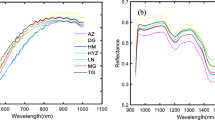
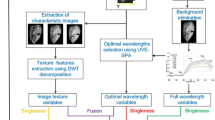
Taking the fat content of six different brands of milk as the research object, hyperspectral reflectance data were obtained using hyperspectral imaging and image processing techniques. The raw data are preprocessed in seven different ways. Combine the three mature bionic algorithms — genetic algorithm (GA), ant colony optimization, and particle swarm optimization — with partial least squares (PLS) and support vector machine regression (SVR) models to filter characteristic bands, and to explore the linear and nonlinear relationship between milk spectral data and fat content. The correlation coefficient method, the uninformative elimination algorithm, the successive projection algorithm, the competitive adaptive reweighting sampling algorithm, four mature feature band selection methods, are compared with the bionic algorithm, and, according to the characteristics of each, are combined. The best combination of characteristic bands is selected to establish a regression model to detect milk fat content accurately. The band combination screened by GA and PLS achieved the best prediction results. A total of 72 bands were selected; the correlation coefficient of prediction was 0.9995, and the root mean square error of prediction was 0.0283. The experimental results show that higher accuracy can be obtained by establishing the PLS model using the characteristic bands screened by the linear relationship between spectral data and milk fat content. The SVR model was established based on the nonlinear relationship between spectral data and milk fat content. The accuracy of the SVR model was slightly lower than that of the PLS model. The selection of characteristic bands can improve the model's prediction accuracy, and the use of hyperspectral technology can realize the accurate detection of milk fat content.
Similar content being viewed by others
References
S. L. Li, K. Yao, Z. J. Cao, C. Q. Liu Y. C. Wang, Z. H. Wang, J. X. Li, J. Q. Wang, H. P. Zhang, and F. L. Kong, Chin. J. Animal Sci., 58, No. 3, 239–244 (2022).
D. M. Whitt, J. Pranata, B. G. Carter, D. M. Barbano, and M. A. Drake, J. Dairy Sci., 105, No. 7, 5700–5713 (2022).
L. Di Marzo, J. Pranata, and D. M. Barbano, J. Dairy Sci., 104, No. 7, 7448–7456 (2021).
Y. Y. Shao, Y. K. Shi, Y. D. Qin, G. T. Xuan, J. Li, Q. K. Li, F. J. Yang, and Z. C. Hu, Food Chem., 386, Article ID 132864 (2022).
Y. Q. Ren and D. W. Sun, Food Chem., 382, Article ID 132346 (2022).
A. Panda, R. B. Pachori, N. Kakkar, M. Joseph John, and N. D. Sinnappah-Kang, Comp. Methods Programs Biomed., 220, Article ID 106836 (2022).
S. Karim, A. Qadir, U. Farooq, M. Shakir, and A. A. Laghari, Curr. Med. Imaging, 19, No. 5, 417–427 (2022), https://doi.org/10.2174/157340561866622051914435.
A. Kayad, F. A. Rodrigues, S. Naranjo, M. Sozzi, F. Pirotti, F. Marinello, U. Schulthess, P. Defourny, B. Gerard, and M. Weiss, Field Crops Res., 282, Article ID 108449 (2022).
L. Zheng, Q. Bao, S. Z. Weng, J. P. Tao, D. Y. Zhang, L. S. Huang, and J. L. Zhao, Spectrochim. Acta A: Mol. Biomol. Spectrosc., 270, Article ID 12084 (2022).
X. Q. Pan, J. B. Jiang, and Y. M. Xiao, Ecolog. Inform., 68 (2022).
H. D. Zhang, J. Luo, S. M. Hou, Z. P. Xu, J. L. Evans, and S. L. He, Appl. Opt., 61, No. 12, 3400–3408 (2022).
J. Lim, G. Kim, C. Mo, M. S. Kim, K. Chao, J. Qin, X. Fu, I. Baek, and B.-K. Cho, Talanta, 151, 183–191 (2016).
Z. J. Zhao, Y. Wei, N. Q. Zhang, R. K. Chang, H. Y. Wu, H. Liu, H. Y. Shan, R. J. Yang, and X. Y. Guo, China Dairy Industry, 46, No. 2, 45–48 (2018).
Y. Wang, Y. Huang, W. Shen, F. Kong, M. Gao, and H. Sun, Spectrosc.. Lett., 54, No. 4, 316–325 (2021).
M. C. Liu, H. R. Xue, J. P. Liu, R. R. Dai, P. W. Hu, Q, Huang, and X. H. Jiang, Spectrosc. Spectr. Anal., 42, No. 5, 1601–1606 (2022).
M. Zhang, S. Zhang, and J. Iqbal, Chemometr. Intell. Lab. Systems, 128, 17–24 (2013).
P. Mishra and E. J. Woltering, Talanta, 224, Article ID 121908 (2021).
R. M. Balabin and S. V. Smirnov, Analyt. Chim. Acta, 692, Nos. 1–2, 63–72 (2011).
M. Radman, M. Moradi, A. Chaibakhsh, M. Kordestani, and M. Saif, IEEE Sensors J., 21, No. 3, 3533–3543 (2021).
V. Centner, D. L. Massart, O. E. de Noord, S. de Jong, B. M. Vandeginste, and C. Sterna, Anal. Chem., 68, No. 21, 3851–3858 (1996).
R. K. H. Galvao, M. C. Ugulino Araujo, W. D. Fragoso, E. C. Silva, G. E. Jose, S. F. Carreiro Soares, and H. M. Paiva, Chemometr. Intell. Lab. Systems, 92, No. 1, 83–91 (2008).
H.-Y. Zhen, R.-J. Ma, Y. Chen, X.-P. Sun, and C.-L. Ma, Spectrosc. Spectr. Anal., 40, No. 5, 1601–1606 (2020).
H. Li, Y. Liang, Q. Xu, and D. Cao, Anal. Chim. Acta, 648, No. 1, 77–84 (2009).
F. Allegrini and A. C. Olivieri, Anal. Chim. Acta, 699, No. 1, 18–25 (2011).
S. Cateni, V. Colla, and M. Vannucci, IEEE, 9th Int. Conf. Intell. Systems Design Appl., 1278–1283 (2009).
R. G. Zhu, H. W. Duan, X. D. Yao, Y. Y. Qiu, B. X. Ma, and C. J. Xu, Spectrosc. Spectr. Anal., 36, No. 9, 2925–2929 (2016).
Z. H. Tu, B. P. Ji, C. Y. Meng, D. Z. Zhu, B. L. Shi, and Z. S. Qing, Spectrosc. Spectr. Anal., 29, No. 10, 2760–2764 (2009).
M. Shamsipur, V. Zare-Shahabadi, B. Hemmateenejad, and M. Akhond, J. Chemometr., 20, No. 3–4, 146–157 (2006).
T. Liu, T. Xu, F. Yu, Q. Yuan, Z. Guo, and B. Xu, Comp. Electron. Agric., 186 (2021).
F. Marini and B. Walczak, Chemometr. Intell. Lab. Systems, 149, 153–165 (2015).
Y. Z. Hou, X. Gao, S. N. Li, X. Cai, P. Li, W. L. Li, and Z. Li, J. Pharmac. Innovation, 17, No. 4 (2022), https://doi.org/10.1007/s12247-022-09620-6.
H. Xu, S. Yu, J. Chen, and X. Zuo, Wireless Personal Commun., 102, No. 4, 2823–2834 (2018).
L. Xu, Y. Mo, Y. Lu, and J. Li, Processes, 9, No. 6, Article ID 1037 (2021).
P.-Y. Diwu, X.-H. Bian, Z.-F. Wang, and W. Liu, Spectrosc. Spectr. Anal., 39, No. 9, 2800–2806 (2019).
M. Li, Y. Feng, Y. Yu, T. Zhang, C. Yan, H. Tang, Q. Sheng, and H. Li, Spectrochim. Acta A: Mol. Biomol. Spectrosc., 257 (2021).
M. M. Galera, D. P. Zamora, J. L. M. Vidal, and A. G. Frenich, Analyt. Lett., 35, No. 5, 921–941 (2002).
Author information
Authors and Affiliations
Corresponding author
Additional information
Abstract of article is published in Zhurnal Prikladnoi Spektroskopii, Vol. 90, No. 4, p. 663, July–August, 2023.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, Q., Xu, Z.P., Jiang, X.H. et al. Nondestructive Detection of Milk Fat Content Based on Hyperspectral Technology. J Appl Spectrosc 90, 947–954 (2023). https://doi.org/10.1007/s10812-023-01617-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10812-023-01617-4