Abstract
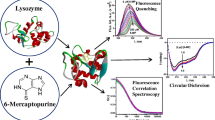
The interactions between tetrasulfophthalocyanines and lysozyme were studied using fluorescence spectroscopic and computational analyses. Lysozyme has been found to be widely studied as an anticancer agent, however, there are few reports of its interaction with phthalocyanines. Fe(III) tetrasulfophthalocyanine (FeTSPc) and free base tetrasulfophthalocyanine (TSPc) used in this study, were synthesized by our research group. Experimental results suggested that the metalled complex FeTSPc has a much higher affinity than TSPc. The binding stoichiometry between each tetrasulfophthalocyanine and lysozyme was 1:1. Stern−Volmer analysis suggested that the fluorescence quenching proceedes through a static process. Binding thermodynamics (ΔG, ΔH and ΔS) confirmed that mainly hydrogen bonds, van der Waals, and electrostatic forces are responsible for the binding process. We carried out molecular dynamics simulations, molecular docking, and binding energy calculations. Molecular dynamics simulations yielded the most populated cluster of lysozyme structures, and a representative structure from this cluster was used for the docking studies with these phthalocyanines. 1000 poses were generated for each ligand. The strudtures of the resulting complexes revealed that Arg 73 and Arg 112 are important for the binding affinity of the tetrasulfophthalocyanines, generating mainly an electrostatic favorable environment for the SO3− groups. In addition, hydrophobic contacts were involved with Trp 62, Trp 63 and Trp 108, explaining the fluorescence quenching observed experimentally. Binding energies were determined for these models, confirming that the interactions with lysozyme were more favorable for FeTSPc compared to TSPc. The understanding of the molecular mechanisms is relevant to characterize the nature of tetrasulfophthalocyanines in photodynamic therapy.






Similar content being viewed by others
Abbreviations
- A(280 nm, 1%) :
-
masss extinction coefficient
- APBS:
-
Adaptive Poisson-Boltzmann Solver
- ΔGCoul :
-
Coulombic energy
- ΔGelec :
-
electrostatic energy
- ΔGcalc :
-
binding free energy
- ΔG:
-
Gibbs free energy change
- ΔGnon-elec :
-
non-electrostatic energy
- ΔGsolv :
-
solvation energy
- ΔH:
-
enthalpy change
- ΔS:
-
entropy change
- ΔSASA:
-
solvent-accessible surface area change
- FeTSPc:
-
Fe(III) tetrasulfophthalocyanine
- F0 or F:
-
fluorescence intensities
- γ:
-
interfacial tension coefficient
- Kb :
-
binding equilibrium constant
- KSV :
-
Stern-Volmer constant
- kq :
-
quenching rate constant
- λexc :
-
excitation wavelength
- λem :
-
emission wavelength
- NAMD:
-
Not Another Molecular Dynamics Program
- n :
-
number of binding sites
- Q:
-
molar concentration
- R:
-
gas constant
- RMSD:
-
Root-Mean-Square Deviation
- T:
-
absolute temperature
- τ0 :
-
average lifetime of biomolecules
- TSPc:
-
free base tetrasulfophthalocyanine
- UV-Vis:
-
Ultraviolet-Visible
- VMD:
-
Visual Molecular Dynamics
References
Örstan A, Wojcik JF (1987) Spectroscopic determination of protein-ligand binding constants. J Chem Educ 64:814–816. https://doi.org/10.1021/ed064p814
Marcoline AT, Elgren TE (1988) A thermodynamic study of azide binding myoglobin. J Chem Educ 75:1622–1623. https://doi.org/10.1021/ed075p1622
Banerjee S, Choudhury SD, Dasgrupta S, Basu S (2008) Photoinduced electron transfer between hen egg white lysozyme and anticancer drug menadione. J Lumin 128:437–444. https://doi.org/10.1016/j.jlumin.2007.09.020
Mitra P, Banerjee M, Biswas S, Basu S (2013) Protein interactions of merocyanine 540: spectroscopic and crystallographic studies with lysozyme as a model protein. J Photochem Photobiol B Biol 121:46–56. https://doi.org/10.1016/j.jphotobiol.2013.02.010
García-Sánchez MA, Rojas-González F, Menchaca-Campos EC, Tello-Solís SR, Quiroz-Segoviano RIY, Díaz-Alejo LA, Salas-Bañales E, Campero C (2013) Crossed and linked histories of tetrapyrrolic macrocycles and their use for engineering pores within sol-gel matrices. Molecules 18:588–653. https://doi.org/10.3390/molecules18010588
Yano S, Hirohara S, Obata M, Hagiya Y, Ogurad SI, Ikeda A, Kataoka H, Tanaka M, Joh T (2011) Current states and future views in photodynamic therapy. J Photochem Photobiol C 12:46–67. https://doi.org/10.1016/j.jphotochemrev.2011.06.001
Castano AP, Demidova TN, Hamblin M (2004) Mechanisms in photodynamic therapy: part one-photosensitizers, photochemistry and cellular localization. Photodiagn Photodyn Ther 1:279–293. https://doi.org/10.1016/S1572-1000(05)00007-4
Li S, Li D (2011) Investigation on the pH-dependent binding of benzocaine and lysozyme by fluorescence and absorbance. Spectrochim Acta A Mol Biomol Spectrosc 82:396–405. https://doi.org/10.1016/j.saa.2011.07.069
Nash JA, Ballard TNS, Weaver TE, Akinbi HT (2006) The peptidoglycan-degrading property of lysozyme is not required for bactericidal activity in vivo. J Immunol 177:519–526. https://doi.org/10.4049/jimmunol.177.1.519
Ghosh KS, Sahoo BK, Dasgupta S (2008) Spectrophotometric studies on the interaction between (−)-epigallocatechin gallate and lysozyme. Chem Phys Lett 452:193–197. https://doi.org/10.1016/j.cplett.2007.12.018
Wang W, Min W, Chen J, Wu X, Hu Z (2011) Binding study of diprophylline with lysozyme by spectroscopic methods. J Lumin 131:354–359. https://doi.org/10.1016/j.jlumin.2010.12.010
Li D, Zhu J, Jin J (2007) Spectrophotometric studies on the interaction between nevadensin and lysozyme. J Photochem Photobiol A 189:114–120. https://doi.org/10.1016/j.jphotochem.2007.01.017
Kovalska V, Chernii S, Cherepanov V, Losytskyy M, Chernii V, Varzatskii O, Naumovets A, Yarmoluk S (2017) The impact of binding of macrocyclic metal complexes on amyloid fibrillization of insulin and lysozyme. J Mol Recognit 30:1–10. https://doi.org/10.1002/jmr.2622
Coahuila-Hernández MI, García-Sánchez MA, Soto Estrada AM, Campero A (2006) Lanthanide tetrasulfophthalocyanines incorporated to SiO2 gels. J Sol-Gel Sci Techn 37:117–120. https://doi.org/10.1007/s10971-006-6429-8
Ito Y, Yoshikawa A, Hotani T, Fukuda S, Sugimura K, Imoto T (1999) Amino acid sequences of lysozymes newly purified from invertebrates imply wide distribution of a novel class in the lysozyme family. Eur J Biochem 259:456–461. https://doi.org/10.1046/j.1432-1327.1999.00064.x
Andrade SM, Costa SM (2002) Spectroscopic studies on the interaction of a water soluble porphyrin and two drug carrier proteins. Biophys J 82:1607–1619. https://doi.org/10.1016/S0006-3495(02)75512-4
Samari F, Hemmateenejad B, Shamsipur M, Rashidi M, Samouei H (2012) Affinity of two novel five-coordinated anticancer Pt(II) complexes to human and bovine serum albumins: a spectroscopic approach. Inorg Chem 51:3454–3464. https://doi.org/10.1021/ic202141g
Zhao P, Huang JW, Ji LN (2012) Cationic pyridinium porphyrins appending different peripheral substituents: spectroscopic studies on their interactions with bovine serum albumin. Spectrochim Acta A 88:130–136. https://doi.org/10.1016/j.saa.2011.12.017
Mansouri M, Pirouzi M, Saberi MR, Ghaderabad M, Chamani J (2013) Investigation on the interaction between cyclophosphamide and lysozyme in the presence of three different kind of cyclodextrins: determination of the binding mechanism by spectroscopic and molecular modeling techniques. Molecules 18:789–813. https://doi.org/10.3390/molecules18010789
Wang J, Dauter M, Alkire R, Joachimiak A, Dauter Z (2007) Triclinic lysozyme at 0.65 a resolution. Acta Crystallogr. Sect D: Biol Crystallogr 63:1254–1268. https://doi.org/10.1107/S0907444907054224
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA, Im W (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12:405–413. https://doi.org/10.1021/acs.jctc.5b00935
Wu EL, Cheng X, Jo S, Rui H, Song KC, Dávila-Contreras EM, Im W (2014) CHARMM-GUI membrane builder toward realistic biological membrane simulations. J Comput Chem 35:1997–2004. https://doi.org/10.1002/jcc.23702
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865. https://doi.org/10.1002/jcc.20945
Phillips JC, Braun R, Wang W, Gumbar J, Tajkhorshid E, Villa E, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802. https://doi.org/10.1002/jcc.20289
Huang J, MacKerell AD Jr (2013) CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comput Chem 34:2135–2145. https://doi.org/10.1002/jcc.23354
Mezei M (2010) Simulaid: a simulation facilitator and analysis program. J Comput Chem 31:2658–2668. https://doi.org/10.1002/jcc.21551
Trott O, Olson AJ (2010) AutoDockVina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
ChemAxon (2019). Marvin 19.20. Retrieved from www.chemaxon.com
Schrödinger L (2019). Maestro. Schrödinger Release 2019–3. New York, NY
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci U S A 98:10037–11004. https://doi.org/10.1073/pnas.181342398
Lira-De León KI, García-Gutiérrez P, Serratos IN, Palomera-Cárdenas M, Figueroa-Corona MP, Campos-Peña V, Meraz-Ríos MA (2013) Molecular mechanism of tau aggregation induced by anionic and cationic dyes. J Alzheimers Dis 35:319–334. https://doi.org/10.3233/JAD-121765
Moral-Rodríguez AI, Leyva-Ramos R, Ocampo-Pérez R, Mendoza-Barron J, Serratos-Alvarez IN, Salazar-Rabago JJ (2016) Removal of ronidazole and sulfamethoxazole from wáter solutions by adsorption on granular activated carbon: equilibrium and intra particle diffusion mechanisms. Adsorption 22:89–103. https://doi.org/10.1007/s10450-016-9758-0
Bustos-Terrones V, Serratos IN, Castañeda-Villa N, Vicente Escobar JO, Romero Romo MA, Córdoba G, Uruchurtu Chavarín J, Menchaca Campos C, Esparza Schulz JM, Domínguez Ortiz A (2018) Functionalized coatings based on organic polymer matrix against the process of corrosion of mild steel in neutral medium. Prog Org Coat 119:221–229. https://doi.org/10.1016/j.porgcoat.2017.12.011
Bustos-Terrones V, Serratos IN, Vargas R, Landeros-Rivera BC, Bustos-Terrones YA, Soto Estrada AM, Vicente Escobar JO, Romero Romo MA, Uruchurtu J, Menchaca C, Esparza Schulz JM, Domínguez A (2018) SBA15–fluconazole as a protective approach against mild steel corrosion: synthesis, characterization, and computational studies. ChemistryOpen 7:984–994. https://doi.org/10.1002/open.201800201
Serratos IN, Olayo R, Millán-Pacheco C, Morales-Corona J, Vicente-Escobar JO, Soto-Estrada AM, Córdoba-Herrera JG, Uribe O, Gómez-Quintero T, Arroyo-Ornelas MA, Godínez-Fernández R (2019) Modeling integrin and plasma-polymerized pyrrole interactions: chemical diversity relevance for cell regeneration. Sci Rep 9:7009. https://doi.org/10.1038/s41598-019-43286-4
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: An automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res Spec Publ 32:W665–W667. https://doi.org/10.1093/nar/gkh381
MacKerell AD Jr, Bashford D, Bellott M et al (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102:3586–3616. https://doi.org/10.1021/jp973084f
Sondergaard CR, Olsson MH, Rostkowski M, Jensen JH (2011) Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J Chem Theory Comput 7:2284–2295. https://doi.org/10.1021/ct200133y
Sitkoff D, Sharp KA, Honig B (1994) Accurate calculation of hydration free energies using macroscopic solvent models. J Phys Chem 98:1978–1988. https://doi.org/10.1021/j100058a043
Levy RM, Zhang LY, Gallicchio E, Felts AK (2003) On the nonpolar hydration free energy of proteins: surface area and continuum solvent models for the solute-solvent interaction energy. J Am Chem Soc 125:9523–9530. https://doi.org/10.1021/ja029833a
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
An W, Jiao Y, Dong C, Yang C, Inoue I, Shuang S (2009) Spectroscopic and molecular modeling of the binding of meso-tetrakis(4-hydroxyphenyl)porphyrin to human serum albumin. Dyes Pigments 81:1–9. https://doi.org/10.1016/j.dyepig.2008.08.004
Hu Y, Zhang G, Yan J (2014) Detection of interaction between lysionotin and bovine serum albumin using spectroscopic techniques combined with molecular modeling. Mol Biol Rep 41:1693–1702. https://doi.org/10.1007/s11033-013-3018-0
Ross PD, Subramanian S (1981) Thermodynamics of protein association reactions: forces contributing to stability. Biochemistry 20:3096–3102. https://doi.org/10.1021/bi00514a017
Jiménez-Pérez C, Tello-Solís SR, Gómez-Castro CZ, Alatorre-Santamaría S, Gómez-Ruíz L, Rodríguez-Serrano G, Cruz-Borbolla J, García-Garibay M, Cruz-Guerrero A (2020) Spectroscopic studies and molecular modelling of aflatoxin M1-bovine α-lactalbumin complex formation. J Photoch Photobio B 111957:1-8 (online). https://doi.org/10.1016/j.jphotobiol.2020.111957
Lakowicz JR (2006) Introduction to fluorescence, in: principles of fluorescence spectroscopy, 3rd edn. Springer, New York
Delavari B, Saboury AA, Atri MS, Ghasemi A, Bigdeli B, Khammari A, Maghami P, Moosavi-Movahedi AA, Haertlé T, Goliaei B (2015) Alpha-lactalbumin: a new carrier for vitamin D3 food enrichment. Food Hydrocoll 45:124–131. https://doi.org/10.1016/j.foodhyd.2014.10.017
Suryawanshi VD, Walekar LS, Gore AH, Anbhule PV, Kolekar GB (2016) Spectroscopicanalysis on the binding interaction of biologically active pyrimidine derivative with bovine serum albumin. J Pharm Anal 6:56–63. https://doi.org/10.1016/j.jpha.2015.07.001
Du X, Li Y, Xia YL, Ai SM, Liang J, Sang P, Ji XL, Liu SQ (2016) Insights into protein-ligand interactions: mechanisms, models, and methods. Int J Mol Sci 17:1–34. https://doi.org/10.3390/ijms17020144
Acknowledgments
The authors thank Dr. Nina Pastor Colón, for helpful discussions and advise as well as her help with the revision of the English language. J.O.V.E thanks CONACyT for the posgraduate scholarship No. 130442 for the studies of Master of Sciences (Chemistry). The Laboratorio de Visualización y Cómputo Paralelo at Universidad Autónoma Metropolitana Iztapalapa is acknowledged for computing time.
Availability of Data and Material
The authors have no any objection for availability of data and materials.
Code Availability
Hen egg-white Lysozyme was purchased from Sigma Chemical Co. (lot. 73H7045).
Crystal structure of Hen egg-white lysozyme from Gallus Gallus. https://www.rcsb.org/structure/2VB1
ACD/ChemSketch Freeware http://www.acdlabs.com/resources/freeware/chemsketch/
Visual Molecular Dynamics (VMD) https://www.ks.uiuc.edu/Research/vmd/
NAMD https://www.ks.uiuc.edu/Research/namd/
Autodock Vina http://vina.scripps.edu
Maestro https://www.schrodinger.com/products/maestro
Charmm-gui web server http://www.charmm-gui.org
Funding
This research was supported by Universidad Autónoma Metropolitana Iztapalapa.
Author information
Authors and Affiliations
Contributions
Jonathan Osiris Vicente-Escobar: investigation, analysis, writing; Miguel Ángel García-Sánchez: investigation, review, formal analysis, resources; Iris N. Serratos: conceptualization, visualization, investigation, software, formal analysis, writing, review & editing, César Millán-Pacheco: investigation, software, formal analysis; Salvador R. Tello-Solís: conceptualization, project administration, visualization, methodology, investigation, formal analysis, writing, review, editing, & resources.
Corresponding author
Ethics declarations
Ethics Approval
“Not applicable”.
Consent to Participate
All authors gave their consent to participate in the research.
Consent for Publication
All authors gave their consent to participate in the publication of the research.
Conflicts of Interest/Competing Interests
All authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vicente-Escobar, J.O., García-Sánchez, M.Á., Serratos, I.N. et al. Binding of Two Tetrasulfophthalocyanines (Fe(III) and Metal-Free) to Lysozyme: Fluorescence Spectroscopic and Computational Approach. J Fluoresc 31, 787–796 (2021). https://doi.org/10.1007/s10895-021-02710-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-021-02710-7