Abstract
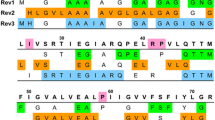
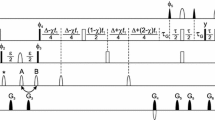
G-matrix FT projection NMR spectroscopy was employed for resonance assignment of the 79-residue subunit c of the Escherichia coli F1F0 ATP synthase embedded in micelles formed by lyso palmitoyl phosphatidyl glycerol (LPPG). Five GFT NMR experiments, that is, (3,2)D HNNCO, L-(4,3)D HNNC αβ C α, L-(4,3)D HNN(CO)C αβ C α, (4,2)D HACA(CO)NHN and (4,3)D HCCH, were acquired along with simultaneous 3D 15N, 13Caliphatic, 13Caromatic-resolved [1H,1H]-NOESY with a total measurement time of ∼43 h. Data analysis resulted in sequence specific assignments for all routinely measured backbone and 13Cβ shifts, and for 97% of the side chain shifts. Moreover, the use of two G2FT NMR experiments, that is, (5,3)D HN{N,CO}{C αβ C α} and (5,3)D {C αβ C α}{CON}HN, was explored to break the very high chemical shift degeneracy typically encountered for membrane proteins. It is shown that the 4D and 5D spectral information obtained rapidly from GFT and G2FT NMR experiments enables one to efficiently obtain (nearly) complete resonance assignments of membrane proteins.




Similar content being viewed by others
Abbreviations
- GFT:
-
G-matrix Fourier transformation
- LPPG:
-
1-Palmitoyl-2-hydroxy-sn-glycero-3-[phospho-RAC-(1-glycerol)]
- RAC:
-
Racemic
References
Atreya HS, Szyperski T (2004) G-matrix Fourier transform NMR spectroscopy for complete protein resonance assignment. Proc Natl Acad Sci USA 101:9642–9647
Atreya HS, Szyperski T (2005) Rapid NMR data collection. Methods Enzymol 394:78–108
Atreya HS, Eletsky A, Szyperski T (2005) Resonance assignment of proteins with high shift degeneracy based on 5D spectral information encoded in G2FT NMR experiments. J Am Chem Soc 127:4554–4555
Bartels C, Xia T, Billeter M, Guntert P, Wüthrich K (1995) The program XEASY for computer-supported NMR spectral analysis of biological macromolecules. J Biomol NMR 6:1–10
Cavanagh J, Fairbrother WJ, Palmer AG III, Rance M, Skelton NJ (2007) Protein NMR Spectroscopy. Elsevier Academic Press, San Diego, CA
Chill JH, Louis JM, Miller C, Bax A (2006) NMR study of the tetrameric KcsA potassium channel in detergent micelles. Protein Sci 15:684–698
Dmitriev OY, Fillingame RH (2001) Structure of Ala(20) → Pro/Pro(64) → Ala substituted subunit c of Escherichia coli ATP synthase in which the essential proline is switched between transmembrane helices. J Biol Chem 276:27449–27454
Dmitriev OY, Abildgaard F, Markley JL, Fillingame RH (2002) Structure of Ala24/Asp61 → Asp24/Asn61 substituted subunit c of Escherichia coli ATP synthase: implications for the mechanism of proton transport and rotary movement in the F0 complex. Biochemistry 41:5537–5547
Eghbalnia HR, Bahrami A, Tonelli M, Hallenga K, Markley JL (2005) High-resolution iterative frequency identification for NMR as a general strategy for multidimensional data collection. J Am Chem Soc 127:12528–12536
Eletsky A, Atreya HS, Liu G, Szyperski T (2005) Probing structure and functional dynamics of (large) proteins with aromatic rings: L-GFT-TROSY (4,3)D HCCH NMR spectroscopy. J Am Chem Soc 127:14578–14579
Fernandez C, Hilty C, Wider G, Güntert P, Wüthrich K (2004) NMR structure of the integral membrane protein OmpX. J Mol Biol 336:1211–1221
Gao FP, Cross TA (2005) Recent developments in membrane protein structural genomics. Genome Biol 6:244
Girvin ME, Rastogi VK, Abildgaard F, Markley JL, Fillingame RH (1998) Solution structure of the transmembrane H+-transporting subunit c of the F1F0 ATP synthase. Biochemistry 37:8817–8824
Granseth E, Seppälä S, Rapp M, Daley DO, Von Heijne G (2007) Membrane protein structural biology—how far can the bugs take us? Mol Membr Biol 24:329–332
Güntert P, Dötsch V, Wider G, Wüthrich K (1992) Processing of multi-dimensional NMR data with the new software PROSA. J Biomol NMR 2:619–629
Hiller S, Fiorito F, Wüthrich K, Wider G (2005) Automated projection spectroscopy (APSY). Proc Natl Acad Sci USA 102:10876–10881
Kim S, Szyperski T (2003) GFT NMR, a new approach to rapidly obtain precise high-dimensional NMR spectral information. J Am Chem Soc 125:1385–1393
Krueger-Koplin RD, Sorgen PL, Krueger-Koplin ST, Rivera-Torres AO, Cahill SM, Hicks DB, Grinius L, Krulwich TA, Girvin ME (2004) An evaluation of detergents for NMR structural studies of membrane proteins. J Biomol NMR 28:43–57
Kupce E, Freeman R (2004) Projection-reconstruction technique for speeding up multidimensional NMR spectroscopy. J Am Chem Soc 126:6429–6440
Liu GH, Shen Y, Atreya HS, Parish D, Shao Y, Sukumaran DK, Xiao R, Yee A, Lemak A, Bhattacharya A, Acton T, Arrowsmith C, Montelione G, Szyperski T (2005) NMR data collection and analysis protocol for high-throughput protein structure determination. Proc Natl Acad Sci USA 102:10487–10492
Lundstrom K (2007) Structural genomics and drug discovery. J Cell Mol Med 11: 224–238
Matthey U, Kaim G, Braun D, Wüthrich K, Dimroth P (1999) NMR studies of subunit c of the ATP synthase from Propionigenium modestum in dodecylsulphate micelles. Eur J Biochem 261:459–467
Matthey U, Braun D, Dimroth P (2002) NMR investigations of subunit c of the ATP synthase from Propionigenium modestum in chloroform/methanol/water (4:4:1). Eur J Biochem 269:1942–1946
Nakano T, Ikegami T, Suzuki T, Yoshida M, Akutsu H (2006) A new solution structure of ATP synthase subunit c from thermophilic Bacillus PS3, suggesting a local conformational change for H+-translocation. J Mol Biol 358:132–144
Norwood TJ, Crawford DA, Steventon ME, Driscoll PC, Campbell ID (1992) Heteronuclear 1H-15N nuclear magnetic resonance studies of the c subunit of the Escherichia coli F1F0 ATP synthase: assignment and secondary structure. Biochemistry 31:6285–6290
Rastogi VK, Girvin ME (1999a) 1H, 13C, and 15N assignments and secondary structure of the high pH form of subunit c of the F1F0 ATP synthase. J Biomol NMR 13:91–92
Rastogi VK, Girvin ME (1999b) Structural changes linked to proton translocation by subunit c of the ATP synthase. Nature 402:263–268
Rivera-Torres IO, Krueger-Koplin RD, Hicks DB, Cahill SM, Krulwich TA, Girvin ME (2004) pKa of the essential Glu54 and backbone conformation for subunit c from the H+-coupled F1F0 ATP synthase from an alkaliphilic Bacillus. FEBS Lett 575:131–135
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423
Shen Y, Atreya HS, Liu G, Szyperski T (2005) G-matrix Fourier transform NOESY based protocol for high-quality protein structure determination. J Am Chem Soc 127:9085–9099
Sorgen PL, Hu Y, Guan L, Kaback HR, Girvin ME (2002) An approach to membrane protein structure without crystals. Proc Natl Acad Sci USA 99:14037–14040
Spera S, Bax A (1991) Empirical correlation between protein backbone conformation and C-Alpha and C-Beta C-13 nuclear-magnetic-resonance chemical shifts. J Am Chem Soc 113:5490–5492
Szyperski T, Atreya HS (2006) Principles and applications of GFT projection NMR spectroscopy. Magn Reson Chem 44:S51–S60
Szyperski T, Yeh DC, Sukumaran DK, Moseley HNB, Montelione GT (2002) Reduced-dimensionality NMR spectroscopy for high-throughput protein resonance assignment. Proc Natl Acad Sci USA 99:8009–8014
Tamm LK, Abildgaard F, Arora A, Blad H, Bushweller JH (2003) Structure, dynamics and function of the outer membrane protein A (OmpA) and influenza hemagglutinin fusion domain in detergent micelles by solution NMR. FEBS Lett 555:139–143
Tourasse NJ, Li WH (2000) Selective constraints, amino acid composition, and the rate of protein evolution. Mol Biol Evol 17:656–664
Xia YL, Yee A, Arrowsmith CH, Gao XL (2003) H-1(C) and H-1(N) total NOE correlations in a single 3D NMR experiment. N-15 and C-13 time-sharing in t(1) and t(2) dimensions for simultaneous data acquisition. J Biomol NMR 27:193–203
Zhang HY, Neal S, Wishart DS (2003) RefDB: a database of uniformly referenced protein chemical shifts. J Biomol NMR 25:173–195
Acknowledgements
This work was supported by the National Institutes of Health (U54 GM074958-01) and the National Science Foundation (MCB 0416899). We thank Mr. Yu-Chieh Lin for help with the NMR sample preparation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Qi Zhang, Hanudatta S. Atreya, Douglas E. Kamen, Mark E. Girvin and Thomas Szyperski—New York Consortium on Membrane Protein Structure.
Rights and permissions
About this article
Cite this article
Zhang, Q., Atreya, H.S., Kamen, D.E. et al. GFT projection NMR based resonance assignment of membrane proteins: application to subunit c of E. coli F1F0 ATP synthase in LPPG micelles. J Biomol NMR 40, 157–163 (2008). https://doi.org/10.1007/s10858-008-9224-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-008-9224-8