Abstract
Purpose
Newborn screening (NBS) quantifies T cell receptor excision circles (TREC) and identifies infants with T cell lymphopenia (TCL). This study elucidates the demographics, laboratory characteristics, genetics, and clinical outcomes following live viral vaccine administration of term infants with transient or persistent idiopathic TCL.
Methods
A single-center retrospective analysis was performed from September 2010 through June 2018. Laboratory variables were compared with Mann-Whitney tests. Correlations between initial TREC levels and T cell counts were determined by Spearman tests.
Results
Twenty-two transient and 21 persistent TCL infants were identified. Males comprised 68% of the transient and 52% of the persistent TCL cohorts. Whites comprised 23% of the transient and 29% of the persistent cohorts. Median initial TREC levels did not differ (66 vs. 60 TRECs/μL of blood, P = 0.58). The transient cohort had higher median initial CD3+ (2135 vs. 1169 cells/μL, P < 0.001), CD4+ (1460 vs. 866 cells/μL, P < 0.001), and CD8+ (538 vs. 277 cells/μL, P < 0.001) counts. The median age of resolution for the transient cohort was 38 days. Genetic testing revealed 2 genes of interest which warrant further study and several variants of uncertain significance in immunology-related genes in the persistent cohort. 19 transient and 14 persistent subjects received the initial rotavirus and/or MMRV immunization. No adverse reactions to live viral vaccines were reported in either cohort.
Conclusion
Transient and persistent TCL infants differ by demographic, laboratory, and clinical characteristics. Select transient and persistent TCL patients may safely receive live attenuated viral vaccines, but larger confirmatory studies are needed.




Similar content being viewed by others
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Guthrie R, Susi A. A simple phenylalanine method for detecting phenylketonuria in large populations of newborn infants. Pediatrics. 1963;32(3):338–43.
Routes JM, Grossman WJ, Verbsky J, Laessig RH, Hoffman GL, Brokopp CD, et al. Statewide newborn screening for severe T-cell lymphopenia. JAMA. 2009;302(22):2465–70.
Comeau AM, Hale JE, Pai SY, Bonilla FA, Notarangelo LD, Pasternack MS, et al. Guidelines for implementation of population-based newborn screening for severe combined immunodeficiency. J Inherit Metab Dis. 2010;33(Suppl 2):S273–81.
Gerstel-Thompson JL, Wilkey JF, Baptiste JC, Navas JS, Pai SY, Pass KA, et al. High-throughput multiplexed T-cell-receptor excision circle quantitative PCR assay with internal controls for detection of severe combined immunodeficiency in population-based newborn screening. Clin Chem. 2010;56(9):1466–74.
Accetta D, Syverson G, Bonacci B, Reddy S, Bengtson C, Surfus J, et al. Human phagocyte defect caused by a Rac2 mutation detected by means of neonatal screening for T-cell lymphopenia. J Allergy Clin Immunol. 2011;127(2):535–8.e1–2.
Kwan A, Abraham RS, Currier R, Brower A, Andruszewski K, Abbott JK, et al. Newborn screening for severe combined immunodeficiency in 11 screening programs in the United States. JAMA. 2014;312(7):729–38.
Vogel BH, Bonagura V, Weinberg GA, Ballow M, Isabelle J, DiAntonio L, et al. Newborn screening for SCID in New York State: experience from the first two years. J Clin Immunol. 2014;34(3):289–303.
Buelow BJ, Verbsky JW, Routes JM. Newborn screening for SCID: lessons learned. Expert Rev Hematol. 2016;9(6):579–84.
Amatuni GS, Currier RJ, Church JA, Bishop T, Grimbacher E, Nguyen AA, et al. Newborn Screening for Severe Combined Immunodeficiency and T-cell Lymphopenia in California, 2010–2017. Pediatrics. 2019;143(2).
Serana F, Chiarini M, Zanotti C, Sottini A, Bertoli D, Bosio A, et al. Use of V(D)J recombination excision circles to identify T- and B-cell defects and to monitor the treatment in primary and acquired immunodeficiencies. J Transl Med. 2013;11:119.
Verbsky JW, Baker MW, Grossman WJ, Hintermeyer M, Dasu T, Bonacci B, et al. Newborn screening for severe combined immunodeficiency; the Wisconsin experience (2008-2011). J Clin Immunol. 2012;32(1):82–8.
Verbsky J, Thakar M, Routes J. The Wisconsin approach to newborn screening for severe combined immunodeficiency. J Allergy Clin Immunol. 2012;129(3):622–7.
Kwan A, Puck JM. History and current status of newborn screening for severe combined immunodeficiency. Semin Perinatol. 2015;39(3):194–205.
Puck JM. Newborn screening for severe combined immunodeficiency and T-cell lymphopenia. Immunol Rev. 2019;287(1):241–52.
Albin-Leeds S, Ochoa J, Mehta H, Vogel BH, Caggana M, Bonagura V, et al. Idiopathic T cell lymphopenia identified in New York State Newborn Screening. Clin Immunol. 2017;183:36–40.
Aluri J, Gupta MR, Dalvi A, Mhatre S, Kulkarni M, Desai M, et al. Lymphopenia and severe combined immunodeficiency (SCID) - think before you ink. Indian J Pediatr. 2019;86:584–9.
Rios X, Chinn IK, Orange JS, Hanson CI, Rider NL. T-cell lymphopenia detected by newborn screening in two siblings with an Xq13.1 duplication. Front Pediatr. 2017;5:156.
Patrawala M, Kobrynski L. Nonsevere combined immunodeficiency T-cell lymphopenia identified through newborn screening. Curr Opin Allergy Clin Immunol. 2019;19(6):586–93.
Kobrynski LJ. Identification of non-severe combined immune deficiency T-cell lymphopenia at newborn screening for severe combined immune deficiency. Ann Allergy Asthma Immunol. 2019;123(5):424–7.
Gholamin M, Bazi A, Abbaszadegan MR. Idiopathic lymphopenia. Curr Opin Hematol. 2015;22(1):46–52.
Shearer WT, Rosenblatt HM, Gelman RS, Oyomopito R, Plaeger S, Stiehm ER, et al. Lymphocyte subsets in healthy children from birth through 18 years of age: the Pediatric AIDS Clinical Trials Group P1009 study. J Allergy Clin Immunol. 2003;112(5):973–80.
Baldeyron C, Soria G, Roche D, Cook AJ, Almouzni G. HP1alpha recruitment to DNA damage by p150CAF-1 promotes homologous recombination repair. J Cell Biol. 2011;193(1):81–95.
Rodriges Blanko E, Kadyrova LY, Kadyrov FA. DNA mismatch repair interacts with CAF-1- and ASF1A-H3-H4-dependent histone (H3-H4)2 tetramer deposition. J Biol Chem. 2016;291(17):9203–17.
Takahashi D, Hase K, Kimura S, Nakatsu F, Ohmae M, Mandai Y, et al. The epithelia-specific membrane trafficking factor AP-1B controls gut immune homeostasis in mice. Gastroenterology. 2011;141(2):621–32.
Davies AA, Masson JY, McIlwraith MJ, Stasiak AZ, Stasiak A, Venkitaraman AR, et al. Role of BRCA2 in control of the RAD51 recombination and DNA repair protein. Mol Cell. 2001;7(2):273–82.
Xia F, Taghian DG, DeFrank JS, Zeng ZC, Willers H, Iliakis G, et al. Deficiency of human BRCA2 leads to impaired homologous recombination but maintains normal nonhomologous end joining. Proc Natl Acad Sci U S A. 2001;98(15):8644–9.
Bergmann C, Fliegauf M, Brüchle NO, Frank V, Olbrich H, Kirschner J, et al. Loss of nephrocystin-3 function can cause embryonic lethality, Meckel-Gruber-like syndrome, situs inversus, and renal-hepatic-pancreatic dysplasia. Am J Hum Genet. 2008;82(4):959–70.
Moylett EH, Wasan AN, Noroski LM, Shearer WT. Live viral vaccines in patients with partial DiGeorge syndrome: clinical experience and cellular immunity. Clin Immunol. 2004;112(1):106–12.
Waters V, Peterson KS, LaRussa P. Live viral vaccines in a DiGeorge syndrome patient. Arch Dis Child. 2007;96(6):519–20.
Markert ML. Defects in thymic development. In: Sullivan KE, Stiehm RE, editors. Stiehm’s Immune Deficiencies. Second ed. London: Elsevier; 2020. p. 357–80.
Perez EE, Bokszczanin A, McDonald-McGinn D, Zackai EH, Sullivan KE. Safety of live viral vaccines in patients with chromosome 22q11.2 deletion syndrome (DiGeorge syndrome/velocardiofacial syndrome). Pediatrics. 2003;122(4):e325.
Azzari C, Gambineri E, Resti M, Moriondo M, Betti L, Saldias LR, et al. Safety and immunogenicity of measles-mumps-rubella vaccine in children with congenital immunodeficiency (DiGeorge syndrome). Vaccine. 2005;23(14):1668–71.
Davis CM, Kancherla VS, Reddy A, Chan W, Yeh HW, Noroski LM, et al. Development of specific T-cell responses to Candida and tetanus antigens in partial DiGeorge syndrome. J Allergy Clin Immunol. 2008;122(6):1194–9.
Koup RA, Douek DC. Vaccine design for CD8 T lymphocyte responses. Cold Spring Harb Perspect Med. 2011;1(1):a007252.
https://www.health.ny.gov/statistics/vital_statistics/ accessed 6/20/2020.
Gans MD, Gavrilova T. Retrospective analysis of a New York newborn screen severe combined immunodeficiency referral center. J Clin Immunol. 2020;40(3):456–65.
Cheloufi S, Elling U, Hopfgartner B, Jung YL, Murn J, Ninova M, et al. The histone chaperone CAF-1 safeguards somatic cell identity. Nature. 2015;528(7581):218–24.
Ng C, Aichinger M, Nguyen T, Au C, Najar T, Wu L, et al. The histone chaperone CAF-1 cooperates with the DNA methyltransferases to maintain. Genes Dev. 2019;33(11–12):669–83.
Bosticardo M, Yamazaki Y, Cowan J, Giardino G, Corsino C, Scalia G, et al. Heterozygous FOXN1 variants cause low TRECs and severe T cell lymphopenia, revealing a crucial role of FOXN1 in supporting early thymopoiesis. Am J Hum Genet. 2019;105(3):549–61.
Quinn J, Modell V, Holle J, Truty R, Aradhya S, Johnson B, et al. Jeffrey’s insights: Jeffrey Modell Foundation’s global genetic sequencing pilot program to identify specific primary immunodeficiency defects to optimize disease management and treatment. Immunol Res. 2020;68(3):126–34.
Chinn IK, Chan AY, Chen K, Chou J, Dorsey MJ, Hajjar J, et al. Diagnostic interpretation of genetic studies in patients with primary immunodeficiency diseases: a working group report of the Primary Immunodeficiency Diseases Committee of the American Academy of Allergy, Asthma & Immunology. J Allergy Clin Immunol. 2020;145(1):46–69.
Sullivan KE. The scary world of variants of uncertain significance (VUS): a hitchhiker’s guide to interpretation. J Allergy Clin Immunol. 2020. https://doi.org/10.1016/j.jaci.2020.06.011.
Zhang S, Elshaigi O, Daian F, Bae E, Innamorato A, Navetta-Modrov B, et al. Describing single nucleotide polymorphisms (SNPs) transient T cell lymphopenia in the United States Immunodeficiency Network (USIDNET) following infants with low lymphocytes (FILL) program and a single referral center from 2010-2017. J Clin Immunol. 2019;39(Suppl 1):S13.
Gans MD, Saavedra-Matiz CA, Bernstein L. A single nucleotide polymorphism in the T-cell receptor excision circle. J Allergy Clin Immunol Pract. 2020;8(2):803–5.e1.
Dorsey MJ, Dvorak CC, Cowan MJ, Puck JM. Treatment of infants identified as having severe combined immunodeficiency by means of newborn screening. J Allergy Clin Immunol. 2017;139(3):733–42.
Amatuni GS, Sciortino S, Currier RJ, Naides SJ, Church JA, Puck JM. Reference intervals for lymphocyte subsets in preterm and term neonates without immune defects. J Allergy Clin Immunol. 2019;144(6):1674–83.
Shoenfeld Y, Alkan ML, Asaly A, Carmeli Y, Katz M. Benign familial leukopenia and neutropenia in different ethnic groups. Eur J Haematol. 1988;41(3):273–7.
Gitlin D, Janeway CA. Agammaglobulinemia, congenital, acquired and transient forms. Prog Hematol. 1956;1:318–29.
Zonios DI, Falloon J, Bennett JE, Shaw PA, Chaitt D, Baseler MW, et al. Idiopathic CD4+ lymphocytopenia: natural history and prognostic factors. Blood. 2008;112(2):287–94.
Busch MP, Valinsky JE, Paglieroni T, Prince HE, Crutcher GJ, Gjerset GF, et al. Screening blood donors for idiopthic CD4+ T-lymphocyotpenia. Transfusion. 1994;34(3):192–7.
Lisco A, Freeman AF, Sereti I. Idiopathic CD4 lymphopenia. In: Sullivan KE, Stiehm RE, editors. Stiehm’s immune deficiencies. Second ed. London: Elsevier; 2020. p. 381–92.
Walker UA, Warnatz K. Idiopathic CD4 lymphocytopenia. Curr Opin Rheumatol. 2006;18(4):389–95.
Régent A, Autran B, Carcelain G, Cheynier R, Terrier B, Charmeteau-De Muylder B, et al. Idiopathic CD4 lymphocytopenia: clinical and immunologic characteristics and follow-up of 40 patients. Medicine (Baltimore). 2014;93(2):61–72.
Kuo CY, Garcia-Lloret MI, Slev P, Bohnsack JF, Chen K. Profound T-cell lymphopenia associated with prenatal exposure to purine antagonists detected by TREC newborn screening. J Allergy Clin Immunol Pract. 2017;5(1):198–200.
Acknowledgments
The authors thank Professor Christopher League for assistance with preparing figures for the manuscript.
Authorship Contributions
AMJ, RS, DWR, and VRB designed the study. AMJ, OE, EH, SZ, FD, EB, AI, CC, BNM, RS, DWR, and VRB contributed to data collection and analysis. AMJ, EH, RS, DWR, and VRB contributed to manuscript preparation. All authors reviewed the manuscript.
Funding
This was supported by a grant from the Immune Deficiency Foundation to Dr. Jongco.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
The Northwell Health Institutional Review Board approved this study, Protocol 19-0150-CCMC.
Consent to Participate
The Northwell Health Institution Review Board granted a waiver of informed consent and assent for this study.
Consent to Publish
Not applicable.
Disclaimer
The Immune Deficiency Foundation had no input on study design, data collection or analysis, and manuscript preparation.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Earlier versions of this data were presented as posters and platform presentations at the annual meetings of the American Academy of Allergy, Asthma & Immunology, and the Clinical Immunology Society.
Supplementary Information
Supplemental Figure 1
Initial laboratory parameters. Median CD19+ (A), CD16/56+ (B) counts. (PNG 4948 kb).
Supplemental Figure 2
Initial TREC and T cell counts do not correlate. Transient (A) and Persistent (B) cohort. For the transient cohort, Spearman coefficients (r) were as follows: CD3+ r = 0.06, P = 0.81, CD4+ r = −0.01, P = 0.99, CD8+ r = 0.05, P = 0.83. For the persistent cohort, Spearman coefficients were as follows: CD3+ r = 0.27, P = 0.24, CD4+ r = 0.32, P = 0.15, CD8+ r = 0.13, P = 0.57. TREC = T cell receptor excision circle (PNG 4948 kb).
Supplemental Figure 3
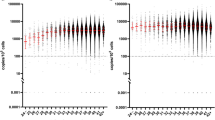
Longitudinal B and NK cell counts for transient and persistent cohorts plotted against the 10th and 90th percentiles for aged matched controls. Transient cohort values are plotted for CD19+ B (A) and CD16/56+ NK (B). Persistent cohort values are plotted for CD19+ B (C) and CD16/56+ NK (D). Subject T22 did not have his CD16/56+ NK cells measured and thus is not represented on Plot B. NK = natural killer (PNG 4948 kb).
Supplemental Table 1
(DOCX 152 kb).
Supplemental Table 2
(DOCX 40.8 kb).
Supplemental Table 3
Longitudinal CD3+, CD4+, CD8+, CD19+ and CD16/56+ counts for the persistent (Tab 1) and Transient (Tab 2) TCL cohorts. Blacked out boxes indicate the absence of data for that subject. WNL indicates that the records stated that the T cell counts had normalized for that subject. (XLSX 34 kb).
Supplemental Table 4
(DOCX 32.5 kb).
Supplemental Table 5
(DOCX 108 kb).
Rights and permissions
About this article
Cite this article
Jongco, A.M., Sporter, R., Hon, E. et al. Characterization of Infants with Idiopathic Transient and Persistent T Cell Lymphopenia Identified by Newborn Screening—a Single-Center Experience in New York State. J Clin Immunol 41, 610–620 (2021). https://doi.org/10.1007/s10875-020-00957-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-020-00957-6