Abstract
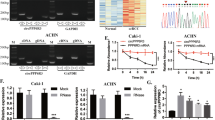
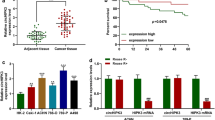
Clear cell renal cell carcinoma (ccRCC) is a prevalent urological carcinoma with high metastatic risk. Circular RNAs (circRNAs) have been identified as effective diagnostic and therapeutic biomarkers for ccRCC. This research aims to disclose the effect and regulatory mechanism of circRNA ribosomal protein L23a (circ_RPL23A) in ccRCC. We performed quantitative real-time polymerase chain reaction (qRT-PCR) to examine circ_RPL23A, microRNA-1233 (miR-1233) and acetyl-coenzyme A acetyltransferase 2 (ACAT2). Cell cycle progression, apoptosis, cell viability, invasion and migration, which were respectively conducted by using flow cytometry, 3-(4, 5-dimethylthiazol-2-y1)-2, 5-diphenyl tetrazolium bromide (MTT), transwell assays. The levels of ACAT2 protein and cell cycle proteins, proliferation-associated protein, and epithelial-mesenchymal transition (EMT) associated proteins were measured by western blot. Target relationship was analyzed via dual-luciferase reporter assay and RNA pull down assay. The animal model was used to study how circ_RPL23A affects in vivo. Circ_RPL23A was lower expressed in ccRCC tissues and cells. The elevated circ_RPL23A suppressed cell cycle progression, proliferation, migration and invasion but promoted apoptosis in ccRCC cells. MiR-1233 was a target of circ_RPL23A and direct targeted to ACAT2. Besides, circ_RPL23A exerted its anti-tumor effect by sponging miR-1233, and then relieved the inhibition effect of miR-1233 on ACAT2. Overexpression of circ_RPL23A also curbed ccRCC tumor growth in vivo. Circ_RPL23A inhibited ccRCC progression by upregulating ACAT2 expression by competitively binding miR-1233, which might provide an in-depth cognition for ccRCC pathogenesis and circ_RPL23A might be a promising biomarker in ccRCC diagnosis and treatment.








Similar content being viewed by others
Abbreviations
- ccRCC:
-
Clear cell renal cell carcinoma
- ACAT2 :
-
acetyl-coenzyme A acetyltransferase 2
- circ_RPL23A :
-
circRNA ribosomal protein L23a
References
Barata P, Sood AK, Hong DS (2016) RNA-targeted therapeutics in cancer clinical trials: current status and future directions. Cancer Treat Rev 50:35–47. https://doi.org/10.1016/j.ctrv.2016.08.004
Cai H, Li Y, Niringiyumukiza JD, Su P, Xiang W (2019) Circular RNA involvement in aging: an emerging player with great potential. Mech Ageing Dev 178:16–24. https://doi.org/10.1016/j.mad.2018.11.002
Chen B, Huang S (2018) Circular RNA: an emerging non-coding RNA as a regulator and biomarker in cancer. Cancer Lett 418:41–50. https://doi.org/10.1016/j.canlet.2018.01.011
Chen T, Shao S, Li W, Liu Y, Cao Y (2019) The circular RNA hsa-circ-0072309 plays anti-tumour roles by sponging miR-100 through the deactivation of PI3K/AKT and mTOR pathways in the renal carcinoma cell lines. Artif Cells Nanomed Biotechnol 47:3638–3648. https://doi.org/10.1080/21691401.2019.1657873
Chen Q, Liu T, Bao Y, Zhao T, Wang J, Wang H, Wang A, Gan X, Wu Z, Wang L (2020a) CircRNA cRAPGEF5 inhibits the growth and metastasis of renal cell carcinoma via the miR-27a-3p/TXNIP pathway. Cancer Lett 469:68–77. https://doi.org/10.1016/j.canlet.2019.10.017
Chen Z, Xiao K, Chen S, Huang Z, Ye Y, Chen T (2020b) Circular RNA hsa_circ_001895 serves as a sponge of microRNA-296-5p to promote clear cell renal cell carcinoma progression by regulating SOX12. Cancer Sci 111:713–726. https://doi.org/10.1111/cas.14261
Dias F, Teixeira AL, Ferreira M, Adem B, Bastos N, Vieira J, Fernandes M, Sequeira MI, Maurício J, Lobo F, Morais A, Oliveira J, Kok K, and Medeiros R (2017). Plasmatic miR-210, miR-221 and miR-1233 profile: potential liquid biopsies candidates for renal cell carcinoma. Oncotarget 8, 103315-103326. https://doi.org/10.18632/oncotarget.21733
Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M (2016) CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 13:34–42. https://doi.org/10.1080/15476286.2015.1128065
Franz A, Ralla B, Weickmann S, Jung M, Rochow H, Stephan C, Erbersdobler A, Kilic E, Fendler A, Jung K (2019). Circular RNAs in clear cell renal cell carcinoma: their microarray-based identification, analytical validation, and potential use in a Clinico-genomic model to improve prognostic accuracy. Cancers (Basel) 11. https://doi.org/10.3390/cancers11101473
Hammond SM (2015) An overview of microRNAs. Adv Drug Deliv Rev 87:3–14. https://doi.org/10.1016/j.addr.2015.05.001
He YH, Chen C, Shi Z (2018) The biological roles and clinical implications of microRNAs in clear cell renal cell carcinoma. J Cell Physiol 233:4458–4465. https://doi.org/10.1002/jcp.26347
Hsieh JJ, Le V, Cao D, Cheng EH, Creighton CJ (2018) Genomic classifications of renal cell carcinoma: a critical step towards the future application of personalized kidney cancer care with pan-omics precision. J Pathol 244:525–537. https://doi.org/10.1002/path.5022
Huang Y, Zhang Y, Jia L, Liu C, Xu F (2019) Circular RNA ABCB10 promotes tumor progression and correlates with pejorative prognosis in clear cell renal cell carcinoma. Int J Biol Markers 34:176–183. https://doi.org/10.1177/1724600819842279
Ibrahim A, Shafie NH, Mohd Esa N, Shafie SR, Bahari H, and Abdullah MA (2020). Mikania micrantha extract inhibits HMG-CoA Reductase and ACAT2 and ameliorates hypercholesterolemia and lipid peroxidation in high cholesterol-fed rats. Nutrients 12:3077. https://doi.org/10.3390/nu12103077
Jiang HM, Wei JH, Zhang ZL, Fang Y, Zhou BF, Chen ZH, Lu J, Liao B, Zhou FJ, Luo JH, Chen W (2016) Does chromophobe renal cell carcinoma have better survival than clear cell renal cell carcinoma? A clinical-based cohort study and meta-analysis. Int Urol Nephrol 48:191–199. https://doi.org/10.1007/s11255-015-1161-3
Lee YT, Tan YJ, Oon CE (2018) Molecular targeted therapy: treating cancer with specificity. Eur J Pharmacol 834:188–196. https://doi.org/10.1016/j.ejphar.2018.07.034
Lee BK, Kim MH, Lee SY, Son SJ, Hong CH, and Jung YS (2020). Downregulated platelet miR-1233-5p in patients with Alzheimer's pathologic change with mild cognitive impairment is associated with Aβ-induced platelet activation via P-Selectin. J Clin Med 9:1642. https://doi.org/10.3390/jcm9061642
Li QK, Pavlovich CP, Zhang H, Kinsinger CR, Chan DW (2019) Challenges and opportunities in the proteomic characterization of clear cell renal cell carcinoma (ccRCC): a critical step towards the personalized care of renal cancers. Semin Cancer Biol 55:8–15. https://doi.org/10.1016/j.semcancer.2018.06.004
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Ljungberg B, Campbell SC, Choi HY, Jacqmin D, Lee JE, Weikert S, Kiemeney LA (2011) The epidemiology of renal cell carcinoma. Eur Urol 60:615–621. https://doi.org/10.1016/j.eururo.2011.06.049
Ma C, Qin J, Zhang J, Wang X, Wu D, Li X (2020) Construction and analysis of circular RNA molecular regulatory networks in clear cell renal cell carcinoma. Mol Med Rep 21:141–150. https://doi.org/10.3892/mmr.2019.10811
Pavlova NN, Thompson CB (2016) The emerging hallmarks of Cancer metabolism. Cell Metab 23:27–47. https://doi.org/10.1016/j.cmet.2015.12.006
Peng L, Han L, Li XN, Miao YF, Xue F, Zhou C (2020) The predictive value of microRNA-134 and microRNA-1233 for the early diagnosis of acute exacerbation of chronic obstructive pulmonary disease with acute pulmonary embolism. Int J Chron Obstruct Pulmon Dis 15:2495–2503. https://doi.org/10.2147/copd.s266021
Qiu Y, Pu C, Li Y, Qi B (2020) Construction of a circRNA-miRNA-mRNA network based on competitive endogenous RNA reveals the function of circRNAs in osteosarcoma. Cancer Cell Int 20:48. https://doi.org/10.1186/s12935-020-1134-1
Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, Sun W, Dou K, Li H (2015) Circular RNA: a new star of noncoding RNAs. Cancer Lett 365:141–148. https://doi.org/10.1016/j.canlet.2015.06.003
Shintani S, O'huigin C, Toyosawa S, Michalová V, Klein J (1999) Origin of gene overlap: the case of TCP1 and ACAT2. Genetics 152:743–754
Siegel RL, Miller KD, Jemal A (2017) Cancer statistics, 2017. CA Cancer J Clin 67:7–30. https://doi.org/10.3322/caac.21387
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30. https://doi.org/10.3322/caac.21442
Sun X, Ge X, Xu Z, Chen D (2020) Identification of circular RNA-microRNA-messenger RNA regulatory network in hepatocellular carcinoma by integrated analysis. J Gastroenterol Hepatol 35:157–164. https://doi.org/10.1111/jgh.14762
Wang K, Sun Y, Tao W, Fei X, Chang C (2017a) Androgen receptor (AR) promotes clear cell renal cell carcinoma (ccRCC) migration and invasion via altering the circHIAT1/miR-195-5p/29a-3p/29c-3p/CDC42 signals. Cancer Lett 394:1–12. https://doi.org/10.1016/j.canlet.2016.12.036
Wang ZY, Zhong T, Wang Y, Song FM, Yu XF, Xing LP, Zhang WY, Yu JH, Hua SC, Yu XF (2017b) Human Enterovirus 68 interferes with the host cell cycle to facilitate viral production. Front Cell Infect Microbiol 7:29. https://doi.org/10.3389/fcimb.2017.00029
Wang G, Xue W, Jian W, Liu P, Wang Z, Wang C, Li H, Yu Y, Zhang D, Zhang C (2018) The effect of Hsa_circ_0001451 in clear cell renal cell carcinoma cells and its relationship with clinicopathological features. J Cancer 9:3269–3277. https://doi.org/10.7150/jca.25902
Wu G, Zhou W, Lin X, Sun Y, Li J, Xu H, Shi P, Gao L, Tian X (2020) circRASSF2 acts as ceRNA and promotes papillary thyroid carcinoma progression through miR-1178/TLR4 signaling pathway. Mol Ther Nucleic Acids 19:1153–1163. https://doi.org/10.1016/j.omtn.2019.11.037
Wulfken LM, Moritz R, Ohlmann C, Holdenrieder S, Jung V, Becker F, Herrmann E, Walgenbach-Brünagel G, Von Ruecker A, Müller SC, Ellinger J (2011) MicroRNAs in renal cell carcinoma: diagnostic implications of serum miR-1233 levels. PLoS One 6:e25787. https://doi.org/10.1371/journal.pone.0025787
Xu W, Zhou B, Wu J, Jiang P, Chen H, Yan F (2020) Circular RNA hsa-circ-0007766 modulates the progression of gastric carcinoma via miR-1233-3p/GDF15 axis. Int J Med Sci 17:1569–1583. https://doi.org/10.7150/ijms.46261
Xue D, Wang H, Chen Y, Shen D, Lu J, Wang M, Zebibula A, Xu L, Wu H, Li G, Xia L (2019) Circ-AKT3 inhibits clear cell renal cell carcinoma metastasis via altering miR-296-3p/E-cadherin signals. Mol Cancer 18:151. https://doi.org/10.1186/s12943-019-1072-5
Yadav S, Khandelwal M, Seth A, Saini AK, Dogra PN, Sharma A (2017) Serum microRNA expression profiling: potential diagnostic implications of a panel of serum microRNAs for clear cell renal cell Cancer. Urology 104:64–69. https://doi.org/10.1016/j.urology.2017.03.013
Yin Y, Long J, He Q, Li Y, Liao Y, He P, Zhu W (2019) Emerging roles of circRNA in formation and progression of cancer. J Cancer 10:5015–5021. https://doi.org/10.7150/jca.30828
Yu J, Zhang L, Ren P, Zhong T, Li Z, Wang Z, Li J, Liu X, Zhao K, Zhang W, Yu XF (2015) Enterovirus 71 mediates cell cycle arrest in S phase through non-structural protein 3D. Cell Cycle 14:425–436. https://doi.org/10.4161/15384101.2014.980631
Zhao ZJ, Shen J (2017) Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol 14:514–521. https://doi.org/10.1080/15476286.2015.1122162
Zhao Z, Lu J, Han L, Wang X, Man Q, Liu S (2016) Prognostic significance of two lipid metabolism enzymes, HADHA and ACAT2, in clear cell renal cell carcinoma. Tumour Biol 37:8121–8130. https://doi.org/10.1007/s13277-015-4720-4
Availability of data and materials
The data that support the findings of this study are available on request from the corresponding author.
Funding
This work was supported by Provincial Department of Education Fundamental Research Operating Expenses of Basic Research Projects (Grant No. 2018-KYYWF-0968).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interest.
Ethics approval and consent to participate
The design of this protocol follows the tenets of the Declaration of Helsinki, approved by the Ethics Committee of The First Affiliated Hospital of Jiamusi University.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Fig. 1
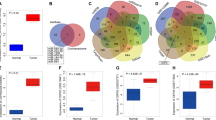
Linear RPL23A was downregulated in ccRCC samples. The qRT-PCR was performed to determine the expression of RPL23A in 60 ccRCC and normal samples. Data are reported as mean ± SD.*P < 0.05 vs normal para-carcinoma tissues. (PNG 2780 kb)
Supplementary Fig. 2
Circ_RPL23A depletion accelerated tumor progression in ccRCC cells. (A-B) The transfection efficiency was assessed by qRT-PCR after 786-O and 769-P cells were transfected with si-circ_RPL23A or si-NC. (C-D) Cell cycle progression in transfected cells was detected by PI-flow cytometry assay. (E-F) Western blot assay was administrated for determining cell cycle proteins CyclinD1 and CDK4 upon transfection. (G-H) Cell viability upon transfection was examined by MTT assay. (I-J) The protein levels of cell proliferation marker (Ki67, PCNA) upon transfection were detected by western blot. (K) The apoptosis rate of transfected cells was examined using Annexin V/PI-flow cytometry. (L-M) Transwell assay was used to measure the migration and invasion of transfected cells. (N-O) The determination of snail and E-cadherin proteins was carried out via western blot. Data are reported as mean ± SD, n = 3.*P < 0.05 vs si-NC group (ANOVA). (PNG 28450 kb)
Supplementary Fig. 3
The direct interaction between circ_RPL23A and miR-1233. RNA pull down assay was employed to analyze the enrichment of miR-1233 in 786-O and 769-P cells with biotin-labeled circ_RPL23A-WT or circ_RPL23A-MUT probe. Data are reported as mean ± SD, n = 3. *P < 0.05 vs bio-NC group (ANOVA). (PNG 1379 kb)
Rights and permissions
About this article
Cite this article
Cheng, L., Cao, H., Xu, J. et al. Circ_RPL23A acts as a miR-1233 sponge to suppress the progression of clear cell renal cell carcinoma by promoting ACAT2. J Bioenerg Biomembr 53, 415–428 (2021). https://doi.org/10.1007/s10863-021-09901-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-021-09901-8