Abstract
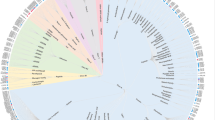
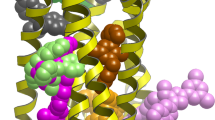
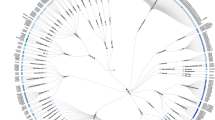
Targeting the allosteric sites on G-protein coupled receptors (GPCRs) for drug discovery is attracting increased interest. Given a GPCR target, identifying the allosteric binding sites in it remains a challenge. Previous works from our and other labs suggest the intracellular region below the middle of the transmembrane (TM) domain that spatially overlaps with the G-protein binding site could contain a common allosteric site for all GPCRs. We performed several bioinformatics analyses on this site for more than 100 representative human GPCR structures. Results of the studies confirmed that the proposed region contains an allosteric site that is druggable for 89% of the GPCRs and is not 100% identical between a GPCR and its most similar homolog for 94% of the GPCRs. The physico-chemical properties and amino acid composition of this site vary among and within GPCR classes. Since this proposed region occupies the space existing in all GPCRs of known structure, it could represent a common host of an allosteric site for all GPCRs that can be targeted for structure-based allosteric drug design.





Similar content being viewed by others
Data availability
All the data generated in this work is freely available in the supplementary materials or by contacting the corresponding author.
References
Congreve M, de Graaf C, Swain NA, Tate CG (2020) Impact of GPCR Structures on Drug Discovery. Cell 181(1):81–91
Foord SM, Bonner TI, Neubig RR, Rosser EM, Pin JP, Davenport AP, Spedding M, Harmar AJ (2005) International Union of pharmacology. XLVI. G protein-coupled receptor list. Pharmacol. Rev. 57(2):279–288
Sriram K, Insel PA (2018) G protein-coupled receptors as targets for approved drugs: how many targets and how many drugs? Mol Pharmacol 93(4):251–258
Topiol S (2018) Current and future challenges in GPCR drug discovery. Methods in Mol. Biol. 1705:1–21
Conn PJ, Christopoulos A, Lindsley CW (2009) Allosteric modulators of GPCRs: a novel approach for the treatment of CNS disorders. Nature Rev Drug Disco 8(1):41–54
Lütjens R, Rocher J-P (2017) Recent advances in drug discovery of GPCR allosteric modulators for neurodegenerative disorders. Curr Opin Pharmacol 32:91–95
Harrington PE, Fotsch C (2007) Calcium sensing receptor activators: calcimimetics. Curr Med Chem 14(28):3027–3034
Dorr P, Westby M, Dobbs S, Griffin P, Irvine B, Macartney M, Mori J, Rickett G, Smith-Burchnell C, Napier C, Webster R, Armour D, Price D, Stammen B, Wood A, Perros M (2005) Maraviroc (UK-427,857), a potent, orally bioavailable, and selective small-molecule inhibitor of chemokine receptor CCR5 with broad-spectrum anti-human immunodeficiency virus type 1 activity. Antimicrob Agents Chemother 49(11):4721–4732
Wootten D, Miller LJ, Koole C, Christopoulos A, Sexton PM (2016) Allostery and biased agonism at class B G protein-coupled receptors. Chem Rev 117(1):111–138
Pupo AS, Duarte DA, Lima V, Teixeira LB, Parreiras-E-Silva LT, Costa-Neto CM (2016) Recent updates on GPCR biased agonism. Pharmacol Res 112:49–57
Chan HCS, Li Y, Dahoun T, Vogel H, Yuan S (2019) New binding sites, new opportunities for GPCR drug discovery. Trends Biochem Sci 44(4):312–330
Thal DM, Glukhova A, Sexton PM, Christopoulos A (2018) Structural insights into G-protein-coupled receptor allostery. Nature 559(7712):45–53
Zheng Y, Qin L, Zacarias NV, de Vries H, Han GW, Gustavsson M, Dabros M, Zhao C, Cherney RJ, Carter P, Stamos D, Abagyan R, Cherezov V, Stevens RC, IJzerman, A. P., Heitman, L. H., Tebben, A., Kufareva, I., & Handel, T. M. (2016) Structure of CC chemokine receptor 2 with orthosteric and allosteric antagonists. Nature 540(7633):458–461
Oswald C, Rappas M, Kean J, Dore AS, Errey JC, Bennett K, Deflorian F, Christopher JA, Jazayeri A, Mason JS, Congreve M, Cooke RM, Marshall FH (2016) Intracellular allosteric antagonism of the CCR9 receptor. Nature 540(7633):462–465
Liu X, Ahn S, Kahsai AW, Meng KC, Latorraca NR, Pani B, Venkatakrishnan AJ, Masoudi A, Weis WI, Dror RO, Chen X, Lefkowitz RJ, Kobilka BK (2017) Mechanism of intracellular allosteric beta2AR antagonist revealed by X-ray crystal structure. Nature 548(7668):480–484
Song G, Yang D, Wang Y, de Graaf C, Zhou Q, Jiang S, Liu K, Cai X, Dai A, Lin G, Liu D, Wu F, Wu Y, Zhao S, Ye L, Han GW, Lau J, Wu B, Hanson MA et al (2017) Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature 546(7657):312–315
Liu K, Wu L, Yuan S, Wu M, Xu Y, Sun Q, Li S, Zhao S, Hua T, Liu Z-J (2020) Structural basis of CXC chemokine receptor 2 activation and signalling. Nature 585(7823):135–140
Redij T, Ma J, Li Z, Hua X, Li Z (2019) Discovery of a potential positive allosteric modulator of glucagon-like peptide 1 receptor through virtual screening and experimental study. J Comput Aided Mol Des 33(11):973–981
Pándy-Szekeres G, Esguerra M, Hauser AS, Caroli J, Munk C, Pilger S, Keserű GM, Kooistra AJ, Gloriam DE (2022) The G protein database, GproteinDb. Nucleic Acids Res 50(D1):D518–D525
Halgren T (2007) New method for fast and accurate binding-site identification and analysis. Chem Biol Drug Des 69(2):146–148
Hussein HA, Borrel A, Geneix C, Petitjean M, Regad L, Camproux A-C (2015) Pock drug-server: a new web server for predicting pocket druggability on holo and apo proteins. Nucleic Acids Res 43(W1):W436-442
UniProt Consortium (2019) UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res 47(D1):D506–D515
García-Nafría J, Tate CG (2020) Cryo-electron microscopy: moving beyond x-ray crystal structures for drug receptors and drug development. Annu Rev Pharmacol Toxicol 60:51–71
Sanchez-Reyes OB, Cooke ALG, Tranter DB, Rashid D, Eilers M, Reeves PJ, Smith SO (2017) G protein-coupled receptors contain two conserved packing clusters. Biophys J 112(11):2315–2326
Katritch V, Cherezov V, Stevens RC (2012) Diversity and modularity of G protein-coupled receptor structures. Trends Pharmacol Sci 33(1):17–27
Santos R, Ursu O, Gaulton A, Bento AP, Donadi RS, Bologa CG, Karlsson A, Al-Lazikani B, Hersey A, Oprea TI, Overington JP (2017) A comprehensive map of molecular drug targets. Nature Rev Drug Discov 16(1):19–34
Jumper J, Evans R, Pritzel A et al (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589
Syrovatkina V, Alegre KO, Dey R, Huang X-Y (2016) Regulation, signaling, and physiological functions of G-proteins. J Mol Biol 428(19):3850–3868
Fauman EB, Rai BK, Huang ES (2011) Structure-based druggability assessment—identifying suitable targets for small molecule therapeutics. Curr Opin Chem Biol 15(4):463–468
Hauser AS, Kooistra AJ, Munk C, Heydenreich FM, Veprintsev DB, Bouvier M, Babu MM, Gloriam DE (2021) GPCR activation mechanisms across classes and macro/microscales. Nature Struct Mol Biol 28(11):879–888
Acknowledgements
The authors thank Dr. Michael Bruist at University of the Sciences in Philadelphia for proofreading the manuscript and helpful comments. This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award Number [R15GM140406]. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Malik, F., Li, Z. Is there a common allosteric binding site for G-protein coupled receptors?. J Comput Aided Mol Des 36, 405–413 (2022). https://doi.org/10.1007/s10822-022-00454-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-022-00454-5