our results represent a promissory step for future processing time reduction in big data.






Similar content being viewed by others
References
Lichtman, J.W., Pfister, H., and Shavit, N., The big data challenges of connectomics. Nat. Neurosci. 17(11):1448–1454, 2014.
Barkhof, F., Haller, S., and Rombouts, S.A., Resting-state functional MR imaging: A new window to the brain. Radiology. 272(1):29–49, 2014.
Worbe, Y., Neuroimaging signature of neuropsychiatric disorders. Curr. Opin. Neurol. 28(4):358–364, 2015.
Zhou, J., and Seeley, W.W., Network dysfunction in Alzheimer's disease and frontotemporal dementia: Implications for psychiatry. Biol. Psychiatry. 75(7):565–573, 2014.
Sharp, D.J., Scott, G., and Leech, R., Network dysfunction after traumatic brain injury. Nat. Rev. Neurol. 10(3):156–166, 2014.
Craddock, R.C., Tungaraza, R.L., and Milham, M.P., Connectomics and new approaches for analyzing human brain functional connectivity. Gigascience. 4:13, 2015.
Marder, E., Understanding brains: details, intuition, and big data. PLoS Biol. 13(5):e1002147, 2015.
Boubela, R.N., et al., Big data approaches for the analysis of large-scale fMRI data using apache spark and GPU processing: A demonstration on resting-state fMRI data from the human connectome project. Front Neurosci. 9:492, 2015.
Conturo, T.E., Lori, N.F., Cull, T.S., Akbudak, E., Snyder, A.Z., et al., Tracking neuronal fiber pathways in the living human brain. Proc. Natl. Acad. Sci. U. S. A. 96:10422–10427, 1999.
Lori, N.F., Akbudak, E., Shimony, J.S., Cull, T.S., Snyder, A.Z., et al., Diffusion tensor fiber tracking of human brain connectivity: aquisition methods, reliability analysis and biological results. NMR Biomed. 15:494―515, 2002.
Tuch, D.S., Q-ball imaging. Magn. Reson. Med. 52:1358–1372, 2004.
Behrens, T.E.J., Berg, H.J., Jbabdi, S., Rushworth, M.F.S., and Woolrich, M.W., Probabilistic diffusion tractography with multiple fibre orientations: What can we gain? Neuroimage. 34:144–155, 2007.
Wedeen, V.J., Wang, R.P., Schmahmann, J.D., Benner, T., Tseng, W.Y.I., et al., Diffusion spectrum magnetic resonance imaging (DSI) tractography of crossing fibers. Neuroimage. 41:1267–1277, 2008.
Raffelt, D., Tournier, J.D., Rose, S., Ridgway, G.R., Henderson, R., et al., Apparent Fibre Density: A novel measure for the analysis of diffusion-weighted magnetic resonance images. Neuroimage. 59:3976–3994, 2012.
Wedeen, V.J., Rosene, D.L., Wang, R., Dai, G., Mortazavi, F., et al., The geometric structure of the brain fiber pathways. Science. 335:1628–1634, 2012.
Dani, A., Huang, B., Bergan, J., Dulac, C., and Zhuang, X., Superresolution imaging of chemical synapses in the brain. Neuron. 68:843–856, 2010.
Hawrylycz, M.J., Lein, E.S., Guillozet-Bongaarts, A.L., Shen, E.H., Ng, L., et al., An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 489:391–399, 2012.
Tuch, D.S., Reese, T.G., Wiegell, M.R., and Wedeen, V.J., DMRI of complex neural architecture. Neuron. 40:885–895, 2003.
Hill, S.L., Wang, Y., Riachi, I., Schürmann, F., and Markram, H., Statistical connectivity provides a sufficient foundation for specific functional connectivity in neocortical neural microcircuits. Proc. Natl. Acad. Sci. U. S. A. 109:E2885–E2894, 2012.
Wang, R., Benner, T., Sorensen, A.G., and Wedeen, V.J., Diffusion toolkit: A software package for diffusion imaging data processing and tractography. Proc. Intl. Soc. Mag. Reson. Med. 15:3720, 2007.
Assaf, Y., Blumenfeld-Katzir, T., Yovel, Y., and Basser, P.J., AxCaliber: a method for measuring axon diameter distribution from dMRI. Magn. Reson. Med. 59:1347–1354, 2008.
Milne, M.L., and Conradi, M.S., Multi-exponential signal decay from diffusion in a single compartment. J. Magn. Reson. 197:87–90, 2009.
U.C.L.A. (n.d.) LONI Image Data Archive (IDA). Available: https://ida.loni.ucla.edu/login.jsp. Accessed 16 November 2012. (2012)
Zhang, Y., Brady, M., and Smith, S., Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans. Med. Imaging. 20:45–57, 2001.
Acknowledgments
We thank the financial support by QREN, FEDER, COMPETE, Investigador FCT, FCT Ciencia 2007, FCT PTDC/SAU-BEB/100147/2008, FCT Project Scope UID/CEC/00319/2013, and the ERASMUS projects (FCT stands for “Fundação para a Ciência e Tecnologia”). We are thankful the relevant scientific conversations with Alard Roebroeck, Rainer Goebel, Van Wedeen, and Gina Caetano. Data collection for this work was in part from “Human Connectome Project” (HCP; Principal Investigators: Bruce Rosen, M.D., Ph.D., Arthur W. Toga, Ph.D., Van J. Weeden, MD). HCP funding was provided by the National Institute of Dental and Craniofacial Research (NIDCR), the National Institute of Mental Health (NIMH), and the National Institute of Neurological Disorders and Stroke (NINDS). HCP data are disseminated by the Laboratory of Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Systems-Level Quality Improvement
An erratum to this article is available at http://dx.doi.org/10.1007/s10916-016-0683-2.
Rights and permissions
About this article
Cite this article
Lori, N.F..., Ibañez, A., Lavrador, R. et al. Processing Time Reduction: an Application in Living Human High-Resolution Diffusion Magnetic Resonance Imaging Data. J Med Syst 40, 243 (2016). https://doi.org/10.1007/s10916-016-0594-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-016-0594-2