Abstract
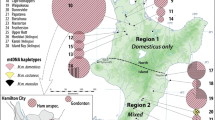
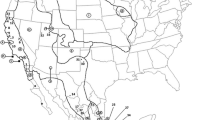
The two currently recognized species of kangaroo mice, Microdipodops megacephalus and M. pallidus, inhabit sandy soils of the Great Basin Desert in western North America. Given their habitat specificity and the fluctuating climate throughout the Pleistocene, kangaroo mice likely endured a turbulent biogeographic history that resulted in disjunct distributions and isolation of genetic lineages. Recent phylogenetic investigations using mitochondrial data have revealed several mitochondrial clades within this genus that may represent cryptic species. These mitochondrial clades are genetically unique, occupy relatively small distributions, and, as such, may be at an increased risk of extinction due to climate change and extensive recent habitat alteration. Herein, we apply haplotype network, population genetic, and historical demographic analyses to mitochondrial data of each Micropdipodops species and mitochondrial clade to assess conservation genetics within kangaroo mice. Results indicate that each mitochondrial clade is a distinct lineage with little to no gene flow occurring among clades. Additionally, historical demographic analyses support past population expansions and identify locations of past refugium for each distinct lineage. Although mitochondrial data indicate that the clades appear to be in approximate genetic equilibrium and have not suffered any extreme bottlenecks over time, there is still concern for the survival of smaller and more vulnerable Microdipodops subpopulations due to impending habitat threats in the Great Basin Desert.




Similar content being viewed by others
References
Antevs E (1952) Cenozoic climates of the Great Basin. Geol Rundsch 40:94–108
Austin GT, Murphy DD (1987) Zoogeography of Great Basin butterflies: patterns of distribution and differentiation. Gt Basin Nat Mem 47:186–201
Beever EA, Brussard PF, Berger J (2003) Patterns of apparent extirpation among isolated populations of pikas (Ochotona princeps) in the Great Basin. J Mammal 84:37–54
Beever EA, Ray C, Wilkening JL, Brussard PF, Mote PW (2011) Contemporary climate change alters the pace and rivers of extinction. Global Change Biol 17:2054–2070
Benson LV (1981) Paleoclimatic significance of lake-level fluctuations in the Lahontan Basin. Quaternary Res 16:390–403
Benson LV, Currey DR, Dorn RI, Lajoie KR, Oviatt CG, Robinson SW, Smith GI, Stine S (1990) Chronology of expansion and contraction of the four Great Basin lake systems during the past 35,000 years. Palaeogeog Palaeoclimatol Palaeoecol 78:241–286
Blois JL, Hadly EA (2009) Mammalian response to Cenozoic climatic change. Annu Rev Earth Planet Sci 37:181–208
Britten HB, Brussard PF, Murphy DD, Austin GT (1994) Colony isolation and isozyme variability of the Western Fritillary, Speyeria nokomis apacheana (Nymphalidae) in the western Great Basin. Great Basin Nat 54:97–105
Britten HB, Brussard PF, Murphy DD, Ehrlich PR (1995) A test for isolation-by-disance in central Rocky Mountain and Great Basin populations of Edith’s Checkerspot Butterfly (Euphydryas editha). J Hered 86:204–210
Chaplin SJ, Gerrard RA, Watson HM, Master LL, Flack SR (2000) The geography of imperilment: targeting conservation toward critical biodiversity areas. In: Stein BA, Kutner LS, Adams JS (eds) Precious Hertitage: The Status of Biodiversity in the United States. The Nature Conservancy and Association for Biodiversity Information, Oxford University Press, Inc., New York, pp 159–199
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Cronquist A, Holmgren AH, Holmgren NH, Reveal JL (1972) Intermountain Flora: Vascular Plants of the Intermountain West, U.S.A., vol 1. Hafner, New York
Davis EB (2005) Mammalian beta diversity in the Great Basin, western USA: palaeontological data suggest deep origin of modern macroecological structure. Global Ecol Biogeog 14:479–490
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192
Dupanloup L, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Epps TM, Britten HB, Rust RW (1998) Historical biogeography of Eusattus muricatus (Coleoptera: Tenebrionidae) within the Great Basin, western North America. J Biogeogr 25:957–968
Excoffier L, Heckel G (2006) Computer programs for population genetics data analysis: a survival guide. Nat Rev Genet 7:745–758
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform 1:47–50
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances using DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics 131:479–491
Fiero B (1986) The Geology of the Great Basin. University of Nevada Press, Reno
Fleishman E, Austin GT, Murphy DD (2001) Biogeography of Great Basin butterflies: revisiting patterns, paradigms, and climate change scenarios. Biol J Linn Soc 74:501–515
Fleishman E, Dobkin DS (2009) Current and potential elevational distributions of birds associated with pinyon-juniper woodlands in the central Great Basin, U.S.A. Restor Ecol 17:731–739
Floyd CH, Van Vuren DH, May B (2005) Marmots on Great Basin mountaintops: using genetics to test a biogeographic paradigm. Ecology 86:2145–2153
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Grayson DK (1993) The Desert’s Past: A Natural Prehistory of the Great Basin. Smithsonian Institution Press, Washington, DC
Grayson DK (2005) A brief history of Great Basin pikas. J Biogeogr 32:2103–2111
Grayson DK (2006) The late Quaternary biogeographic histories of some Great Basin mammals (western USA). Quaternary Sci Rev 25:2964–2991
Hafner DJ, Hafner JC (1998a) Microdipodops megacephalus Merriam 1891 Dark kangaroo mouse. In: Hafner DJ, Yensen E, Kirkland Jr. GL (eds) North American Rodents: Status Survey and Conservation Action Plan. IUCN (The World Conservation Union), Glan, Switzerland and Cambridge, UK, pp 79–80
Hafner DJ, Hafner JC (1998b) Microdipodops pallidus Merriam 1901 Pale kangaroo mouse. In: Hafner DJ, Yensen E, Kirkland Jr. GL (eds) North American Rodents: Status Survey and Conservation Action Plan. IUCN (The World Conservation Union), Gland, Switzerland and Cambridge, UK, pp 80–81
Hafner DJ, Hafner JC, Hafner MS (1979) Systematic status of kangaroo mice, genus Microdipodops: morphometric, chromosomal, and protein analyses. J Mammal 60:1–10
Hafner JC (1981) Evolution, Systematics, and Historical Biogeography of Kangaroo Mice, Genus Microdipodops. Dissertation, University of California, Berkeley, 269 pp
Hafner JC, Hafner DJ, Hafner MS (1996) Habitat selection and coexistence of species of kangaroo mice (Microdipodops). In: Genoways HH, Baker RJ (eds) Contributions in Mammalogy: A Memorial Volume Honoring Dr. J. Knox Jones, Jr. Museum of Texas Tech University, Lubbock, pp 249–259
Hafner JC, Hafner MS (1983) Evolutionary relationships of heteromyid rodents. Great Basin Nat Mem 7:3–29
Hafner JC, Light JE, Hafner DJ, Hafner MS, Reddington E, Rogers DS, Riddle BR (2007) Basal clades and molecular systematics of heteromyid rodents. J Mammal 87:1129–1145
Hafner JC, Reddington E, Craig MT (2006) Kangaroo mice (Microdipodops megacephalus) of the Mono Basin: phylogeography of a peripheral isolate. J Mammal 87:1204–1217
Hafner JC, Upham NS (2011) Phylogeography of the dark kangaroo mouse, Microdipodops megacephalus: cryptic lineages and dispersal routes in North America's Great Basin. J Biogeogr 38:1077–1097
Hafner JC, Upham NS, Reddington E, Torres CW (2008) Phylogeography of the pallid kangaroo mouse, Microdipodops pallidus: a sand-obligate endemic of the Great Bason, western North America. J Biogeogr 35:2101–2118
Hall ER (1941) Revision of the rodent genus Microdipodops. Field Mus Nat Hist Zool Ser 27:233–277
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos T Roy Soc B 351:1291–1298
Harpending RC (1994) Signature of ancient population growth in a low-resolution mitochondria DNA mismatch distribution. Hum Biol 66:591–600
Heled J, Drummond AJ (2008) Bayesian inference of population size history from multiple loci. BMC Evol Biol 8:289
Ho SYW, Shapiro B (2011) Skyline-plot methods for estimating demographic history from nucleotide sequences. Mol Ecol Resour 11:423–434
Johnson NK (1975) Controls of number of bird species on montane islands in the Great Basin. Evolution 29:545–567
Johnson NK (1978) Patterns of avian geography and speciation in the Intermountain Region. West N Am Naturalist 2:137–159
Johnson NK, Marten JA (1992) Macrogeographic patterns of morphometric and genetic variation in the sage sparrow complex. Condor 94:1–19
Knapp PA (1996) Cheatgrass (Bromus tectorum L.) dominance in the Great Basin Desert. Global Environ Change 6:37–52
Kramer AT, Fant JB, Ashley MV (2011) Influences of landscape and pollinators on population genetic structure: examples from three Penstemon (Plantaginaceae) species in the Great Basin. Am J Bot 98:109–121
Lance SL, Light JE, Jones KL, Hagen C, Hafner JC (2010) Isolation and characterization of 17 polymorphic microsatellite loci in the kangaroo mouse, genus Microdipodops (Rodentia: Heteromyidae). Conserv Genet Resour 2:139–141
Lawlor TE (1998) Biogeography of Great Basin mammals: paradigm lost? J Mammal 79:1111–1130
Linzey AV, Hammerson G (2008) Microdipodops megacephalus. IUCN 2011. IUCN Red List of Threatened Species. Version 2011.1. http://www.iucnredlist.org (accessed 22 September 2011)
Linzey AV, Hammerson G, Morefield J (2008) Microdipodops pallidus. IUCN 2011. IUCN Red List of Threatened Species. Version 2011.1. http://www.iucnredlist.org (accessed 22 September 2011)
McDonald KA, Brown JH (1992) Using montane mammals to model extinctions due to global change. Conserv Biol 6:409–415
Mehringer PJ (1986) Prehistoric environments. In: D’Azevedo WL (ed) Great Basin: Handbook of North American Indians, vol 11. Smithsonian Institution, Washington, DC, pp 31–50
Miller MP (2005) Alleles in Space (AIS): computer software for the joint analysis of interindividual spatial and genetic information. J Hered 96:722–724
Miller RF, Rose JA (1999) Fire history and western juniper encroachment in sagebrush steppe. J Range Manage 52:550–559
Moritz C (1994) Defining ‘evolutionary signficant units’ for conservation. Trends Ecol Evol 9:373–375
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Patton JL (2005) Family Heteromyidae. In: Wilson DE, Reeder DM (eds) Mammal Species of the World: A Taxonomic and Geographic Reference. 3rd edn. Johns Hopkins University Press, Baltimore, pp 844–858
Pellant M, Abbey B, Karl S (2004) Restoring the Great Basin Desert, USA: integrating science, management, and people. Environ Monit Assess 99:169–179
Porter JL, Rust RW (1996) Allozyme variation within five species of Aegialia (Coleoptera: Scarabaeidae). Ann Entomol Soc Am 89:710–721
Posada D, Crandall KA (2001) Intraspecific gene genealogies: trees grafting into networks. Trends Ecol Evol 16:7–45
Rambaut A, Drummond AJ (2007) Tracer v1.4, Available from http://beast.bio.ed.ac.uk/Tracer
Ray N, Currat M, Excoffier L (2003) Intra-deme molecular diversity in spatially expanding populations. Mol Biol Evol 20:76–86
Reveal JL (1979) Biogeography of the Intermountain Region: a speculative appraisal. Mentzelia 4:1–92
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol 9:552–569
Rogers MA (1991a) Evolutionary differentation within the northern Great Basin pocket gopher, Thomomys townsendii. I. Morphological variation. Great Basin Nat 51:109–126
Rogers MA (1991b) Evolutionary differentiation within the northern Great Basin pocket gopher, Thomomys townsendii. II. Genetic variation and biogeographic considerations. Great Basin Nat 51:127–152
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Simpkin JL, Britten HB, Brussard PF (2000) Effects of habitat fragmentation and differing mobility on the population structures of a Great Basin dragonfly (Sympetrum corruptum) and damselfly (Enallagma carunculatum). West N Am Naturalist 60:320–332
Smith GR (1978) Biogeography of intermountain fishes. Gt Basin Nat Mem 2:17–42
Smith RSU (1982) Sand dunes in the North American deserts. In: Bender GL (ed) Reference Handbook of the Deserts of North America. Greenwood Press, Westport, CT, pp 481–526
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Team RDC (2011) R: A language and environment for statistical computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org/
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132:619–633
Visser ME (2008) Keeping up with the warming world; assessing the rate of adaptation to climate change. Proc R Soc Lond Ser B 275:649–659
Weir BS, Cockerham CC (1984) Estimating F statistics for the analysis of population structure. Evolution 28:1358–1370
Wells PV (1983) Paleobiogeography of montane islands in the Great Basin since the Last Glaciopluvial. Ecol Monogr 53:341–382
Whisenant SG (1990) Changing fire frequencies of Idaho’s Snake River Plains: ecological and management implications. Proceedings of the symposium on cheatgrass invastion, shrub die-off, and other aspects of shrub biology and managment, vol General Technical Report INT-276
Wilcox BA, Murphy DD, Ehrlich PR, Austin GT (1986) Insular biogeography of the montane butterfly faunas in the Great Basin: comparison with birds and mammals. Oceologia 69:199–194
Yandell UG (1992) An allozyme analysis of whitebark pine (Pinus albicaulis Engl.). M.Sc. Thesis, University of Nevada, 59 pp
Acknowledgements
We thank F. Burbrink for advice regarding analytical methods. This is publication number 208 of the Center for Biosystematics and Biodiversity, at Texas A&M University. Support for this study was provided in part by the Nevada Department of Wildlife (contracts 05-21 and 08-15 to J.C.H.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Light, J.E., Hafner, J.C., Upham, N.S. et al. Conservation Genetics of Kangaroo Mice, Genus Microdipodops . J Mammal Evol 20, 129–146 (2013). https://doi.org/10.1007/s10914-012-9193-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10914-012-9193-2