Abstract
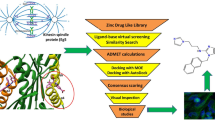
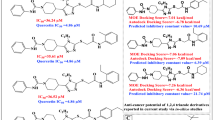
Eg5, a mitotic kinesin exclusively involved in the formation and function of the mitotic spindle has attracted interest as an anticancer drug target. Eg5 is co-crystallized with several inhibitors bound to its allosteric binding pocket. Each of these occupies a pocket formed by loop 5/helix α2 (L5/α2). Recently designed inhibitors additionally occupy a hydrophobic pocket of this site. The goal of the present study was to explore this hydrophobic pocket with our MED-SuMo fragment-based protocol, and thus discover novel chemical structures that might bind as inhibitors. The MED-SuMo software is able to compare and superimpose similar interaction surfaces upon the whole protein data bank (PDB). In a fragment-based protocol, MED-SuMo retrieves MED-Portions that encode protein-fragment binding sites and are derived from cross-mining protein-ligand structures with libraries of small molecules. Furthermore we have excluded intra-family MED-Portions derived from Eg5 ligands that occupy the hydrophobic pocket and predicted new potential ligands by hybridization that would fill simultaneously both pockets. Some of the latter having original scaffolds and substituents in the hydrophobic pocket are identified in libraries of synthetically accessible molecules by the MED-Search software.





Similar content being viewed by others
Abbreviations
- SCF:
-
Surface Chemical Feature
- PDB:
-
Protein Data Bank
- KSP:
-
Kinesin Spindle Protein
- HYD:
-
Hydrophobic
- L5/α2:
-
loop 5/helix α2
- Å:
-
Angström
- Du:
-
MED-Portion dummy atom
- ROC:
-
Receiver Operating Characteristic
References
Mitchison TJ, Salmon ED (2001) Nat Cell Biol 3:E17. doi:10.1038/35050656
Mitchison T, Kirschner M (1984) Nature 312:237. doi:10.1038/312237a0
Mitchison TJ (1989) J Cell Biol 109:637. doi:10.1083/jcb.109.2.637
Vale RD, Fletterick RJ (1997) Annu Rev Cell Dev Biol 13:745. doi:10.1146/annurev.cellbio.13.1.745
Amos LA, Cross RA (1997) Curr Opin Struct Biol 7:239. doi:10.1016/S0959-440X(97)80032-2
Wood KW, Cornwell WD, Jackson JR (2001) Curr Opin Pharmacol 1:370. doi:10.1016/S1471-4892(01)00064-9
Mayer TU, Kapoor TM, Haggarty SJ, King RW, Schreiber SL, Mitchison TJ (1999) Science 286:971. doi:10.1126/science.286.5441.971
Kwok BH, Kapoor TM (2007) Curr Opin Cell Biol 19:36. doi:10.1016/j.ceb.2006.12.003
Turner J, Anderson R, Guo J, Beraud C, Fletterick R, Sakowicz R (2001) J Biol Chem 276:25496. doi:10.1074/jbc.M100395200
Maliga Z, Kapoor TM, Mitchison TJ (2002) Chem Biol 9:989. doi:10.1016/S1074-5521(02)00212-0
Yan Y, Sardana V, Xu B, Homnick C, Halczenko W, Buser CA, Schaber M, Hartman GD, Huber HE, Kuo LC (2004) J Mol Biol 335:547. doi:10.1016/j.jmb.2003.10.074
Cox CD, Breslin MJ, Mariano BJ, Coleman PJ, Buser CA, Walsh ES, Hamilton K, Huber HE, Kohl NE, Torrent M, Yan Y, Kuo LC, Hartman GD (2005) Bioorg Med Chem Lett 15:2041. doi:10.1016/j.bmcl.2005.02.055
Cox CD, Torrent M, Breslin MJ, Mariano BJ, Whitman DB, Coleman PJ, Buser CA, Walsh ES, Hamilton K, Schaber MD, Lobell RB, Tao W, South VJ, Kohl NE, Yan Y, Kuo LC, Prueksaritanont T, Slaughter DE, Li C, Mahan E, Lu B, Hartman GD (2006) Bioorg Med Chem Lett 16:3175. doi:10.1016/j.bmcl.2006.03.040
Fraley ME, Steen JT, Brnardic EJ, Arrington KL, Spencer KL, Hanney BA, Kim Y, Hartman GD, Stirdivant SM, Drakas BA, Rickert K, Walsh ES, Hamilton K, Buser CA, Hardwick J, Tao W, Beck SC, Mao X, Lobell RB, Sepp-Lorenzino L, Yan Y, Ikuta M, Munshi SK, Kuo LC, Kreatsoulas C (2006) Bioorg Med Chem Lett 16:6049. doi:10.1016/j.bmcl.2006.08.118
Tarby CM, Kaltenbach RF3, Huynh T, Pudzianowski A, Shen H, Ortega-Nanos M, Sheriff S, Newitt JA, McDonnell PA, Burford N, Fairchild CR, Vaccaro W, Chen Z, Borzilleri RM, Naglich J, Lombardo LJ, Gottardis M, Trainor GL, Roussell DL (2006) Bioorg Med Chem Lett 16:2095. doi:10.1016/j.bmcl.2006.01.056
Kim KS, Lu S, Cornelius LA, Lombardo LJ, Borzilleri RM, Schroeder GM, Sheng C, Rovnyak G, Crews D, Schmidt RJ, Williams DK, Bhide RS, Traeger SC, McDonnell PA, Mueller L, Sheriff S, Newitt JA, Pudzianowski AT, Yang Z, Wild R, Lee FY, Batorsky R, Ryder JS, Ortega-Nanos M, Shen H, Gottardis M, Roussell DL (2006) Bioorg Med Chem Lett 16:3937. doi:10.1016/j.bmcl.2006.05.037
Garcia-Saez I, DeBonis S, Lopez R, Trucco F, Rousseau B, Thuery P, Kozielski F (2007) J Biol Chem 282:9740. doi:10.1074/jbc.M608883200
Jambon M, Imberty A, Deleage G, Geourjon C (2003) Proteins 52:137. doi:10.1002/prot.10339
Jambon M, Andrieu O, Combet C, Deleage G, Delfaud F, Geourjon C (2005) Bioinformatics 21:3929. doi:10.1093/bioinformatics/bti645
Doppelt O, Moriaud F, Bornot A, de Brevern AG (2007) Bioinformation 1:357
Moriaud F, Doppelt-Azeroual O, Martin L et al. (2009) J Chem Inf Model
Doppelt-Azeroual O, Moriaud F, Delfaud F (2009) Infect Disord Drug Targets
The PubChem Project (2008) http://pubchem.ncbi.nlm.nih.gov. Accessed 9 June 2008
Jambon M (2003) A bioinformatic system for searching functional similarities in 3D structures of proteins. Université Claude Bernard Lyon 1, Lyon
Chen X, Lin Y, Liu M, Gilson MK (2002) Bioinformatics 18:130. doi:10.1093/bioinformatics/18.1.130
Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK (2007) Nucleic Acids Res 35:D198. doi:10.1093/nar/gkl999
Nicholls A (2008) J Comput Aided Mol Des 22:239–255
Gehlhaar DK, Verkhivker GM, Rejto PA, Sherman CJ, Fogel DB, Fogel LJ, Freer ST (1995) Chem Biol 2:317. doi:10.1016/1074-5521(95)90050-0
Czerminski R (2005) Conference presentation, http://www.eyesopen.com/about/events/cup6/
Jenkins JL, Glick M, Davies JW (2004) J Med Chem 47:6144. doi:10.1021/jm049654z
Bemis GW, Murcko MA (1996) J Med Chem 39:2887. doi:10.1021/jm9602928
The Open Babel Package (2008) http://openbabel.sourceforge.net/. Accessed 5 July 2008
Accelrys Software Inc.: 10188 Telesis Court, Suite 100 San Diego, CA 92121, USA San Diego
Halgren TA (1999) J Comput Chem 20:720. doi:10.1002/(SICI)1096-987X(199905)20:7<720::AID-JCC7>3.0.CO;2-X
Brooks BR, Bruccoleri RE, Olafson BD et al. (1983) J Comp Chem 187
Gehlhaar DK, Bouzida D, Rejto PA (1992) American Chemical Society: Washington DC 292
Jain AN (1996) J Comput Aided Mol Des 10:427. doi:10.1007/BF00124474
Krammer A, Kirchhoff PD, Jiang X, Venkatachalam CM, Waldman M (2005) J Mol Graph Model 23:395. doi:10.1016/j.jmgm.2004.11.007
Muegge I (2006) J Med Chem 49:5895. doi:10.1021/jm050038s
Muegge I, Martin YC (1999) J Med Chem 42:791. doi:10.1021/jm980536j
Acknowledgments
We thank Raphaël Guerois for providing us useful comments and suggestions for the initiation of this study (Commissariat à l’Energie Atomique (CEA), Institut de Biologie et Technologies de Saclay, and Centre National de la Recherche Scientifique (CNRS), Gif-sur-Yvette, F-91191, France). Olivia Doppelt-Azeroual was funded by the ANRT. This work was supported by the Carriocas collaborative project (http://www.carriocas.org/) and funded by the French office “Direction Générale des Entreprises”.
Author information
Authors and Affiliations
Corresponding author
Additional information
Ksenia Oguievetskaia and Laetitia Martin-Chanas contributed equally to this work.
Rights and permissions
About this article
Cite this article
Oguievetskaia, K., Martin-Chanas, L., Vorotyntsev, A. et al. Computational fragment-based drug design to explore the hydrophobic sub-pocket of the mitotic kinesin Eg5 allosteric binding site. J Comput Aided Mol Des 23, 571–582 (2009). https://doi.org/10.1007/s10822-009-9286-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-009-9286-z