Abstract
Background
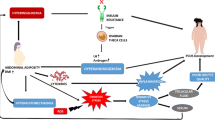
In recent years, alterations in lipid metabolism are currently considered a hallmark feature of many diseases. However, the role in women with reproductive dysfunction (WRD) remains to be fully elucidated. Here, this study aimed to explore the effect of lipid metabolism-related genes (LMRGs) on endometrial receptivity of WRD.
Methods
This study retrospectively analyzed endometrial receptivity array (ERA) in GEO database. The differential expression genes (DEGs) were obtained by limma differential analysis, and the core genes and corresponding predicted microRNA were obtained through protein–protein interaction (PPI) analysis and TargetScan database, so as to predict the chemical targets of drug therapy. Through the intersection of DEGs and LMRGs, the target gene expression profile was obtained for subsequent consensus clustering analysis and immune analysis. In addition, the immune cell infiltration was assessed by applying the ESTIMATE and MCPcounter algorithm and potential drug targets were obtained from the HERB website.
Results
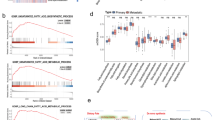
1473 genes showed differential expression between the groups of WRD and fertile women, and then a large number of lipid metabolism-related pathways and immune-related pathways were enriched. Twelve core genes and corresponding predicted miR-134-3p were obtained; most importantly, we found that these 12 genes were all LMRGs. Through drug target prediction, we obtained three drugs that regulate lipid metabolism and improve blood circulation, namely lovastatin, estrogen, and quercetin. EHHADH (AUC = 0.85) and PTEN (AUC = 0.82) have the best diagnostic performance. UMAP and heatmap revealed large differences between three clusters. LMRGs revealed specific manifestations of WRD in endometrial receptivity and immune microenvironment.
Conclusions
Our study explored the expression pattern of LMRGs in endometrium of WRD, screened the corresponding biomarkers, and proposed the combination of traditional Chinese and Western medicine to improve the endometrial receptivity.






Similar content being viewed by others
References
Wilcox AJ, Weinberg CR, O’Connor JF, Baird DD, Schlatterer JP, Canfield RE, et al. Incidence of early loss of pregnancy. N Engl J Med. 1988;319:189–94.
Diedrich K, Fauser BC, Devroey P, Griesinger G. Evian Annual Reproduction Workshop G. The role of the endometrium and embryo in human implantation. Hum Reprod Update. 2007;13:365–77.
Kasius A, Smit JG, Torrance HL, Eijkemans MJ, Mol BW, Opmeer BC, et al. Endometrial thickness and pregnancy rates after IVF: a systematic review and meta-analysis. Hum Reprod Update. 2014;20:530–41.
Strowitzki T, Germeyer A, Popovici R, von Wolff M. The human endometrium as a fertility-determining factor. Hum Reprod Update. 2006;12:617–30.
Altmae S, Esteban FJ, Stavreus-Evers A, Simon C, Giudice L, Lessey BA, et al. Guidelines for the design, analysis and interpretation of ‘omics’ data: focus on human endometrium. Hum Reprod Update. 2014;20:12–28.
Achache H, Revel A. Endometrial receptivity markers, the journey to successful embryo implantation. Hum Reprod Update. 2006;12:731–46.
Hernandez-Vargas P, Munoz M, Dominguez F. Identifying biomarkers for predicting successful embryo implantation: applying single to multi-OMICs to improve reproductive outcomes. Hum Reprod Update. 2020;26:264–301.
Bulun SE. Aromatase and estrogen receptor alpha deficiency. Fertil Steril. 2014;101:323–9.
Kang HJ, Imperato-McGinley J, Zhu YS, Rosenwaks Z. The effect of 5alpha-reductase-2 deficiency on human fertility. Fertil Steril. 2014;101:310–6.
Rosenwaks Z, Adashi EY. Introduction. Fertility in the face of genetically determined steroidogenic dysfunction. Fertil Steril. 2014;101:299–300.
DeAngelis AM, Roy-O’Reilly M, Rodriguez A. Genetic alterations affecting cholesterol metabolism and human fertility. Biol Reprod. 2014;91:117.
Taminau J, Meganck S, Lazar C, Steenhoff D, Coletta A, Molter C, et al. Unlocking the potential of publicly available microarray data using inSilicoDb and inSilicoMerging R/Bioconductor packages. BMC Bioinformatics. 2012;13:335.
Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8:118–27.
Jassal B, Matthews L, Viteri G, Gong C, Lorente P, Fabregat A, et al. The reactome pathway knowledgebase. Nucleic Acids Res. 2020;48:D498–503.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50.
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45:D362–8.
Wilkerson MD, Hayes DN. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26:1572–3.
Zeng D, Ye Z, Shen R, Yu G, Wu J, Xiong Y, et al. IOBR: multi-omics immuno-oncology biological research to decode tumor microenvironment and signatures. Front Immunol. 2021;12:687975.
Yoshihara K, Shahmoradgoli M, Martinez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612.
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F, et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016;17:218.
Horvath S, Dong J. Geometric interpretation of gene coexpression network analysis. PLoS Comput Biol. 2008;4:e1000117.
Karizbodagh MP, Rashidi B, Sahebkar A, Masoudifar A, Mirzaei H. Implantation window and angiogenesis. J Cell Biochem. 2017;118:4141–51.
Author information
Authors and Affiliations
Contributions
SZ: conceptualization.
YL: data curation, methodology, investigation, writing—review and editing.
YY, HS: methodology, validation.
JZ, HL: data curation, investigation.
SW, TZ, MM: data curation, investigation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., Yao, Y., Sun, H. et al. Lipid metabolism-related genes as biomarkers and therapeutic targets reveal endometrial receptivity and immune microenvironment in women with reproductive dysfunction. J Assist Reprod Genet 39, 2179–2190 (2022). https://doi.org/10.1007/s10815-022-02584-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-022-02584-z