Abstract
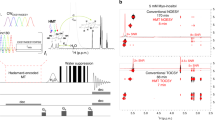
The paper presents a set of two-dimensional experiments that utilize direct 13C detection to provide proton–carbon, carbon–carbon and carbon–nitrogen correlations in the bases of nucleic acids. The set includes a 13C-detected proton–carbon correlation experiment for the measurement of 13C–13C couplings, the CaCb experiment for correlating two quaternary carbons, the HCaCb experiment for the 13C–13C correlations in cases where one of the carbons has a proton attached, the HCC-TOCSY experiment for correlating a proton with a network of coupled carbons, and a 13C-detected 13C–15N correlation experiment for detecting the nitrogen nuclei that cannot be detected via protons. The IPAP procedure is used for extracting the carbon–carbon couplings and/or carbon decoupling in the direct dimension, while the S3E procedure is preferred in the indirect dimension of the carbon–nitrogen experiment to obtain the value of the coupling constant. The experiments supply accurate values of 13C and 15N chemical shifts and carbon–carbon and carbon–nitrogen coupling constants. These values can help to reveal structural features of nucleic acids either directly or via induced changes when the sample is dissolved in oriented media.










Similar content being viewed by others
Abbreviations
- AP:
-
anti-phase
- DIPAP:
-
Double IPAP
- DSS:
-
4,4-dimethyl 4-silapentane sodium sulfonate
- HSQC:
-
Heteronuclear single-quantum correlation
- INEPT:
-
Insensitive nuclei enhancement by polarization transfer
- IP:
-
In-phase
- PCM:
-
Polarizable continuum model
- RDC:
-
Residual dipolar coupling
- S3E:
-
Spin-state selective excitation
References
Andersson P, Weigelt J, Otting G (1998) Spin-state selection filters for the measurement of heteronuclear one-bond coupling constants. J Biomol NMR 12:435–441
Balayssac S, Bertini I, Luchinat C, Parigi G, Piccioli M (2006) 13C direct detected NMR increases the detectability of residual dipolar couplings. J Am Chem Soc 128:15042–15043
Bermel W, Bertini I, Duma L, Felli IC, Emsley L, Pierattelli R, Vasos PR (2005a) Complete assignment of heteronuclear protein resonances by protonless NMR spectroscopy. Angew Chem Intl Ed 44:3089–3092
Bermel W, Bertini I, Felli IC, Pierattelli R, Vasos PR (2005b) A selective experiment for the sequential protein backbone assignment from 3D heteronuclear spectra. J Magn Reson 172:324–328
Bermel W, Bertini I, Felli IC, Kummerle R, Pierattelli R (2006a) Novel 13C direct detection experiments, including extension to the third dimension, to perform the complete assignment of proteins. J Magn Rerson 178:56–64
Bermel W, Bertini I, Felli IC, Lee YM, Luchinat C, Pierattelli R (2006b) Protonless NMR experiments for sequence-specific assignment of backbone nuclei in unfolded proteins. J Am Chem Soc 128:3918–3919
Bermel W, Bertini I, Felli IC, Piccioli M, Pierattelli R (2006c) 13C-detected protonless NMR spectroscopy of proteins in solution. Prog NMR Spectrosc 48:25–45
Boisbouvier J, Bryce DL, O’Neil-Cabello E, Nikonowicz EP, Bax A (2004) Resolution-optimized NMR measurement of 1DCH, 1DCC and 2DCH residual dipolar couplings in nucleic acid bases. J Biomol NMR 30:287–301
Brutscher B (2001) Accurate measurement of small spin-spin couplings in partially aligned molecules using a novel J-mismatch compensated spin-state-selection filter. J Magn Reson 151:332–338
Burum DP, Ernst RR (1980) Net polarization transfer via a J-ordered state for signal enhancement of low-sensitivity nuclei. J Magn Reson 39:163–168
Cromsigt J, Hilbers CW, Wijmenga SS (2001) Prediction of proton chemical shifts in RNA. Their use in structure refinement and validation. J Biomol NMR 21:11–29
Cromsigt J, Schleucher J, Gustafsson T, Kihlberg J, Wijmenga S (2002) Preparation of partially 2H/13C-labelled RNA for NMR studies. Stereo-specific deuteration of the H5″ in nucleotides. Nucleic Acids Res 30:1639–1645
Duma L, Hediger S, Lesage A, Emsley L (2003) Spin-state selection in solid-state NMR. J Magn Reson 164:187–195
Fiala R, Munzarová ML, Sklenář V (2004) Experiments for correlating quaternary carbons in RNA bases. J Biomol NMR 29:477–490
Fürtig B, Richter C, Wohnert J, Schwalbe H (2003) NMR spectroscopy of RNA. Chembiochem 4:936–962
Fürtig B, Richter C, Bermel W, Schwalbe H (2004) New NMR experiments for RNA nucleobase resonance assignment and chemical shift analysis of an RNA UUCG tetraloop. J Biomol NMR 28:69–79
Jaroniec CP, Boisbouvier J, Tworowska I, Nikonowicz EP, Bax A (2005) Accurate measurement of 15N–13C–13 residual dipolar couplings in nucleic acids. J Biomol NMR 31:231–241
Kadkhodaie M, Rivas O, Tan M, Mohebbi A, Shaka AJ (1991) Broad-band homonuclear cross polarization using flip-flop spectroscopy. J Magn Reson 91:437–443
Kovacs H, Moskau D, Spraul M (2005) Cryogenically cooled probes––a leap in NMR technology. Prog NMR Spectrosc 46:131–155
Meissner A, Duus JØ, Sørensen OW (1997a) Integration of spin-state-selective excitation into 2D NMR correlation experiments with heteronuclear ZQ/2Q p rotations for 1JXH-resolved E.COSY-type measurement of heteronuclear coupling constants in proteins. J Biomol NMR 10:89–94
Meissner A, Duus JØ, Sørensen OW (1997b) Spin-state-selective excitation. Application for E.COSY-type measurement of JHH coupling constants. J Magn Reson 128:92–97
Meissner A, Schulte-Herbruggen T, Sørensen OW (1998) Relaxation artifacts and their suppression in multidimensional E.COSY-type NMR experiments for measurement of J coupling constants in 13C- or 15N-labeled proteins. J Am Chem Soc 120:7989–7990
Mohebbi A, Shaka AJ (1991) Improvements in 13C broad-band homonuclear cross-polarization for 2D and 3D NMR. Chem Phys Lett 178:374–378
Mueller L, Legault P, Pardi A (1995) Improved RNA structure determination by detection of NOE contacts to exchange-broadened amino protons. J Am Chem Soc 117:11043–11048
Nikonowicz EP (2001). Preparation and use of 2H-labeled RNA oligonucleotides in nuclear magnetic resonance studies. Meth Enzymol 338:320–341
Ottiger M, Delaglio F, Bax A (1998) Measurement of J and dipolar couplings from simplified two-dimensional NMR spectra. J Magn Reson 131:373–378
Reiter NJ, Nikstad LJ, Allmann AM, Johnson RJ, Butcher SE (2003) Structure of the U6 RNA intramolecular stem-loop harboring an S-P-phosphorothioate modification. RNA 9:533–542
Riek R, Pervushin K, Fernandez C, Kainosho M, Wuthrich K (2001) [13C,13C]- and [13C,1H]-TROSY in a triple resonance experiment for ribose-base and intrabase correlations in nucleic acids. J Am Chem Soc 123:658–664
Serber Z, Richter C, Dotsch V (2001) Carbon-detected NMR experiments to investigate structure and dynamics of biological macromolecules. Chembiochem 2:247–251
Serber Z, Richter C, Moskau D, Bohlen JM, Gerfin T, Marek D, Haberli M, Baselgia L, Laukien F, Stern AS, Hoch JC, Dotsch V (2000) New carbon-detected protein NMR experiments using CryoProbes. J Am Chem Soc 122:3554–3555
Shaka AJ, Barker PB, Freeman R (1985) Computer-optimized decoupling scheme for wideband applications and low-level operation. J Magn Reson 64:547–552
Shimba N, Stern AS, Craik CS, Hoch JC, Dotsch V (2003) Elimination of 13C alpha splitting in protein NMR spectra by deconvolution with maximum entropy reconstruction. J Am Chem Soc 125:2382–2383
Tolbert TJ, Williamson JR (1996) Preparation of specifically deuterated RNA for NMR studies using a combination of chemical and enzymatic synthesis. J Am Chem Soc 118:7929–7940
Tolbert TJ, Williamson JR (1997) Preparation of specifically deuterated and 13C-labeled RNA for NMR studies using enzymatic synthesis. J Am Chem Soc 119:12100–12108
Wijmenga SS, Kruithof M, Hilbers CW (1997) Analysis of 1H chemical shifts in DNA: assessment of the reliability of H-1 chemical shift calculations for use in structure refinement. J Biomol NMR 10:337–350
Wijmenga SS, van Buuren BNM (1998) The use of NMR methods for conformational studies of nucleic acids. Prog Nucl Magn Reson Spectrosc 32:287–387
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15N chemical-shift referencing in biomolecular NMR. J Biomol NMR 6:135–140
Žídek L, Wu HH, Feigon J, Sklenář V (2001) Measurement of small scalar and dipolar couplings in purine and pyrimidine bases. J Biomol NMR 21:153–160
Žídek L, Padrta P, Chmelík J, Sklenář V (2003) Internal consistency of NMR data obtained in partially aligned biomacromolecules. J Magn Reson 162:385–395
Acknowledgments
The work was supported by the grants MSM0021622413 to RF and LC06030 to VS provided by the Ministry of Education of the Czech Republic and by the FSG-V-RNA project of the 6th Framework Program of the European Union, Contract No. 503455. We are indebted to David Staple and Sam Butcher for the U6 ISL sample, to Wolfgang Bermel and Helena Kovacs for stimulating discussions and help in the implementation of the pulse sequences, and to Petr Novák for an assistance in the evaluation of scalar couplings.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fiala, R., Sklenář, V. 13C-detected NMR experiments for measuring chemical shifts and coupling constants in nucleic acid bases. J Biomol NMR 39, 153–163 (2007). https://doi.org/10.1007/s10858-007-9184-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-007-9184-4