Abstract
Tumor organoids mimic the architecture and heterogeneity of in vivo tumors and enable studies of collective interactions between tumor cells as well as with their surrounding microenvironment. Although tumor organoids hold significant promise as cancer models, they are also more costly and labor-intensive to cultivate than traditional 2D cell culture. We sought to identify critical factors regulating organoid growth ex vivo, and to use these observations to develop a more efficient organoid expansion method. Using time-lapse imaging of mouse mammary tumor organoids in 3D culture, we observed that outgrowth potential varies non-linearly with initial organoid size. Maximal outgrowth occurred in organoids with a starting size between ~10 to 1000 cells. Based on these observations, we developed a suspension culture method that maintains organoids in the ideal size range, enabling expansion from 1 million to over 100 million cells in less than 2 weeks and less than 3 hours of hands-on time. Our method facilitates the rapid, cost-effective expansion of organoids for CRISPR based studies and other assays requiring a large amount of organoid starting material.
Similar content being viewed by others
Introduction
A wealth of studies have demonstrated that 3D organotypic culture systems can faithfully recapitulate diverse aspects of the tissue architecture, heterogeneity, and spatial complexity of the in vivo microenvironment [30, 55, 57, 59, 60]. In recent years, organoids have been usefully applied to understanding diverse developmental and disease processes ranging from early embryogenesis [53, 62] to COVID-19 viral replication in different tissue types [64] to complex genotype-phenotype processes in degenerative brain diseases [19, 58].
Organoids have also emerged as important models for cancer [15, 37, 55, 60]. Tumor organoids can be isolated from experimental mouse models as well as human patient primary and metastatic tumors, allowing for the representation of a wide range of tumor phenotypes and genotypes. To observe and perturb complex interactions amongst tumor cells, normal cells, and stroma, multicellular tumor organoids are embedded in a variety of 3D matrix platforms [2, 11, 12, 20, 32, 43,44,45]. In this way, investigators have used tumor organoids in 3D culture to uncover essential aspects of tumor formation, metastasis, intercellular tumor-tumor and tumor-microenvironmental interactions, and therapeutic drug resistance [4,5,6, 10, 24, 46, 61].
Though tumor organoids hold significant promise, they also face technical challenges distinct from studies in immortalized 2D cell lines. 2D culture of cell lines enables robust, cost-effective expansion of starting material. This is particularly important for experiments that require a high starting input, such as genome-wide or drug library screening. In contrast, 3D organoid culture typically requires passaging variably sized organoids in a 3D gel, usually a basement membrane-rich extract such as Matrigel, every 1–2 weeks. The logistical and financial burdens of 3D culture are sometimes under-appreciated, but are important factors affecting the adoption of organoid techniques into mainstream use.
In this study, we characterize the parameters affecting tumor organoid outgrowth. We find that organoid growth is optimal within an intermediate size range. We find that commonly used protocols for passaging by partial enzymatic digestion do not maintain organoids in this ideal range, resulting in suboptimal outgrowth. Importantly, we identify conditions that allow tumor organoids to expand robustly in suspension, precluding the need for plating or depolymerization of solid Matrigel droplets, and allowing us to expand organoid lines by over 100-fold in less than 2 weeks.
Results
Intermediate Size Organoids Expand Optimally
Individual organoids vary in their starting size in 3D culture, but how starting organoid size affects eventual outgrowth potential is not well understood. Here we performed time-lapse imaging of tumor organoids isolated from the MMTV-PyMT mouse model of aggressive breast cancer, which has invasive and metastatic properties that are robustly modeled in organoid culture [6, 7, 13, 22, 36, 46, 47]. We embedded freshly derived MMTV-PyMT tumor organoids in basement membrane-rich extract (3D Matrigel) and directly observed the growth kinetics of 522 individual organoids (Fig. 1a, b). For each organoid, we measured the starting and ending areas to obtain an estimate of outgrowth. Importantly, organoid outgrowth varied considerably; some organoids had high growth rates in 3D Matrigel while others grew poorly or even shrank over time (Movie S1). Closer examination revealed that single tumor cells frequently underwent cell death, consistent with prior observations [7, 61]. More surprisingly, very small organoids (< 300 μm2 area) and very large organoids (>30,000 μm2) also showed inferior outgrowth compared with organoids at intermediate sizes (~300–30,000 μm2) (Fig. 1c).
Maximal organoid growth occurs within an intermediate size range which promotes proliferation and minimizes cell death. a Time lapse imaging of MMTV-PyMT organoids in 3D Matrigel. Right, summary of growth as change in area. See also Movie S1. b Organoid initial area and final area (after 4 days of time lapse imaging in 3D Matrigel) were measured to estimate growth over time. n = 522 organoids from 8 mice. Red line = fold change of 1 (no growth). c Organoid area growth rate ([final area-initial area]/initial area) from 4 days of time-lapse imaging in 3D Matrigel binned by initial organoid area. n = 8 mice, 522 organoids. P values = unpaired t-tests. d Mitotic rate assessed by phospho-Histone H3 immunofluorescence in organoids of varying sizes in 3D Matrigel. Each dot represents the % of cells pHH3+ in organoids derived from one biological replicate, n = 349 organoids, n = 14 to 65 organoids per mouse. Red line = median. P values = unpaired t-tests. e Propidium iodide (PI) staining of non-viable cells in organoids of varying sizes in 3D Matrigel. The PI+ core area was measured and converted to a volume to estimate the % of cells in the non-viable core region, relative to the total organoid volume. Each dot is an organoid (n = 267 organoids, n = 2 mice). Line = non-linear regression. Band = 95% CI of non-linear regression. f Organoid cell number growth rate was calculated from 4 days of time lapse imaging in Matrigel ([final volume-initial volume]/initial volume). Based on the non-linear regression calculated in E, growth rates were corrected to account for organoid core death (detailed equations in Methods section). Blue line = non-linear regressions. Pink line = rolling average. Band = 95% CIs of regressions. n = 522 organoids from 8 mice
To extend this observation, we measured cell proliferation and cell death in individual organoids embedded in 3D Matrigel. First, we measured mitotic rate in organoids by staining for nuclear mitosis marker phospho-Histone H3. Consistent with our outgrowth estimates, we observed that intermediate-sized organoids had the highest proportion of cells in mitosis (Fig. 1d). Next, we treated organoids after 4 days in 3D Matrigel with cell-impermeable propidium iodide to assess organoid viability. We observed that very large organoids frequently had extensive non-viable regions within the organoid core (Fig. 1e). No observed organoids below 10,000 um2 had such non-viable core regions.
Taken together, these data indicate that organoid expansion is highly nonlinear and dependent on initial organoid size. Single MMTV-PyMT cells rarely grow in 3D culture, and frequently undergo cell death. Intermediate sized organoids have increased mitotic rates, increased viability, and strong outgrowth in 3D culture. Very large organoids have decreased mitotic rates, increased core cell death, and poorer overall growth rates. To translate starting and final organoid areas from Fig. 1b into approximate starting and ending cell numbers we empirically determined the average organoid cell area, accounted for non-viable cores formation as reported in Fig. 1e, and generated a non-linear regression model (see Methods for details). Based on these calculations, we estimate that the starting size of organoids that generates maximal growth is between ~10 cells (95% CI 10.5–12.9) to ~1000 cells (95% CI 1029.3–1268.2) per organoid (Fig. 1f).
Suspension Culture Supplemented with Basement Membrane-Rich Extract Produces Optimal Organoid Outgrowth
Organoids are commonly cultivated fully surrounded by a 3D matrix, most often encapsulated within a droplet of basement membrane extract such as Matrigel [16, 45, 46]. In these systems, organoids are typically split by gentle enzymatic digestion or by trituration, then re-embedded in fresh 3D matrix. We reasoned that this conventional approach was likely suboptimal because of our finding that intermediately sized organoids have maximal growth potential. Importantly, under-digestion during traditional organoid passaging produces overly large organoids with low growth rates and which are prone to form non-viable cores. Additionally, over-digestion produces many single cells and very small (<<10 cell) clusters prone to cell death and growth arrest, respectively (Fig. 2a: left panel). Because the optimal level of digestion depends on many factors including organoid size, amount of 3D matrix, and cultivation time, the size of digested organoids is difficult to control and to keep consistent from passage to passage.
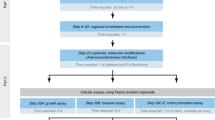
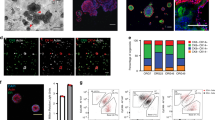
Suspension culture supplemented with basement membrane-rich extract facilitates rapid expansion of tumor organoids. a Schematic of organoid culture methods. Traditionally, organoids are either plated intact in 3D gels or briefly mechanically or enzymatically disrupted to generate organoid fragments, as well as debris and single cells, before plating in 3D culture. Right, we present an alternative culture method where single tumor cells are aggregated in non-adherent suspension supplemented with basement membrane-rich extract to form multicellular aggregates optimally sized for rapid outgrowth. b DIC images (left) and fold growth by viable cell number (right), of MMTV-PyMT organoids after 4 days in organoid media only, media supplemented with 2% Matrigel, media +100 μg/mL laminin/entactin, or media +70 μg/mL laminin/entactin +30 μg/mL collagen IV. Each dot is a biological replicate. P values = unpaired t-tests. c To determine the optimal density to seed MMTV-PyMT single cells for aggregation in 2% Matrigel suspension culture, cells were plated at 0.5 × 105 to 2.5 × 105 viable cells/mL to form small clusters. After 4 days, fold-growth by viable cell number was measured. n = 4 mice, each line is a biological replicate. P values = paired t-tests. d Left, Keratin-14 immunofluorescence images of MMTV-PyMT organoids cultured embedded in 100% Matrigel vs. cultured in suspension supplemented with 2% Matrigel. Right, quantification of the % of cells K14+ (basal). n = 2 mice (suspension), n = 3 mice (embedded), n = 4161 cells analyzed from 34 organoids. P-value = unpaired t-test. e Images of organoids after 4 days in culture embedded in 100% Matrigel or seeded in suspension culture supplemented with 2% Matrigel. Right, summary of organoid morphology at D4. “Rounded” refers to organoids with smooth spherical borders, vs. “budded” organoids with multicellular budded protrusions. No organoids scored had hollow, cystic morphology in either condition. n = 3 mice, n = 285 organoids in suspension, 259 organoids embedded in 100% Matrigel. P-value = unpaired t-test. f DIC images of MMTV-PyMT organoids 0 days, 1 day, and 4 days after initial plating in the methods described in (a). Whole organoids or briefly dissociated (5 minutes Accumax treatment) organoids were plated in 3D matrigel, or single cells were allowed to aggregated at 150,000 viable cells/mL in organoid media +2% Matrigel. g From the area measurements of Fig. 2f (see Table 2) and the growth parameters identified in Fig. 1f, estimated fold growth for each culture method was predicted (see Methods section for piecewise function). Then actual fold growth of MMTV-PyMT organoids by viable cell number after 4 days in each of the 3 culture methods was measured. Each dot is a mouse. P-values = unpaired t-tests
We hypothesized that a suspension culture system could more optimally generate organoids in the ideal intermediate size range for optimal outgrowth (Fig. 2a: right panel). In this system, we dissociate organoids to single cells and then reaggregate them in suspension at specific cell densities to form intermediate sized clusters (Fig. 1f, ~10–1000 cells). By digesting completely to single cells, the degree of digestion and the concentration of cells can be highly controlled from passage to passage, while allowing for reaggregation in suspension to reform clusters. To optimize our suspension protocol, we first determined whether basement membrane matrix was required for outgrowth (Fig. 2b). The extracellular matrix surrounding tumor cells fulfills two key roles: supplying signaling molecules such as integrin ligands and growth factors, and providing a physical framework that supports and exerts force on cells [28]. 98% of the protein content of Matrigel is composed of laminin, collagen IV, and entactin [2]. To determine whether these components were sufficient to induce organoid growth, we tested outgrowth of MMTV-PyMT organoids in organoid media only; media supplemented with 2% Matrigel, which remains liquid; or media supplemented with recombinant laminin, entactin, and collagen IV. Outgrowth was highest using media +2% Matrigel compared with all other conditions (Fig. 2b). These findings indicate that (1) matrix structure provided by a rigid polymerized gel is dispensable for robust outgrowth, (2) the presence of additional components in Matrigel, beyond laminin/entactin/collagen IV, are required for organoid expansion, and that (3) a 2% concentration of Matrigel provides the necessary signals for outgrowth and allows easy sample manipulation as the media remains completely fluid.
Having devised a suspension culture system which could support high organoid outgrowth, we next sought to identify the ideal seeding density that would maximize growth by forming optimally sized aggregates. We tested cell seeding at 0.5 × 105 to 2.5 × 105 viable single cells per mL in non-adherent 6 well plates. 150,000 viable cells/mL consistently delivered the highest outgrowth in matched organoid cultures from the same mice (Fig. 2c). Organoids in either suspension or 100% Matrigel had similar proportions of cells expressing Keratin-14 (Fig. 2d), confirming that this suspension culture method does not interrupt the formation of K14+ basal cells in tumor organoids. No significant differences in morphology were observed in organoids cultivated in suspension vs. 100% Matrigel (Fig. 2e), using previously published morphologic criteria for organoids [25]. In both conditions most organoids formed dense, budded structures and no hollow, cystic structures were observed. Seeding at 150,000 viable cells/mL also resulted in robust growth of normal mouse mammary organoids as well as organoids from the MMTV-Neu and C3(1)TAg mouse models of breast cancer (Table 1). In summary, we find that aggregating tumor organoid cells at 150,000 viable cells/mL in media supplemented with 2% Matrigel generates the optimal rate of outgrowth per passage.
Finally, we sought to compare our suspension culture method head-to-head with traditional embedded 3D culture methods. MMTV-PyMT organoids were either plated in 100% Matrigel droplets, briefly enzymatically dissociated and then plated in 100% Matrigel droplets, or seeded at 150,000 viable cells/mL in suspension +2% Matrigel (Fig. 2f). DIC images were taken of each culture at day 1 and the sizes of tumor organoids, tumor single cells, and suspension aggregates were measured (Table 2). Overall, we observed that the vast majority of whole organoids seeded in 100% Matrigel were above the threshold we predict would yield optimal growth (10–1000 cells). Brief enzymatic dissociation improved the size distribution of organoids embedded in 3D Matrigel, resulting in a higher proportion of cells in optimally sized clusters (Table 2). However, many large organoids which had shed outer layers of cells remained >1000 cells in size. We also observed a large number of single cells shed from organoids after brief enzymatic digestion. Over time, some of these single cells merged with adjacent neighbors to form small clusters, however many remained as single cells and failed to undergo substantial growth (Fig. 2f). When plated using our suspension method at 150,000 viable cells/mL, MMTV-PyMT cells formed many small, optimally sized clusters within 1 day of plating which underwent strong outgrowth over 4 days in culture (Table 2, Fig. 2f).
Using the growth rate prediction model shown in Fig. 1f, we calculated the predicted change in cell number per passage for each of these methods based on organoid area measurements (Table 2). Based on these predictions we estimated the fold-growth per passage by viable cell number to be approximately 4.3-fold, 6.1-fold, and 13.0-fold for whole organoids in 3D Matrigel, partly dissociated organoids in 3D Matrigel, and our suspension culture method, respectively (Fig. 2g). Next, we assessed the actual fold-growth per passage in each culture method using matched organoids from the same mice. The median fold-growth per passage was 1.3-fold, 4.6-fold, and 13.0-fold (by viable cell number) for the whole embedded, partly dissociated, and suspension culture methods, respectively (Fig. 2g). MMTV-PyMT organoid lines consistently generated strong fold-growth in suspension culture after passaging. Individual growth rates varied between lines generated from different mice (Table 1), consistent with heterogeneity of spontaneous tumor initiation and outgrowth of this model in vivo. This variation in growth is an important consideration when planning experiments. We recommend that when possible organoid lines be compared between genetic perturbations, for instance control and shRNA or sgRNA transduced, and that comparisons between organoid lines should take into account differences in intrinsic growth rates. Overall, these real-world data fairly closely match our predictions based on the growth parameters identified in Fig. 1f, and confirm that our suspension culture method supports more optimal growth than traditional 3D culture techniques.
The Suspension Organoid Culture Method Facilitates Large Scale Organoid Expansion and Efficient Lentiviral Transduction
It is relatively simple and inexpensive to expand millions or tens of millions of cells from immortal cancer cell lines in 2D culture. In comparison, when culturing organoids in 3D basement membrane-rich gels it can be tedious and expensive to generate more than a few million viable cells as starting inputs. As a result, commonly used assays that work best with large numbers of cells such as immunoblots, biochemical assays, and medium to high-throughput screens are much more challenging to perform using organoids.
We realized that our suspension culture system had several useful properties supportive of large-scale expansion for such assays. First, our system maintains organoids in suspension. This avoids the requirement to liberate organoids from a polymerized gel, making passaging and sample collection far less time-consuming. As a result, organoid culture could potentially be scaled up simply by increasing the volume of media present. Second, our system expands organoids more rapidly than fully embedded systems, reducing the time to achieve sufficient numbers of cells. Third, our system uses 2% Matrigel, which significantly reduces the cost of basement membrane extract during large-scale experiments.
As a demonstration of the scaling capability of this method, we cultured over 50 mL of organoids using cell culture flasks coated with 1% agarose to generate a non-adherent surface. Using this suspension +2% Matrigel culture method in non-adherent cell culture flasks, we expanded 1 million organoid cells to over 100 million cells in 2 weeks (Fig. 3a). To expand 1 million starting cells to >150 million, the number that has been reported to achieve sufficient coverage for a genome-wide CRISPR screen [27], the estimated cost is much more expensive and much more labor-intensive using traditional 3D culture compared to this suspension culture method (Table 3).
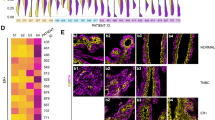
Suspension culture supplemented with basement membrane-rich extract facilitates stable long-term culture and efficient lentiviral transduction for in vivo metastasis studies. a Using the 2% Matrigel suspension culture method, a starting input of 1 million tumor organoid cells (n = 3 mice, each line is a biological replicate, each dot is a passage) were expanded and passaged for 2 weeks. Right, image of >300 million MMTV-PyMT cells expanded from 1 million initial cells after 2 weeks. b Percent viability and mean fold growth by viable cell number of MMTV-PyMT cells at passage number 2, 5, or 7 in 2% Matrigel suspension culture. n = 4 mice. No significant differences were detected between passages for growth or viability (unpaired t-tests, p > 0.05). c Immunofluorescence for basal cell marker Keratin-14 in MMTV-PyMT organoids freshly isolated from primary tumors (passage 0) or after 4 passages in 2% Matrigel suspension culture. Representative images from n = 2 mice. d Log2 tumor/normal copy number variations from low coverage whole genome sequencing (plotted as 100 kilobase bins) of MMTV-PyMT organoids from the same mouse shortly after isolation (passage 0) or after 10 passages in 2% Matrigel suspension culture. e Heatmap of passaged log2 copy number ratios (tumor/normal) minus passage 0 log2 copy number ratios (mean per chromosome). f Passaged log2 copy number ratios (tumor/normal) minus passage 0 log2 copy number ratios (mean per chromosome). Each dot is a chromosome, n = 4 mice, n = 6 passages. Red line = median. g Left, schematic of lentiviral transduction of organoids in suspension. MMTV-PyMT tumor organoids were lentivirally transduced with PGK-EGFP at MOI ~10 in suspension. Then tumor organoids were lentivirally transduced with non-targeting (sgCTRL) or EGFP targeting (sgEGFP) lentiCRISPRv2 constructs. Right, DIC and GFP images after 10 days of puromycin selection showing effective EGFP knockout. h MMTV-PyMT organoids were cultured in suspension +2% Matrigel and transduced with a GFP encoding lentivirus as in G. After puromycin selection, PyMT-GFP clusters were orthotopically transplanted into the mammary fat pads of NSG mice. Tumors were collected 6 weeks later. n = 10 mice, n = 20 tumors. i MMTV-PyMT organoids were cultured in suspension +2% Matrigel and transduced with GFP encoding lentivirus as in G. After puromycin selection, PyMT-GFP clusters were injected by tail vein into NSG mice. 3 weeks later lungs were harvested to assess metastatic outgrowth via stereomicroscopy. n = 8 mice
To expand such large numbers of cells required at least 2 passages, and we anticipated that this protocol would be most useful if organoids could be repeatedly passaged to maintain laboratory stock cultures. Consequently, we sought to determine whether repeated passaging negatively affected important organoid features and phenotypes. We assessed viability and fold growth of matched MMTV-PyMT organoid lines after 2, 5, or 7 passages in 2% Matrigel suspension culture. No significant changes in either viability or fold growth were observed (Fig. 3b). The longest we cultivated organoid lines was 10 passages (approximately 30 population doublings); up to passage 10 organoid lines did not reach growth arrest or experience lower fold-growth per passage than earlier passages (n = 2).
Next, we assessed cell lineage; breast cancer tumors contain Keratin-14 positive basal cells, generally enriched on the organoid periphery, as well as Keratin-8 positive luminal cells [6]. After repeated passaging in suspension, organoids maintained similar levels and distributions of K14 protein expression (Fig. 3c). Lastly, we assessed genomic stability, as long periods of growth and passaging can also introduce genomic alterations [39]. Copy number variations were assessed using low coverage whole genome sequencing, relative to a normal mouse control, in unpassaged and passaged PyMT organoids. We observed only modest changes in copy number variations after passaging (chromosome-level log2 copy # ratio range vs. unpassaged = −0.36 to 0.2, 100 kb bin-level log2 copy # ratio range vs. unpassaged = −1.16 to 1.48), and no chromosome-level duplications or deletions were identified (Fig. 3d–f). Though it is likely that passaging will alter organoids at least to some degree, these results suggest that key properties of MMTV-PyMT organoids, including genotypic stability and basal/luminal lineage specification, remain consistent after repeated passaging in suspension.
Having generated a protocol to rapidly expand very large numbers of organoids, we next sought to test the utility of this protocol for lentiviral transduction, a useful technique which is more challenging in traditional 3D culture [42]. We previously developed magnetic particle transduction methods for organoids as a means to enhance transduction efficiency in limited organoid samples [6, 47]. However, an optimal organoid expansion protocol enables a more cost-efficient transduction method for large numbers of organoids. Lentiviral particles were generated in 293FT cells and concentrated to 1/100th the starting lentiviral supernatant volume. To maximize cell-virus contact, MMTV-PyMT organoids were first dissociated to single cells before plating in suspension +2% Matrigel. Protamine sulfate was then added to the media, a polycation which greatly increases lentiviral transduction efficiency [3, 8]. Using this method, we were able to generate a highly EGFP expressing MMTV-PyMT organoid line (Fig. 3g). Next, we sought to confirm that CRISPR-Cas9 could be used in this system for effective gene targeting. The MMTV-PyMT-EGFP organoids were transduced again, using the same protocol, with either non-targeting or EGFP targeting sgRNAs. Organoids were maintained in media +2% Matrigel with puromycin for 10 days to select for transduced cells and permit enough time for maximal Cas9 activity [54]. After selection, we observed strong EGFP signal reduction only in the sgEGFP transduced condition (Fig. 3g). This experiment provides a proof-of-concept that MMTV-PyMT organoids in long-term suspension culture can be used to generate knockout, knockdown, or overexpression cultures through lentiviral transduction [14, 27]. In summary, we present an optimized protocol for rapid mouse organoid expansion which can facilitate large-scale experiments, such as genome wide lentivirus-based screens and proteomic studies.
Lastly, we sought to confirm that organoids cultivated, passaged, transduced, and selected in 2% suspension culture retained the ability to form primary tumors and metastases in vivo. MMTV-PyMT organoids were generated, passaged in 2% Matrigel suspension, and transduced with a GFP-encoding lentiviral construct. Cells were selected in puromycin for at least 5 days. Selected organoids were then dissociated to single cells and allowed to reaggregate overnight in 2% Matrigel suspension to form small clusters. The following day, reaggregated clusters were orthotopically transplanted into the mammary fat pads of immunocompromised NSG mice which were collected 6 weeks later when they had formed large tumors. We observed 100% take rates (n = 10/10 mice) and tumor sections were confirmed to be GFP+ (Fig. 3h). As a second approach, reaggregated clusters were injected by tail vein into immunocompromised NSG mice, and lungs were collected 3 weeks later. Likewise, we observed 100% take rates (n = 8/8 mice) and numerous lung metastases that were GFP+ by stereomicroscopy (Fig. 3i). We conclude that our propagation method can be used to prepare and transduce large numbers of organoids prior to in vivo experiments, facilitating validation and phenotypic analysis of sgRNA, shRNA, or overexpression organoid lines in pre-clinical models. Together, these methods form an efficient pipeline for large-scale cultivation, ex vivo manipulation, and in vivo analysis of tumor organoids.
Conclusion
Here, we describe a protocol for the rapid cultivation of large numbers of mouse mammary tumor organoids. We show that organoid size is predictive of organoid outgrowth and that maximal outgrowth occurs at an intermediate size corresponding to between ~10 to 1000 cells per organoid. As a result, passaging organoids via partial digestion to generate fragments yields suboptimal outgrowth. To overcome this challenge, we developed a suspension culture protocol in which organoids are fully dissociated to single cells, and then subsequently aggregated to form organoids in the ideal intermediate size range. This protocol now enables the large-scale and rapid outgrowth of mouse mammary tumor organoids for a variety of assays requiring large cell numbers. Although not directly tested here, we also expect that the size vs. outgrowth relationships described may occur in normal organoids, which could be useful for regenerative applications, or during the expansion of fresh or cryopreserved tumor organoids from other mouse models of cancer. We suggest that researchers interested in testing this culture system in their preferred organoid model should compare fold growth, viability, cellular heterogeneity, and genomic stability in traditional 3D culture vs. 2% Matrigel suspension using the techniques we have applied to freshly isolated MMTV-PyMT tumor organoids.
This culture system offers a number of logistical advantages over traditional embedded 3D culture, greatly reducing financial cost and hands-on time for large-scale experiments. It also diverges from other breast cancer suspension culture techniques in several important ways. Mammosphere or tumorsphere assays have also been used to cultivate breast cancer cells in suspension. In these assays, cells are plated as single cells and prevented from re-aggregating, often by addition of methylcellulose to the media [33, 56]. Multiple growth factors are typically needed to coax the clonal outgrowth of spheres of cells. While these conditions are ideal to test the presence of rare cancer stem cell or progenitor cells, our data suggest that this approach will also result in high dropout of individual clones. In contrast, our method encourages aggregation to induce the optimal level of multicellularity supporting organoid outgrowth.
We also acknowledge some limitations to this method. Firstly, the cultivation of tumor organoids in suspension is non-organotypic. Tumor cells do not typically exist in fluid-based environments in vivo, apart from tumor cells in the circulation or in bodily fluids like pleural effusions and malignant ascites. Suspension culture lacks the rigid physical structure of fully-embedded systems, which could modulate mechano-transduction and other signaling pathways relevant for cancer [17, 41], as well as affect hypoxic gradients that could induce EMT and metabolic phenotypes relevant for cancer progression [34, 40, 48]. We therefore suggest using this method primarily for expansion. For organoid analyses, we recommend transferring expanded organoids into fully embedded 3D matrices or animal models to better recapitulate the tumor microenvironment. Secondly, the introduction of additional mutations over time can cause confounding effects on experiments, particularly those assessing the phenotype of shRNAs or sgRNAs. MMTV-PyMT tumors have relatively high genomic stability [49], making organoids from this model ideal for large-scale, long-term propagation. We encourage regular genetic analysis when applying this long-term cultivation method to other organoid models with higher genomic instability. Thirdly, this method is applicable at this time only to organoids isolated from mice. It is well known that human organoids grow significantly slower than mouse organoids. We do not resolve that problem here but suggest this to be an essential direction of future research.
In this study, we show that the size of tumor cell communities can have large non-linear effects on proliferation and cell death. Collective cellular organization is known to promote invasive and metastatic potential in diverse tumor types [1, 5, 7, 18, 21, 23, 29, 38, 46, 61]. An interesting question is whether the growth kinetics observed here as a function of organoid size in 3D culture might also have relevance for the outgrowth of microscopic and macroscopic metastases in vivo, which are known to display different properties at different size regimes [7, 9, 31, 52]. Thus, our study advocates for further systematic analyses of the factors supporting collective organization, signaling, and outgrowth in tumor cells.
Methods
Mouse Mammary Tumor Organoid Culture
Animal protocols were approved by the Fred Hutchinson Institutional Animal Care and Use Committee. FVB/N-Tg(MMTV-PyVT)634Mul/J (MMTV-PyMT) were maintained and tumor growth was monitored every 2 days. Adult female mice were used for all experiments.
In this study, we isolated intact epithelial fragments from tumors to generate freshly derived tumor tissue organoids [47]. A variety of growth factors and ligands can be used in this culture setting to promote growth, but a simple media composed of DMEM/F12, ITS-X, and 2.5 nM FGF is frequently used to maintain both normal mammary and tumor organoids [6, 45, 47, 61]. However other supplements can be included to promote growth or alter the cellular composition of organoids [26], particularly when culturing human tumor organoids which often grow poorly ex vivo [50, 51].
Organoids were isolated from MMTV-PyMT, C3(1)TAg, or MMTV-Neu mouse mammary tumors as previously described [45]. Mammary tumors were dissected, mechanically disrupted with a scalpel, and then digested in a collagenase-trypsin solution for 30–60 min shaking at an angle at 100–150 rpm at 37 °C. The digestion solution (in 20 mL of DMEM/F12) contained 2 mg/mL collagenase (Sigma C2139), 2 mg/mL trypsin (Gibco 27,250–018), 5% fetal bovine serum, 5 μg/mL human insulin (Sigma-Aldrich I9278), 50 μg/mL gentamicin (Gibco 15,750–060). Tumor fragments were centrifuged for 10 min at 1500 rpm, resuspended in 4 mL DMEM/F12 and treated with 40 μL (2000 U/mL) of DNAse (Sigma D4263) for 3 min then pelleted and resuspended in 10 mL DMEM/F12. The solution was centrifuged for 4 s at 1500 rpm (453 g) to isolate multicellular organoids, then resuspended in 10 mL DMEM/F12. This was repeated for a total of 4 washes to remove contaminating single cells and debris, leaving behind primarily multicellular epithelial organoids [45, 47]. Mice were harvested as the largest tumor neared 1.5 cm in diameter.
For 3D culture, 100–200 clusters were embedded in 100 μL of growth-factor reduced Matrigel (Corning 354,230), the Matrigel was allowed to polymerize for 30–60 min at 37 °C, then 1 mL of organoid media (DMEM-F12, 2.5 nM FGF2, insulin-transferrin selenium, & penicillin/streptomycin) was added. For recombinant ECM media supplementation, laminin/entactin (Corning 354259) and collagen IV (Corning 354233) were used. For suspension culture, clusters were cultured in non-adherent 6 well plates (Fisher Scientific 07–200-601) in 2–4 mL organoid media +2% (v/v) Corning Growth Factor Reduced Matrigel. Tips and tubes used to handle organoids were first coated in 2.5% bovine serum albumin (Sigma-Aldrich A9576) in DPBS to prevent loss of material. To passage cells in 2% Matrigel suspension, organoids were pelleted and resuspended in Accumax (Innovative Cell Technologies, AM105) for 20 min in a 37 °C water bath, pipetted every 10 min with a BSA coated pipette to generate a single cell suspension. Cells were counted with Trypan blue on a hemocytometer to ensure high viability and a low number of residual clusters. Cells were pelleted and resuspended in warm OGM + 2% Matrigel at 150,000 viable cells/mL and plated in non-adherent plates or flasks. Matrigel was added to the media at 4 °C, mixed thoroughly, then warmed at 37 °C for at least 10 min before organoid resuspension.
For large-scale suspension culture, T175 flasks were coated in 1% agarose (dissolved in boiling Milli-Q filtered water and then cooled to ~50 °C before pouring along bottom of flask until ~3 mm thick) to prevent cell adhesion. MMTV-PyMT cells were plated at 150,000 viable cells/mL in 75 mL OGM + 2% Matrigel in T-175 flasks coated with 1% agarose and passaged every 4–6 days back to 150,000 viable cells/mL at the desired volume.
Fresh organoid preparations were typically >70% viable. Organoids passaged in 2% Matrigel supplemented suspension maintained higher viability (~80–95%, Fig. 3b).Taking into account single cells that fail to aggregate and small clusters, both of which are more likely to apoptose, we estimate a drop-out rate of <10% after initial passage in suspension culture (Table 2).
Immunofluorescence and Non-viable Cell Labeling
Organoids were fixed with 4% paraformaldehyde in DPBS (10 min), permeabilized 30 min with 0.5% Triton-X, and blocked 1 h at room temperature with 10% FBS/1% BSA/0.1% Triton-X in DPBS. Primary antibodies were added in block solution and incubated at 4 °C overnight. Alexa Fluor conjugated secondary antibodies (1:200) were incubated for 2–3 h at room temperature with 5% host serum. Confocal images were acquired using an Andor CSU-W confocal spinning disk on a Leica DMi8 inverted microscope. Antibodies used: phospho-Histone H3 Ser10 (CST 9701) and Keratin-14 (BioLegend 905301). For live imaging of non-viable cells, propidium iodide (Thermo Scientific P1304MP) was added at 1 μg/mL to the organoid media. At least 15 min later, DsRed and DIC images were acquired. To assess K14-/K14+ cell populations in suspension culture vs. embedded 100% Matrigel culture, organoids cultured in 2% Matrigel suspension for at least 48 h were embedded in 100% Matrigel and fixed immediately for immunofluorescence. K14 scoring was blinded, DAPI+ cells with at least 50% membrane positivity were scored as K14 positive.
Time-Lapse Imaging of Organoids
Single or clustered cells were plated in growth factor reduced basement membrane-rich gels. Differential interference contrast (DIC) images were captured hourly at 10X magnification. Exposure times were < 30 ms for BF/DIC. Temperature was maintained at 37 °C and CO2 at 5%. Images were acquired using a Leica DMi8 TCS SPE. To assess organoid growth, the μm2 area of organoids was assessed at 0 h and 108 h of time lapse imaging in FIJI software.
Lentivirus Production and Transduction
Lentivirus was produced in HEK293FT cells using PsPax (Addgene #12260) and MD2.G (Addgene #12259) packaging plasmids. Supernatants were concentrated using Lenti-X (Takara 631232), resuspended in 1/100th the supernatant volume of PBS, and frozen at -80 °C. Viruses were titered using a ZeptoMetrix p24 ELISA (Fisher Scientific 22–156-700). Tumor organoids were dissociated to single cells in Accumax and resuspended at 150,000 viable cells/mL in organoid media +2% (v/v) Matrigel. Protamine sulfate (Fisher ICN15197105) was added at 8 μg/mL to enhance transduction efficiency before adding concentrated lentivirus. For plasmids with PuroR, 2 days after transduction puromycin was included in the media at 1–2 μg/mL for selection. CRISPR sgRNA transduced cells were kept in puromycin for 10 days before assessing sgRNA efficacy.
Low-Coverage Whole Genome Sequencing
MMTV-PyMT organoids were flash frozen in liquid nitrogen at specific passage numbers (unpassaged organoids = passage 0). MMTV-PyMT tumor organoid genomic DNA was collected using the QIAGEN DNEasy Blood & Tissue Kit (69504), following manufacturer’s instructions. For whole genome analysis of CNV variation, we performed low-coverage whole-genome sequencing of organoid genomic DNA. The NEBNext DNA Library Prep Master Mix Set for Illumina (New England BioLabs; catalog E6040L) was used to generate DNA libraries from 1000 ng genomic DNA. Single-read data (50 bp) were generated using an Illumina HiSeq 2500. Reads were aligned to the mm9 build of the murine genome using the Burrows Wheeler Aligner [35], and we used CNVseq [63] to examine copy numbers.
Orthotopic Transplantation of PyMT-GFP Clusters
MMTV-PyMT-mTomato organoids were transduced with a GFP encoding lentivirus (Transomic NT4) and selected with 1 μg/mL puromycin for at least 5 days. Organoids were then dissociated to single cells using Accumax, and reaggregated overnight at 250,000 viable cells/mL in organoid media +2% Matrigel (using non-adherent 6 well plates). Clusters were then resuspended in 50% Matrigel in DMEM/F12 (vol/vol) on ice for mammary fat pad injection. 9–10 week old Nod scid gamma (NSG) immunocompromised mice were anesthetized with 2.5% isoflurane and the surgical site was sterilized with ethanol and chlorhexidine. A 1 cm midline incision was made, allowing the #4 mammary fat pad to be exposed. 50,000 clustered MMTV-PyMT-GFP cells per gland were injected into the left and right #4 mammary glands. The surgical area was locally infiltrated with 0.25% bupivacaine for pain relief. Surgical wounds were closed with 9 mm autoclips and tissue glue. Triple antibiotic ointment was applied to the incision. Mice were monitored closely with autoclip removal two weeks after surgery. Tumors were measured every 2 days, and mice were sacrificed before tumors reached 15 mm in diameter. At endpoint, mice were euthanized and primary tumors were fixed in 4% PFA for 4 h, then transferred to 25% sucrose in DPBS overnight at 4 °C before embedding in OCT and storing at -80 °C. 50 μm sections stained with DAPI were used to assess retention of GFP expression in MMTV-PyMT tumors.
Tail Vein Injection of PyMT-GFP Clusters
MMTV-PyMT-mTomato organoids were transduced with a GFP encoding lentivirus (Transomic NT4) and selected with 1 μg/mL puromycin for at least 5 days. Organoids were then dissociated to single cells using Accumax, and reaggregated overnight at 150,000 viable cells/mL in media +2% Matrigel (v/v). The following day, after cells had reaggregated, 200,000 viable clustered cells per mouse were injected by tail vein into Nod scid gamma (NSG) immunocompromised mice in 200 μL of PBS using a 27 g needle. 3 weeks after tail vein injection, mice were euthanized and lungs imaged under a dissecting microscope to assess GFP+ PyMT metastatic outgrowth.
Piecewise Function for Estimating Organoid Growth Rate from Initial Cell Number
FIJI software was used to measure the μm2 area of cells or clusters to measure organoid growth rate, defined as [(final organoid volume)-(initial organoid volume)/(initial organoid volume)]. To estimate organoid volume, an approximate radius was calculated from organoid area (A = πr2). Then, approximate organoid volume was calculated (V = (4/3)πr3). From immunofluorescence, we estimated the average MMTV-PyMT cell radius as ~4.92 μm (95% CI: 4.66–5.00). To convert organoid volume to approximate cell number, volume was divided by 498.2 μm3 (volume of one cell with radius 4.92 μm). Growth rate by volume ([final organoid volume-initial organoid volume]/initial organoid volume) was calculated for each organoid to approximate growth rate by cell number. Based on measurements in Fig. 1f, we generated a non-linear regression to estimate the proportion of cells in non-viable organoid cores. Growth rates were adjusted to exclude the predicted number of cells in non-viable cores. A simple linear regression was used to model the growth rates of 1–10 cell organoids (usually growth arrested). For organoids >10 cells, we generated non-linear regression to predict final growth rate. Equations used to model growth summarized below:
Predicted proportion outside non-viable core
Cell # growth rate = \( \frac{\left(\mathrm{final}\ \mathrm{cell}\#\mathrm{corrected}\ \mathrm{for}\ \mathrm{viability}\right)-\left(\mathrm{initial}\ \mathrm{cell}\#\mathrm{corrected}\ \mathrm{for}\ \mathrm{viability}\right)}{\mathrm{initial}\ \mathrm{cell}\#\mathrm{corrected}\ \mathrm{for}\ \mathrm{viability}} \)
Predicted cell number growth rate based on initial organoid size
Simple linear regression for x < 10, 4-part nonlinear regression (variable slope) for x ≥ 10
Statistical Analysis
Graphs, non-linear regressions, and statistical tests were generated using GraphPad Prism 8. Red lines denote medians, unless otherwise noted. Experiments conducted on different days or using samples from different mice were considered biological replicates. All statistical tests are two-sided. p ≤ 0.05 was considered significant. P-values: “ns” p > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
References
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS, Spencer JA, et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell. 2014;158:1110–22.
Aisenbrey EA, Murphy WL. Synthetic alternatives to Matrigel. Nat Rev Mater 2020;5: 539–551. https://doi.org/10.1038/s41578-020-0199-8
Balak JRA, de Graaf N, Zaldumbide A, Rabelink TJ, Hoeben RC, de Koning EJP, et al. Highly efficient ex vivo lentiviral transduction of primary human pancreatic exocrine cells. Sci Rep. 2019;9:15870.
Cattaneo CM, Dijkstra KK, Fanchi LF, Kelderman S, Kaing S, van Rooij N, et al. Tumor organoid–T-cell coculture systems. Nat Protoc. 2020;15:15–39.
Cheung KJ, Ewald AJ. A collective route to metastasis: seeding by tumor cell clusters. Science (New York, NY). 2016;352:167–9.
Cheung KJ, Gabrielson E, Werb Z, Ewald AJ. Collective invasion in breast cancer requires a conserved basal epithelial program. Cell. 2013;155:1639–51.
Cheung KJ, Padmanaban V, Silvestri V, Schipper K, Cohen JD, Fairchild AN, et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc Natl Acad Sci U S A. 2016;113:E854–63.
Cornetta K, Anderson WF. Protamine sulfate as an effective alternative to polybrene in retroviral-mediated gene-transfer: implications for human gene therapy. J Virol Methods. 1989;23:187–94.
Davis RT, Blake K, Ma D, Gabra MBI, Hernandez GA, Phung AT, et al. Transcriptional diversity and bioenergetic shift in human breast cancer metastasis revealed by single-cell RNA sequencing. Nat Cell Biol. 2020;22:310–20.
de Witte CJ, Espejo Valle-Inclan J, Hami N, Lõhmussaar K, Kopper O, Vreuls CPH, et al. Patient-derived ovarian cancer organoids mimic clinical response and exhibit heterogeneous inter- and intrapatient drug responses. Cell Rep. 2020;31:107762.
Debnath J, Brugge JS. Modelling glandular epithelial cancers in three-dimensional cultures. Nat Rev Cancer. 2005;5:675–88.
Debnath J, Muthuswamy SK, Brugge JS. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods. 2003;30:256–68.
Diermeier SD, Chang KC, Freier SM, Song J, El Demerdash O, Krasnitz A, et al. Mammary tumor-associated RNAs impact tumor cell proliferation, invasion, and migration. Cell Rep. 2016;17:261–74.
Dominguez AA, Lim WA, Qi LS. Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nat Rev Mol Cell Biol. 2016;17:5–15.
Drost J, Clevers H. Organoids in cancer research. Nat Rev Cancer. 2018;18:407–18.
Drost J, Karthaus WR, Gao D, Driehuis E, Sawyers CL, Chen Y, et al. Organoid culture systems for prostate epithelial and cancer tissue. Nat Protoc. 2016;11:347–58.
Fattet L, Jung H-Y, Matsumoto MW, Aubol BE, Kumar A, Adams JA, et al. Matrix rigidity controls epithelial-mesenchymal plasticity and tumor metastasis via a mechanoresponsive EPHA2/LYN complex. Dev Cell. 2020;54:302–316.e7.
Friedl P, Gilmour D. Collective cell migration in morphogenesis, regeneration and cancer. Nat Rev Mol Cell Biol. 2009;10:445–57.
Ghatak S, Dolatabadi N, Trudler D, Zhang X, Wu Y, Mohata M, et al. Mechanisms of hyperexcitability in Alzheimer’s disease hiPSC-derived neurons and cerebral organoids vs isogenic controls. eLife. 2019;8:e50333.
Giandomenico SL, Mierau SB, Gibbons GM, Wenger LMD, Masullo L, Sit T, et al. Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat Neurosci. 2019;22:669–79.
Gkountela S, Castro-Giner F, Szczerba BM, Vetter M, Landin J, Scherrer R, et al. Circulating tumor cell clustering shapes DNA methylation to enable metastasis seeding. Cell. 2019;176:98–112.e114.
Guy CT, Cardiff RD, Muller WJ. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol Cell Biol. 1992;12:954–61.
Haeger A, Alexander S, Vullings M, Kaiser FMP, Veelken C, Flucke U, et al. Collective cancer invasion forms an integrin-dependent radioresistant niche. J Exp Med. 2020;217.
Hubert CG, Rivera M, Spangler LC, Wu Q, Mack SC, Prager BC, et al. A three-dimensional organoid culture system derived from human glioblastomas recapitulates the hypoxic gradients and cancer stem cell heterogeneity of tumors found in vivo. Cancer Res. 2016;76:2465–77.
Jamieson PR, Dekkers JF, Rios AC, Fu NY, Lindeman GJ, Visvader JE. Derivation of a robust mouse mammary organoid system for studying tissue dynamics. Development. 2017;144:1065–71.
Jardé T, Lloyd-Lewis B, Thomas M, Kendrick H, Melchor L, Bougaret L, et al. Wnt and Neuregulin1/ErbB signalling extends 3D culture of hormone responsive mammary organoids. Nat Commun. 2016;7:13207.
Joung J, Konermann S, Gootenberg JS, Abudayyeh OO, Platt RJ, Brigham MD, et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat Protoc. 2017;12:828–63.
Kai F, Drain AP, Weaver VM. The extracellular matrix modulates the metastatic journey. Dev Cell. 2019;49:332–46.
Labuschagne CF, Cheung EC, Blagih J, Domart MC, Vousden KH. Cell clustering promotes a metabolic switch that supports metastatic colonization. Cell Metab. 2019;30:720–734.e725.
Lancaster MA, Knoblich JA. Organogenesis in a dish: modeling development and disease using organoid technologies. Science (New York, NY). 2014;345:1247125.
Lawson DA, Bhakta NR, Kessenbrock K, Prummel KD, Yu Y, Takai K, et al. Single-cell analysis reveals a stem-cell program in human metastatic breast cancer cells. Nature. 2015;526:131–5.
Lee GY, Kenny PA, Lee EH, Bissell MJ. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat Methods. 2007;4:359–65.
Lee CH, Yu CC, Wang BY, Chang WW. Tumorsphere as an effective in vitro platform for screening anti-cancer stem cell drugs. Oncotarget. 2016;7:1215–26.
Lee P, Chandel NS, Simon MC. Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat Rev Mol Cell Biol. 2020;21:268–83.
Li H, Durbin R. Fast and accurate long-read alignment with burrows–wheeler transform. Bioinformatics. 2010;26:589–95.
Lin EY, Jones JG, Li P, Zhu L, Whitney KD, Muller WJ, et al. Progression to malignancy in the polyoma middle T oncoprotein mouse breast cancer model provides a reliable model for human diseases. Am J Pathol. 2003;163:2113–26.
Liu K, Newbury PA, Glicksberg BS, Zeng WZD, Paithankar S, Andrechek ER, et al. Evaluating cell lines as models for metastatic breast cancer through integrative analysis of genomic data. Nat Commun. 2019a;10:2138.
Liu X, Taftaf R, Kawaguchi M, Chang YF, Chen W, Entenberg D, et al. Homophilic CD44 interactions mediate tumor cell aggregation and polyclonal metastasis in patient-derived breast cancer models. Cancer Discov. 2019b;9:96–113.
Liu Y, Mi Y, Mueller T, Kreibich S, Williams EG, Van Drogen A, et al. Multi-omic measurements of heterogeneity in HeLa cells across laboratories. Nat Biotechnol. 2019c;37:314–22.
Martinez-Outschoorn UE, Peiris-Pagés M, Pestell RG, Sotgia F, Lisanti MP. Cancer metabolism: a therapeutic perspective. Nat Rev Clin Oncol. 2017;14:11–31.
Martino F, Perestrelo AR, Vinarský V, Pagliari S, Forte G. Cellular mechanotransduction: from tension to function. Front Physiol. 2018;9.
Maru Y, Orihashi K, Hippo Y. Lentivirus-based stable gene delivery into intestinal organoids. Methods in molecular biology (Clifton, NJ). 2016;1422:13–21.
Misra S, Moro CF, Del Chiaro M, Pouso S, Sebestyén A, Löhr M, et al. Ex vivo organotypic culture system of precision-cut slices of human pancreatic ductal adenocarcinoma. Sci Rep. 2019;9:2133.
Neal JT, Li X, Zhu J, Giangarra V, Grzeskowiak CL, Ju J, et al. Organoid modeling of the tumor immune microenvironment. Cell. 2018;175:1972–1988.e1916.
Nguyen-Ngoc KV, Shamir ER, Huebner RJ, Beck JN, Cheung KJ, Ewald AJ. 3D culture assays of murine mammary branching morphogenesis and epithelial invasion. Methods in molecular biology (Clifton, NJ). 2015;1189:135–62.
Padmanaban V, Krol I, Suhail Y, Szczerba BM, Aceto N, Bader JS, et al. E-cadherin is required for metastasis in multiple models of breast cancer. Nature. 2019;573:439–44.
Padmanaban V, Grasset EM, Neumann NM, Fraser AK, Henriet E, Matsui W, et al. Organotypic culture assays for murine and human primary and metastatic-site tumors. Nat Protoc. 2020;15:2413–42.
Rankin EB, Giaccia AJ. Hypoxic control of metastasis. Science (New York, NY). 2016;352:175–80.
Rennhack JP, To, B, Swiatnicki M, Dulak C, Ogrodzinski MP, Zhang Y, et al. Integrated analyses of murine breast cancer models reveal critical parallels with human disease. Nat Commun. 2019;10:3261.
Rosenbluth JM, Schackmann RCJ, Gray GK, Selfors LM, Li CM-C, Boedicker M, et al. Organoid cultures from normal and cancer-prone human breast tissues preserve complex epithelial lineages. Nat Commun. 2020;11:1711.
Sachs N, de Ligt J, Kopper O, Gogola E, Bounova G, Weeber F, et al. A living biobank of breast cancer organoids captures disease heterogeneity. Cell. 2018;172:373–386.e310.
Savage P, Blanchet-Cohen A, Revil T, Badescu D, Saleh SMI, Wang YC, et al. A targetable EGFR-dependent tumor-initiating program in breast cancer. Cell Rep. 2017;21:1140–9.
Shahbazi MN, Jedrusik A, Vuoristo S, Recher G, Hupalowska A, Bolton V, et al. Self-organization of the human embryo in the absence of maternal tissues. Nat Cell Biol. 2016;18:700–8.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science (New York, NY). 2014;343:84–7.
Shamir ER, Ewald AJ. Three-dimensional organotypic culture: experimental models of mammalian biology and disease. Nat Rev Mol Cell Biol. 2014;15:647–64.
Sheridan JM, Ritchie ME, Best SA, Jiang K, Beck TJ, Vaillant F, et al. A pooled shRNA screen for regulators of primary mammary stem and progenitor cells identifies roles for Asap1 and Prox1. BMC Cancer. 2015;15:221.
Simian M, Bissell MJ. Organoids: a historical perspective of thinking in three dimensions. J Cell Biol. 2017;216:31–40.
Smits LM, Reinhardt L, Reinhardt P, Glatza M, Monzel AS, Stanslowsky N, et al. Modeling Parkinson’s disease in midbrain-like organoids. NPJ Parkinson's Disease. 2019;5:5.
Takebe T, Wells JM. Organoids by design. Science (New York, NY). 2019;364:956–9.
Tuveson D, Clevers H. Cancer modeling meets human organoid technology. Science (New York, NY). 2019;364:952–5.
Wrenn EW, Yamamoto A, Moore BM, Huang,Y, McBirney M, Thomas AJ, Greenwood E, Rabena YF, Rahbar H, Partridge SC, et al. Regulation of collective metastasis by nanolumenal signaling. Cell. 2020;183(2):395-410.e19.
Xiang L, Yin Y, Zheng Y, Ma Y, Li Y, Zhao Z, et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature. 2020;577:537–42.
Xie C, Tammi MT. CNV-seq, a new method to detect copy number variation using high-throughput sequencing. BMC Bioinformatics. 2009;10:80.
Zhou J, Li C, Liu X, Chiu MC, Zhao X, Wang D, et al. Infection of bat and human intestinal organoids by SARS-CoV-2. Nat Med. 2020;26:1077–83.
Acknowledgments
We thank Patrick Paddison for providing sgRNA reagents and lentiviral packaging plasmids. We thank Andrew Ewald and Sasha Stanton for providing mouse tumors. We thank David MacPherson and Nan Wu for CNV sample preparation and analysis. This work was supported by grants from the Department of Defense W81XWH-18-1-0098, NIH R37CA234488, the Burroughs Wellcome Fund Career Award for Medical Scientists 1013355.01, the Phi Beta Psi Sorority, the Breast Cancer Research Foundation BCRF-18-035, the V Foundation V2017-014, Seattle Translational Tumor Research, and the Fred Hutch/University of Washington Cancer Consortium (P30 CA015704). This research was also supported by Fred Hutch Comparative Medicine Shared Resources. The authors declare no competing interests.
Author information
Authors and Affiliations
Contributions
EDW and KJC drafted the manuscript. KJC supervised the project and secured funding. EDW, BMM, EG, and MM designed and conducted experiments.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Movie S1
Growth dynamics of mammary tumor organoids embedded in 3D Matrigel. Left, hourly DIC time lapse images of 3 organoids of different sizes (from Fig. 1a) cultured embedded in 100% Matrigel. Right, summary of growth rate ([current area-initial area]/initial area) measured every 10 h. (MP4 12645 kb)
Rights and permissions
About this article
Cite this article
Wrenn, E.D., Moore, B.M., Greenwood, E. et al. Optimal, Large-Scale Propagation of Mouse Mammary Tumor Organoids. J Mammary Gland Biol Neoplasia 25, 337–350 (2020). https://doi.org/10.1007/s10911-020-09464-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10911-020-09464-1