Abstract
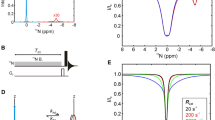
Exchange between conformational states is required for biomolecular catalysis, allostery, and folding. A variety of NMR experiments have been developed to quantify motional regimes ranging from nanoseconds to seconds. In this work, we describe an approach to speed up the acquisition of chemical exchange saturation transfer (CEST) experiments that are commonly used to probe millisecond to second conformational exchange in proteins and nucleic acids. The standard approach is to obtain CEST datasets through the acquisition of a series of 2D correlation spectra where each experiment utilizes a single saturation frequency to 1H, 15N or 13C. These pseudo 3D datasets are time consuming to collect and are further lengthened by reduced signal to noise stemming from the long saturation pulse. In this article, we show how usage of a multiple frequency saturation pulse (i.e., MF-CEST) changes the nature of data collection from series to parallel, and thus decreases the total acquisition time by an integer factor corresponding to the number of frequencies in the pulse. We demonstrate the applicability of MF-CEST on a Src homology 2 (SH2) domain from phospholipase Cγ and the secondary active transport protein EmrE as model systems by collecting 13C methyl and 15N backbone datasets. MF-CEST can also be extended to additional sites within proteins and nucleic acids. The only notable drawback of MF-CEST as applied to backbone 15N experiments occurs when a large chemical shift difference between the major and minor populations is present (typically greater than ~ 8 ppm). In these cases, ambiguity may arise between the chemical shift of the minor population and the multiple frequency saturation pulse. Nevertheless, this drawback does not occur for methyl group MF-CEST experiments or in cases where somewhat smaller chemical shift differences occur are present.




Similar content being viewed by others
References
Akyuz N, Altman RB, Blanchard SC, Boudker O (2013) Transport dynamics in a glutamate transporter homologue. Nature 502:114
Banigan JR, Leninger M, Her AS, Traaseth NJ (2018) Assessing interactions between a polytopic membrane protein and lipid bilayers using differential scanning calorimetry and solid-state NMR. J Phys Chem B 122:2314–2322
Boutin C, Leonce E, Brotin T, Jerschow A, Berthault P (2013) Ultrafast Z-spectroscopy for (129)Xe NMR-based sensors. J Phys Chem Lett 4:4172–4176
Bouvignies G, Kay LE (2012) A 2D (1)(3)C-CEST experiment for studying slowly exchanging protein systems using methyl probes: an application to protein folding. J Biomol NMR 53:303–310
Chen YJ et al (2007) X-ray structure of EmrE supports dual topology model. Proc Natl Acad Sci USA 104:18999–19004
Cho MK, Gayen A, Banigan JR, Leninger M, Traaseth NJ (2014) Intrinsic conformational plasticity of native EmrE provides a pathway for multidrug resistance. J Am Chem Soc 136:8072–8080
Delaforge E et al (2015) Large-scale conformational dynamics control H5N1 influenza polymerase PB2 binding to import in alpha. J Am Chem Soc 137:15122–15134
Farrow NA, Zhang O, Forman-Kay JD, Kay LE (1994) A heteronuclear correlation experiment for simultaneous determination of 15N longitudinal decay and chemical exchange rates of systems in slow equilibrium. J Biomol NMR 4:727–734
Fawzi NL, Ying J, Ghirlando R, Torchia DA, Clore GM (2011) Atomic-resolution dynamics on the surface of amyloid-beta protofibrils probed by solution NMR. Nature 480:268–272
Fizil A, Gaspari Z, Barna T, Marx F, Batta G (2015) “Invisible” conformers of an antifungal disulfide protein revealed by constrained cold and heat unfolding, CEST-NMR experiments, and molecular dynamics calculations. Chemistry 21:5136–5144
Gayen A, Banigan JR, Traaseth NJ (2013) Ligand-induced conformational changes of the multidrug resistance transporter EmrE probed by oriented solid-state NMR spectroscopy. Angew Chem Int Ed Engl 52:10321–10324
Gayen A, Leninger M, Traaseth NJ (2016) Protonation of a glutamate residue modulates the dynamics of the drug transporter EmrE. Nat Chem Biol 12:141–145
Gladkova C et al (2017) An invisible ubiquitin conformation is required for efficient phosphorylation by PINK1. EMBO J 36:3555–3572
Gutowsky HS, Saika A (1953) Dissociation, chemical exchange, and the proton magnetic resonance in some aqueous electrolytes. J Chem Phys 21:1688–1694
Hansen DF, Led JJ (2006) Determination of the geometric structure of the metal site in a blue copper protein by paramagnetic NMR. Proc Natl Acad Sci USA 103:1738–1743
Huang Z et al (2016) Two FGF receptor kinase molecules act in concert to recruit and transphosphorylate phospholipase Cgamma. Mol Cell 61:98–110
Korzhnev DM, Kloiber K, Kanelis V, Tugarinov V, Kay LE (2004) Probing slow dynamics in high molecular weight proteins by methyl-TROSY NMR spectroscopy: application to a 723-residue enzyme. J Am Chem Soc 126:3964–3973
Lee JS, Regatte RR, Jerschow A (2012) Isolating chemical exchange saturation transfer contrast from magnetization transfer asymmetry under two-frequency rf irradiation. J Magn Reson 215:56–63
Levitt MH (1982) Symmetrical composite pulse sequences for NMR population-inversion. 2. Compensation of resonance offset. J Magn Reson 50:95–110
Li Y, Palmer AG III (2009) TROSY-selected ZZ-exchange experiment for characterizing slow chemical exchange in large proteins. J Biomol NMR 45:357 – 60
Long D, Delaglio F, Sekhar A, Kay LE (2015) Probing invisible, excited protein states by non-uniformly sampled pseudo-4D CEST spectroscopy. Angew Chem Int Ed 54:10507–10511
Loria JP, Rance M, Palmer AG III (1999) A relaxation-compensated Carr–Purcell–Meiboom–Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Ma RS et al (2016) Determination of pseudocontact shifts of low-populated excited states by NMR chemical exchange saturation transfer. Phys Chem Chem Phys 18:13794–13798
Mangia S, Traaseth NJ, Veglia G, Garwood M, Michaeli S (2010) Probing slow protein dynamics by adiabatic R(1rho) and R(2rho) NMR experiments. J Am Chem Soc 132:9979–9981
Massi F, Grey MJ, Palmer AG III (2005) Microsecond timescale backbone conformational dynamics in ubiquitin studied with NMR R1rho relaxation experiments. Prot Sci 14:735–742
Matsuki Y, Konuma T, Fujiwara T, Sugase K (2011) Boosting protein dynamics studies using quantitative nonuniform sampling NMR spectroscopy. J Phys Chem B 115:13740–13745
Milojevic J, Esposito V, Das R, Melacini G (2007) Understanding the molecular basis for the inhibition of the Alzheimer’s Abeta-peptide oligomerization by human serum albumin using saturation transfer difference and off-resonance relaxation NMR spectroscopy. J Am Chem Soc 129:4282–4290
Mittermaier AK, Kay LE (2009) Observing biological dynamics at atomic resolution using NMR. Trends Biochem Sci 34:601–611
Montelione GT, Wagner G (1989) 2d chemical-exchange NMR-spectroscopy by proton-detected heteronuclear correlation. J Am Chem Soc 111:3096–3098
Morrison EA et al (2012) Antiparallel EmrE exports drugs by exchanging between asymmetric structures. Nature 481:45–50
Palmer AG III, Massi F (2006) Characterization of the dynamics of biomacromolecules using rotating-frame spin relaxation NMR spectroscopy. Chem Rev 106:1700–1719
Proudfoot A, Frank AO, Ruggiu F, Mamo M, Lingel A (2016) Facilitating unambiguous NMR assignments and enabling higher probe density through selective labeling of all methyl containing amino acids. J Biomol NMR 65:15–27
Rennella E, Huang R, Velyvis A, Kay LE (2015) (13)CHD2-CEST NMR spectroscopy provides an avenue for studies of conformational exchange in high molecular weight proteins. J Biomol NMR 63:187–199
Rosen MK et al (1996) Selective methyl group protonation of perdeuterated proteins. J Mol Biol 263:627–636
Sahu D, Clore GM, Iwahara J (2007) TROSY-based z-exchange spectroscopy: application to the determination of the activation energy for intermolecular protein translocation between specific sites on different DNA molecules. J Am Chem Soc 129:13232–13237
Schuldiner S (2009) EmrE, a model for studying evolution and mechanism of ion-coupled transporters. Biochim Biophys Acta 1794:748 –762
Sekhar A et al (2015) Thermal fluctuations of immature SOD1 lead to separate folding and misfolding pathways. Elife 4:e07296
Tang C, Louis JM, Aniana A, Suh JY, Clore GM (2008) Visualizing transient events in amino-terminal autoprocessing of HIV-1 protease. Nature 455:693-U92
Tate CG, Ubarretxena-Belandia I, Baldwin JM (2003) Conformational changes in the multidrug transporter EmrE associated with substrate binding. J Mol Biol 332:229–242
Traaseth NJ et al (2012) Heteronuclear Adiabatic Relaxation Dispersion (HARD) for quantitative analysis of conformational dynamics in proteins. J Magn Reson 219:75–82
Tugarinov V, Kay LE (2005) Methyl groups as probes of structure and dynamics in NMR studies of high-molecular-weight proteins. Chembiochem 6:1567–1577
Ubarretxena-Belandia I, Baldwin JM, Schuldiner S, Tate CG (2003) Three-dimensional structure of the bacterial multidrug transporter EmrE shows it is an asymmetric homodimer. EMBO J 22:6175–6181
Ulrich EL et al (2008) BioMagResBank. Nucleic Acids Res 36:D402-8
Vallurupalli P, Bouvignies G, Kay LE (2012) Studying “invisible” excited protein states in slow exchange with a major state conformation. J Am Chem Soc 134:8148–8161
Ward KM, Aletras AH, Balaban RS (2000) A new class of contrast agents for MRI based on proton chemical exchange dependent saturation transfer (CEST). J Magn Reson 143:79–87
Xu X, Lee JS, Jerschow A (2013) Ultrafast scanning of exchangeable sites by NMR spectroscopy. Angew Chem Int Ed Engl 52:8281–8284
Zhao B, Guffy SL, Williams B, Zhang Q (2017) An excited state underlies gene regulation of a transcriptional riboswitch. Nat Chem Biol 13:968–974
Zhou JY, van Zijl PCM (2006) PChemical exchange saturation transfer imaging and spectroscopy. Prog Nucl Magn Reson Spectrosc 48:109–136
Acknowledgements
The NMR methodology was supported by NSF award MCB 1506420 (to N.J.T.) and NIH Grant R01 EB016045 (to A.J.). Applications to EmrE and the SH2 domain were funded by NIH Grants R01 AI108889 and GM 117118, respectively. ML was supported from a Dean’s Dissertation Fellowship from New York University and W.M.M. acknowledges support from an NIH career transition award (F99 CA212474). All NMR data were collected with a cryoprobe at NYU that was supported by an NIH S10 Grant (OD016343). We thank Professor Moosa Mohammadi for the SH2 domain plasmid and Dr. Jae-Seung Lee for scientific discussions. We also thank Dr. Jae-Seung Lee for sharing a Python script for generating multiple frequency saturation pulses. This script is freely available from the link: https://github.com/jaeseung16/NMR_Bruker/blob/master/python/multifreq.py.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Leninger, M., Marsiglia, W.M., Jerschow, A. et al. Multiple frequency saturation pulses reduce CEST acquisition time for quantifying conformational exchange in biomolecules. J Biomol NMR 71, 19–30 (2018). https://doi.org/10.1007/s10858-018-0186-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-018-0186-1