Abstract
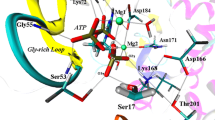
In the present work we have modeled the Michaelis complex of the cyclic-Adenosine Monophosphate Dependent (cAMD) Protein Kinase A (PKA) with Mg2ATP and the heptapeptide substrate Kemptide by classical molecular dynamics. The chosen synthetic substrate is relevant for its high efficiency and small size, and it has not been used in previous theoretical studies. The structural analysis of the data generated along the 6 ns simulation indicates that the modeled substrate–enzyme complex mimics the substrate binding pattern known for PKA. The values of the average prereactive distances obtained from the simulation do not exclude any of the two limiting situations proposed as mechanisms in the literature for the phosphorylation reaction (dissociative and associative) because the system oscillates between configurations compatible with each of them. Furthemore, the results obtained for the average interaction distances between active site residues concord in suggesting the plausability of an alternative third reaction mechanism.







Similar content being viewed by others
Notes
From now on, with the name PKA we will refer solely to the catalytic subunit of the enzyme and not to the entire tetramer.
The numbers of the residues of the substrates or inhibitor are referred to the peptide bound to the enzyme in the crystallographic structures. Thus the first residue of PKI(5–24), of SP20, and of the model substrate is Thr1, and the last one is Asp20 in place of Thr5 and Asp24, respectively. In the same way the first residue of Kemptide is Leu13 and the last one is Gly19.
References
Madhusudan, Akamine P, Xuong NH, Taylor SS (2002) Nat Struct Biol 9:273
Adams JA (2001) Chem Rev 101:2271
Cole PA, Courtney AD, Shen K, Zhang ZS, Qiao YF, Lu W, Williams DM (2003) Acc Chem Res 36:444
Johnson DA, Akamine P, Radzio-Andzelm E, Madhusudan, Taylor SS (2001) Chem Rev 101:2243
Gibbs CS, Zoller MJ (1991) J Biol Chem 266:8923
Bossemeyer D, Engh RA, Kinzel V, Ponstingl H, Huber R (1993) EMBO J 12:849
Gibbs CS, Zoller MJ (1991) Biochemistry 30:5329
Madhusudan, Trafny EA, Xuong NH, Adams JA, Teneyck LF, Taylor SS, Sowadski JM (1994) Protein Sci 3:176
Moore MJ, Adams JA, Taylor SS (2003) J Biol Chem 278:10613
Shabb JB (2001) Chem Rev 101:2381
Brinkworth RI, Horne J, Kobe B (2002) J Mol Recognit 15:104
Kemp BE, Graves DJ, Benjamini E, Krebs EG (1977) J Biol Chem 252:4888
Qamar R, Yoon M-Y, Cook PF (1992) Biochemistry 31:9986
Grant BD, Adams JA (1996) Biochemistry 35:2022
Kemp BE, Benjamini E, Krebs EG (1976) Proc Nat Acad Sci 73:1038
Diaz N, Field MJ (2004) J Am Chem Soc 126:529
Zhou J, Adams JA (1997) Biochemistry 36:2977
Williams DM, Cole PA (2002) J Am Chem Soc 124:5956
Cheng YH, Zhang YK, McCammon JA (2005) J Am Chem Soc 127:1553
Hart JC, Sheppard DW, Hillier IH, Burton NA (1999) Chem Comm 79
Blachut-Okrasinska E, Lesyng B, Briggs JM, McCammon JA, Antosiewicz JM (1999) Eur Biophys J 28:457
Sheppard DW, Burton N, Hillier IH (2000) J Mol Struct 506:35
Hutter MC, Helms V (2003) Int J Quant Chem 95:479
Valiev M, Kawai R, Adams JA, Weare JH (2003) J Am Chem Soc 125:9926
Cheng YH, Zhang YK, McCammon JA (2006) Protein Sci 15:672
Klahn M, Rosta E, Warshel A, Afklaehn M, Rosta E, Warshel A (2006) J Am Chem Soc 128:15310
Vanpatten SM, Ng DC, Thng JPH, Angelos KL, Smith AJ, Walsh DA (1991) Proc Nat Acad Sci 88:5383
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) J Comput Chem 4:187
MacKerell AD Jr, Bashford D, Bellott M Jr, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE III, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiórkiewicz-Kuczera J, Yin D, Karplus M (1998) J Phys Chem B 102:3586
Feng MH, Philippopoulos M, MacKerell AD, Lim C (1996) J Am Chem Soc 118:11265
Batkin M, Shaltiel ISS (2000) Biochemistry 39:5366
Shaltiel S, Cox S, Taylor SS (1998) Proc Nat Acad Sci 95:484
Mildvan AS (1997) Proteins 29:401
Granot J, Mildvan AS, Bramson HN, Kaiser ET (1980) Biochemistry 19:3537
Acknowledgments
We are grateful for financial support from the Spanish “Ministerio de Educación y Ciencia” and the “Fondo Europeo de Desarrollo Regional” through projects CTQ2005-07115/BQU and BIO2004-05879-C02-01, the “Generalitat de Catalunya” (2005SGR00400), and the “Universitat Autònoma de Barcelona” (EME2006-18). M.G.-V. thanks the “Ramon y Cajal” program for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Montenegro, M., Garcia-Viloca, M., González-Lafont, À. et al. Comparative study of the prereactive protein kinase A Michaelis complex with Kemptide substrate. J Comput Aided Mol Des 21, 603–615 (2007). https://doi.org/10.1007/s10822-007-9143-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-007-9143-x