Abstract
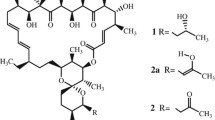
Detailed X-ray structures are presented of the three chemical variants of the antibiotic Oligomycin (Oligomycins A, B and C), all inhibitors of the enzyme ATP synthase, which has itself been the subject of intensive studies in recent years. All three oligomycins crystallized in space group P212121 with Z = 4 molecules per unit cell. Oligomycin A crystallized as the methanol solvate C45H72O11 · CH3OH with unit cell parameters a = 10.476(3), b = 17.342(1), c = 26.825(5) Å; oligomcin B as an acetic acid solvate C45H71O12 · CH3CO2 with unit cell parameters a = 10.351(3), b = 17.305(1), c = 26.929(5) Å; and oligomycin C, C45H74O10, with unit cell parameters a = 10.385(2), b = 11.9510(9), c = 38.007(4) Å. Oligomycin A refined with final R indices [I > 2sigma(I)], R1 = 0.0734, wR2 = 0.1940 and R indices (all data) R1 = 0.1106, wR2 = 0.2100 and absolute structure parameter = −0.7(4); for oligomycin B, final R indices [I > 2sigma(I)] are R1 = 0.0479, wR2 = 0.1388 and R indices (all data) are R1 = 0.0581, wR2 = 0.1435 and absolute structure parameter = −0.2(2); for oligomycin C, final R indices [I > 2sigma(I)] are R1 = 0.0454, wR2 = 0.1130 and R indices (all data) are R1 = 0.1061, wR2 = 0.1221 and absolute structure parameter = 0.1(3). The present study has resulted in providing: (1) corrections to the published chemical structures of the Oligomycins; (2) full descriptions of the absolute configurations; (3) information on regions of the structures with minor but important differences in their three dimensional structures which may create differences between the Oligomycins in their potential to bind to sites on the ATP synthase molecule. These results are all of major importance for future studies designed to establish details of the actual binding of Oligomycins to ATP synthase.
Index Abstract
Rex A. Palmer and Brian S. Potter
The antibiotics Oligomycin A, B and C are chemically very similar and all three are inhibitors of ATP synthases—enzymes, with complex multi-subunited protein structures, that can synthesize adenosine triphosphate (ATP) from adenosine diphosphate (ADP) and inorganic phosphate. In mitochondria, the ATP synthase molecule can be visualized as having a major domain, F1,outside the cell membrane, and a minor domain, FO, embedded within the membrane. FO (written as a subscript “O”, not “zero”) which derives it name from being the Oligomycin binding domain and is also known as OSCP (the oligomycin sensitivity conferal protein).

The X-ray structures of Oligomycins A, B and C, having subtle but important differences from each other as described here may enable the exact location of the binding site in ATP synthase to be characterized and differences in the relative binding characteristics of the Oligomycins to be evaluated






Similar content being viewed by others
Notes
In view of the unusual geometry revealed in this analysis for side chain R4, several complete intensity data sets were measured for Oligomycin A using fresh crystals. The results were always consistent with those reported above.
References
Lardy HA (1980) Pharmacol Ther, 11:649
Fossati G, Moulding DA, Spiller DG, Moots RJ, White MRH, Edwards SW (2003) J Immunol 170:1964
Altamura N, Capitanio N, Bonnefoy N, Papa S, Dujardin G (1996) FEBS Lett 382(1–2):111
Cho HJ, Balasubramanyam M, Chernaya G, Gardner JP, Aviv A, Reeves JP, Dargis PG, Christian EP (1997) Biochem J 324:971
von Glehn M, Norrestam R, Kierkegaard P, Maron L, Ernster L (1972) FEBS Lett 20(3):267
Fernandez-Moran H (1962) Circulation 26:1039
Abrahams JP, Leslie AGW, Lutter R, Walker JE (1994) Nature 370:621
Boyer PD (1993) Biochim Biophys Acta 1140(3):215
Enraf-Nonius (1988) CAD-4 Software. Enraf-Nonius, Delft, Holland
North ACT, Phillips DC, Mathews FS (1968) Acta Cryst A24:351
Sheldrick GM (1986) SHELXS-86: program for the solution of crystal structures. University of Göttingen, Germany
Sheldrick GM (1993) SHELXL-97: program for the refinement of crystal structures. University of Göttingen, Germany
Farrugia LJ (1998) J Appl Cryst 32:837
Spek AL (1990) Acta Crystallogr A46:C34
Farrugia LJ (1997) J Appl Cryst, 30:565 (Based on ORTEP-III (v 1.0.3) by CK Johnson, MN Burnett)
Merrit EA, Bacon DJ (1997) Methods Enzymol 277:505 (Implemented in WinGX (qv) and generated by ORTEP-III for Windows)
Macrae CF, Edgington PR, McCabe P, Pidcock E, Shields GP, Taylor R, Towler M, van de Streek J (2006) J Appl Cryst 39:453
Humphrey W, Dalke A, Schulten K (1996) J Mol Graphics 14.1:33
Ladd MFC, Palmer RA (2003) Structure determination by X-ray crystallography, 4th edn. Klewer-Plenum, NY, p 503
Berdy J, Aszalos A, Bostian M, McNitt K CRC handbook of antibiotic compounds, vol. IV(2), 319, CRC Press Inc., Boca Raton, Florida
Ramirez F, Marecek F, Shu-I TU, Kantor T, Okazaki H (1982) Eur J Biochem 121:275
Flack HD (1983) Acta Cryst A39:876
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Palmer, R.A., Potter, B.S. X-ray Structures and Absolute Configurations of the Antibiotics Oligomycins A, B and C: Inhibitors of ATP Synthase. J Chem Crystallogr 38, 243–253 (2008). https://doi.org/10.1007/s10870-008-9317-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10870-008-9317-y