Abstract
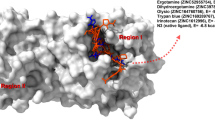
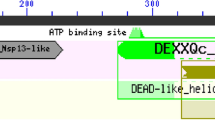
The modeling of the severe acute respiratory syndrome coronavirus helicase ATPase catalytic domain was performed using the protein structure prediction Meta Server and the 3D Jury method for model selection, which resulted in the identification of 1JPR, 1UAA and 1W36 PDB structures as suitable templates for creating a full atom 3D model. This model was further utilized to design small molecules that are expected to block an ATPase catalytic pocket thus inhibit the enzymatic activity. Binding sites for various functional groups were identified in a series of molecular dynamics calculation. Their positions in the catalytic pocket were used as constraints in the Cambridge structural database search for molecules having the pharmacophores that interacted most strongly with the enzyme in a desired position. The subsequent MD simulations followed by calculations of binding energies of the designed molecules were compared to ATP identifying the most successful candidates, for likely inhibitors—molecules possessing two phosphonic acid moieties at distal ends of the molecule.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Wu CH, Apweiler R, Bairoch A, Natale DA, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Mazumder R, O’Donovan C, Redaschi N, Suzek B (2006) Nucleic Acids Res 34(Database issue):D187–D191
Berman H, Henrick K, Nakamura H (2003) Nat Struct Biol 10(12):980
Neuman BW, Stein DA, Kroeker AD, Churchill MJ, Kim AM, Kuhn P, Dawson P, Moulton HM, Bestwick RK, Iversen PL, Buchmeier MJ (2005) J Virol 79(15):9665–9676
http://www.who.int/csr/resources/publications/WHO_CDS_ CSR_ARO_2004_1/en/index.html
Kliger Y, Levanon EY, Gerber D (2005) Drug Discov Today 10(5):345–352
Bujnicki JM, Elofsson A, Fischer D, Rychlewski L (2001) Bioinformatics 17(8):750–751
http://predictioncenter.org/
Ginalski K, Rychlewski L (2003) Proteins 53(Suppl 6):410–417
Ginalski K, Grishin NV, Godzik A, Rychlewski L (2005) Nucleic Acids Res 33(6):1874–1891
Sali A, Blundell TL (1993) J Mol Biol 234(3):779–815
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson JDM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117:5179–5197
Sali A, Potterton L, Yuan F, van Vlijmen H, Karplus M (1995) Proteins 23(3):318–326
Spector FC, Liang L, Giordano H, Sivaraja M, Peterson MG (1998) J Virol 72(9):6979–6987
Ravna AW, Schroder KE, Edvardsen O (1999) Comput Chem 23(5):435–437
Ljungberg KB, Marelius J, Musil D, Svensson P, Norden B, Aqvist J (2001) Eur J Pharm Sci 12(4):441–446
Egan WJ, Walters WP, Murcko MA (2002) Curr Opin Drug Discov Dev 5(4):540–549
Plewczynski D, Pas J, von Grotthuss M, Rychlewski L (2002) Appl Bioinf 1(4):223–225
Shen X, Xue JH, Yu CY, Luo HB, Qin L, Yu XJ, Chen J, Chen LL, Xiong B, Yue LD, Cai JH, Shen JH, Luo XM, Chen KX, Shi TL, Li YX, Hu GX, Jiang HL (2003) Acta Pharmacol Sin 24(6):505–511
Oda A, Yamaotsu N, Hirono S (2005) J Comput Chem 26(8):818–826
Jojart B, Martinek TA, Marki A (2005) J Comput Aided Mol Des 19(5):341–356
Park H, Lee S (2004) J Comput Aided Mol Des 18(6):375–388
Vreven T, Morokuma K, Farkas O, Schlegel HB, Frisch MJ (2003) J Comput Chem 24(6):760–769
Saen-oon S, Kuno M, Hannongbua S (2005) Proteins 61(4):859–869
Wieczorek R, Dannenberg JJ (2005) J Am Chem Soc 127(42):14534–14535
Hoffmann M, Khavrutskii IV, Musaev DG, Morokuma K (2004) Int J Quant Chem 99(6):972–980
Burgoyne NJ, Jackson RM (2006) Bioinformatics 22(11):1335–1342
Nemoto T, Fedorov DG, Uebayasi M, Kanazawa K, Kitaura K, Komeiji Y (2005) Comput Biol Chem 29(6):434–439
Klingenberg M (2005) Biochemistry 44(24):8563–8570
Waldron TT, Schrift GL, Murphy KP (2005) J Mol Biol 346(3):895–905
Nissink JW, Murray C, Hartshorn M, Verdonk ML, Cole JC, Taylor R (2002) Proteins 49(4):457–471
Swegat W, Schlitter J, Kruger P, Wollmer A (2003) Biophys J 84(3):1493–1506
Ginalski K, Pas J, Wyrwicz LS, von Grotthuss M, Bujnicki JM, Rychlewski L (2003) Nucleic Acids Res 31(13):3804–3807
Karplus K, Karchin R, Barrett C, Tu S, Cline M, Diekhans M, Grate L, Casper J, Hughey R (2001) Proteins 5(Suppl): 86–91
Jaroszewski L, Rychlewski L, Li Z, Li W, Godzik A (2005) Nucleic Acids Res 33(Web Server issue):W284–W288
Jones DT (1999) J Mol Biol 287(4):797–815
Fischer, D (2000) Pac Symp Biocomput 5:119–130
Shi J, Blundell TL, Mizuguchi K (2001) J Mol Biol 310(1):243–257
Kelley LA, MacCallum RM, Sternberg MJ (2000) J Mol Biol 299(2):499–520
Hogbom M, Andersson ME, Nordlund P (2001) J Biol Inorg Chem 6(3):315–323
Korolev S, Hsieh J, Gauss GH, Lohman TM, Waksman G (1997) Cell 90(4):635–647
Singleton MR, Dillingham MS, Gaudier M, Kowalczykowski SC, Wigley DB (2004) Nature 432(7014):187–193
Eswar N, John B, Mirkovic N, Fiser A, Ilyin VA, Pieper U, Stuart AC, Marti-Renom MA, Madhusudhan MS, Yerkovich B, Sali A (2003) Nucleic Acids Res 31(13):3375–3380
Marti-Renom MA, Stuart AC, Fiser A, Sanchez R, Melo F, Sali A (2000) Annu Rev Biophys Biomol Struct 29:291–325
Fox T, Kollman PA (1996) Proteins 25(3):315–334
Cheatham TE, Cieplak P, Kollman PA (1999) J Biomol Struct Dyn 16(4):845–862
Weiner SJ, Kollman PA, Nguyen DT, Case DA (1986) J␣Comput Chem 7(2):230–252
Cornell WDCP, Bayly CI, Gould IR, Merz KM, Ferguson DM Jr, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117:5179–5197
Ren P, Ponder JW (2003) J Phys Chem B 107:5933–5947
Ren P, Ponder JW (2002) J Comput Chem 23(16):1497–1506
Pappu RV, Hart RK, Ponder JW (1998) J Phys Chem B 102:9725–9742
Hodsdon ME, Ponder JW, Cistola DP (1996) J Mol Biol 264(3):585–602
Meagher KL, Redman LT, Carlson HA (2003) J Comput Chem 24(9):1016–1025
Taylor RD, Jewsbury PJ, Essex JW (2002) J Comput Aided Mol Des 16(3):151–166
Tanner JA, Watt RM, Chai YB, Lu LY, Lin MC, Peiris JS, Poon LL, Kung HF, Huang JD (2003) J Biol Chem 278(41):39578–39582
Joseph-McCarthy D, Alvarez JC (2003) Proteins 51(2):189–202
Miranker A, Karplus M (1995) Proteins 23(4):472–490
Caflisch A, Miranker A, Karplus M (1993) J Med Chem 36(15):2142–2167
van de Streek J (2006) Acta Crystallogr B 62(Pt 4):567–579
Allen FH (2002) Acta Crystallogr B 58(Pt 3 Pt 1):380–388
Allen FH, Motherwell WDS (2002) Acta Crystallogr B 58(Pt 3):407–422
CDS refcodes of the structures: ABITOU, AEADMP, AMAFAP, BACVAD, BAYHAK, BEBCEQ, BEJRAJ01, BERDIL, BERDOR, BIBTIP, BOLHEP, BUFDEL, BUNYOY, CACLAC, CAGUCP10, CEPBUU, CIDSUD, CITWUX10, CIZCUJ, COWDOH, COYDUP, CYACPH10, DEFLAB, DINYII10, DUMJEA10, DUVDON10, EHOWUT, EJETES, EVOYET, EWAHAL, FELSUK, FILCUY, FILDAF, GLCTSM, GLPCHO, GUPCYT20, HIBYIA, HUNBUN, ISOTEP, LAGXIA, LASDEO, LENCEM, LIHLET, LOTLEL, LUBLID, MAHNAK, NUKFII, PIDPHA10, QAZMIN, QEQNOP, QIXMEP, QURYAD, REBPET, SOYCUE, SPGLUC01, SUSBEN01, TAPFUL, THYTHY10, TICNIC, TOSDOU, TYRPX110, UKOWEW, XABSEZ, XOBMOQ, XUSBOC, YEJJUS, YITJOA
Cramer CJ, Truhlar DG (1999) Chem Rev 99(8):2161–2200
Frisch MJ, Schlegel GWTHB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA Jr, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMWB, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004) Gaussian, Inc, Wallingford
Dewar MJS, Zoebisch EG, Healy EF, Stewart JJP (1985) J␣Am Chem Soc 107(13):3902–3909
Dewar MJS, Reynolds CH (1986) J Comput Chem 7(2):140–143
Dewar MJS, Mckee ML, Rzepa HS (1978) J Am Chem Soc 100(11):3607–3607
Stewart JJP (1989) J Comput Chem 10(2):209–220
Stewart JJP (1989) J Comput Chem 10(2):221–264
Rota PA, Oberste MS, Monroe SS, Nix WA, Campagnoli R, Icenogle JP, Penaranda S, Bankamp B, Maher K, Chen MH, Tong S, Tamin A, Lowe L, Frace M, DeRisi JL, Chen Q, Wang D, Erdman DD, Peret TC, Burns C, Ksiazek TG, Rollin PE, Sanchez A, Liffick S, Holloway B, Limor J, McCaustland K, Olsen-Rasmussen M, Fouchier R, Gunther S, Osterhaus AD, Drosten C, Pallansch MA, Anderson LJ, Bellini WJ (2003) Science 300(5624):1394–1399
Ginalski K, Elofsson A, Fischer D, Rychlewski L (2003) Bioinformatics 19(8):1015–1018
Caruthers JM, McKay DB (2002) Curr Opin Struct Biol 12(1):123–133
Velankar SS, Soultanas P, Dillingham MS, Subramanya HS, Wigley DB (1999) Cell 97(1):75–84
Gorbalenya AE, Koonin EV (1993) Curr Opin Struct Biol 3:419–429
Walker JE, Saraste M, Runswick MJ, Gay NJ (1982) Embo J␣1(8):945–951
Hall MC, Matson SW (1999) Mol Microbiol 34(5):867–877
George JW, Brosh RM Jr, Matson SW (1994) J Mol Biol 235(2):424–435
Story RM, Steitz TA (1992) Nature 355(6358):374–376
Pause A, Methot N, Sonenberg N (1993) Mol Cell Biol 13(11):6789–6798
Ivanov KA, Thiel V, Dobbe JC, van der Meer Y, Snijder EJ, Ziebuhr J (2004) J Virol 78(11):5619–5632
Borowski P, Lang M, Niebuhr A, Haag A, Schmitz H, zur Wiesch JS, Choe J, Siwecka MA, Kulikowski T (2001) Acta Biochim Pol 48(3):739–744
Borowski P, Deinert J, Schalinski S, Bretner M, Ginalski K, Kulikowski T, Shugar D (2003) Eur J Biochem 270(8):1645–1653
Shin JM, Cho DH (2005) Nucleic Acids Res 33(Database issue):D238–241
Humphrey W, Dalke A, Schulten K (1996) J Mol Graph 14(1):33–38–27–38
Acknowledgment
Financial support from European committee grant no. SP22-CT-2004–003831 and Polish Ministry for Science is gratefully acknowledged. K. E. and M. v. G. thank the Foundation for Polish Science for a fellowship. Calculations were performed in Poznan Supercomputing and Networking Center (PCSS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hoffmann, M., Eitner, K., von Grotthuss, M. et al. Three dimensional model of severe acute respiratory syndrome coronavirus helicase ATPase catalytic domain and molecular design of severe acute respiratory syndrome coronavirus helicase inhibitors. J Comput Aided Mol Des 20, 305–319 (2006). https://doi.org/10.1007/s10822-006-9057-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-006-9057-z