Abstract
The present study shows the existence of PII-acetyl-CoA carboxylase interaction in the cyanobacterium Synechococcus sp. PCC 7942 and the possible adverse impact of nitrogen starvation on this interaction. The in silico and in vitro analysis of PII-acetyl-CoA carboxylase interaction revealed that the biotin carboxyl carrier protein subunit of acetyl-CoA carboxylase enzyme actually interacts with the T-loop of PII protein. However, exposure of the cyanobacterium to nitrogen-starved condition showed a higher expression and activity of acetyl-CoA carboxylase at the intracellular level which denoted the impairment of PII-acetyl-CoA carboxylase interaction. A similar stimulatory effect of nitrogen starvation has also been noticed in the PII mutant of Synechococcus PCC 7942. Further, the physiological study reflected that nitrogen starvation–caused reactive oxygen species generation in the wild-type and PII mutant strains and lipid was increased in both strains of Synechococcus sp. PCC 7942. Proteomic analysis showed the upregulation of glycogen synthase, biotin carboxylase, and antioxidative enzymes and the deregulation of proteins involved in photosynthesis, energy metabolism, and protein synthesis. Interestingly, enhanced accumulation of transcripts of few tricarboxylic acid cycle genes was also noticed in the wild type. Although oxidative stress and lipid production were enhanced in both the test strains under nitrogen starvation, the impacts were more prominent in the mutant strain. Our results suggest that the nitrogen starvation–induced oxidative stress possibly relieved the PII-mediated inhibition of acetyl-CoA carboxylase and led to increased lipid synthesis in Synechococcus PCC 7942.










Similar content being viewed by others
References
Allen NM (1984) Cyanobacterial cell inclusions. Annu Rev Microbiol 38:1–25
Berman-Frank I, Bidle KD, Haramaty L, Falkowsky PG (2004) The demise of the marine cyanobacterium, Trichodesmium spp., via an autocatalyzed cell death pathway. Limnol Oceanogr 49:997–1005
Carrieri D, Lombardi T, Paddock T, Cano M, Goodney GA, Nag A, Old W, Maness PC, Seibert M, Ghirardi M, Yua J (2017) Transcriptome and proteome analysis of nitrogen starvation responses in Synechocystis 6803 ΔglgC, a mutant incapable of glycogen storage. Algal Res 21:64–75
Chang IF, Curran A, Woolsey R, Quilici D, Cushman JC, Mittler R, Harmon A, Harper JF (2009) Proteomic profiling of tandem affinity purified 14-3-3 protein complexes in Arabidopsis thaliana. Proteomics 9:2967–2985
Cronan JE Jr (2002) Interchangeable enzyme modules. J Biol Chem 277:22520–22527
Curci PL, Cigliano RA, Zuluaga DL, Janni M, Sanseverino W, Sonnante G (2017) Transcriptomic response of durum wheat to nitrogen starvation. Sci Rep 7:1176
Davis MS, Jr Cronan JE (2001) Inhibition of Escherichia coli acetyl coenzyme a carboxylase byacyl-acyl carrier protein. J Bacteriol 183:1499–1503
Deschoenmaeker F, Facchini R, Leroy B, Badri H, Zhang CC, Wattiez R (2014) Proteomic and cellular views of Arthrospira sp. PCC 8005 adaptation to nitrogen depletion. Microbiol 160:1224–1236
Ding Y, Gan N, Li J, Sedmak B, Song L (2012) Hydrogen peroxide induces apoptotic-like cell death in Microcystis aeruginosa (Chroococcales, Cyanobacteria) in a dose-dependent manner. Phycologia 51:567–575
Dong W, Wang M, Xu F, Quan T, Peng K, Xiao L, Xia G (2013) Wheat oxophytodienoate reductase gene TaOPR1 confers salinity tolerance via enhancement of abscisic acid signaling and reactive oxygen species scavenging. Plant Physiol 161:1217–1228
Feria Bourrellier AB, Valot B, Guillot A, Ambard-Bretteville F (2010) Chloroplast acetyl-CoA carboxylase activity is 2-oxoglutarate regulated by interaction of PII with the biotin carboxyl carrier subunit. Proc Natl Acad Sci 107:502–507
Folch J, Lees M, Sloane-Stanley GH (1957) A simple method for the isolation and purification of total lipids from animal tissues. J Biol Chem 226:497–509
Fulda S, Mikkat S, Huang F, Huckauf J, Marin K, Norling B, Hagemann M (2006) Proteome analysis of salt stress response in the cyanobacterium Synechocystis sp. strain PCC 6803. Proteomics 6:2733–2745
García-Domínguez M, Florencio FJ (1997) Nitrogen availability and electron transport control the expression of glnB gene (encoding PII protein) in the cyanobacterium Synechocystis sp. PCC 6803. Plant Mol Biol 35:723–734
Gerhardt EC, Rodrigues TE, Müller-Santos M, Pedrosa FO (2015) The bacterial signal transduction protein GlnB regulates the committed step in fatty acid biosynthesis by acting as a dissociable regulatory subunit of acetyl-CoA carboxylase. Mol Microbiol 95:1025–1035
Gordillo FJL, Jiménez C, Figueroa FL, Niell FX (1999) Effects of increased atmospheric CO2 and N supply on photosynthesis, growth and cell composition of the cyanobacterium Spirulina platensis (Arthrospira). J Appl Phycol 10:461–469
Hasunuma T, Kikuyama F, Matsuda M, Aikawa S, Izumi Y, Kondo A (2013) Dynamic metabolic profiling of cyanobacterial glycogen biosynthesis under conditions of nitrate depletion. J Exp Bot 64:2943–2954
Hauf W, Schmid K, Gerhardt ECM, Huergo LF (2016) Interaction of the nitrogen regulatory protein GlnB(PII) with biotin carboxyl carrier protein (BCCP) controls acetyl-CoA levels in the cyanobacterium Synechocystis sp. PCC 6803. Front Microbiol 7:1700
He YY, Hader DP (2002) UV-B-induced formation of reactive oxygen species and oxidative damage of the cyanobacterium Anabaena sp.: protective effects of ascorbic acid and N acetyl cysteine. J Photochem Photobiol 66:115–124
Hickman JW, Kotovic KM, Miller C, Warrener P, Kaiser B, Jurista T, Budde M, Cross F, Roberts JM, Carleton M (2013) Glycogen synthesis is a required component of the nitrogen stress response in Synechococcus elongatus PCC 7942. Algal Res 2:98–106
Huergo LF, Chandra G, Merrick M (2013) PII signal transduction proteins: nitrogen regulation and beyond. FEMS Microbiol Rev 37:251–283
Kaczmarzyk D, Fulda M (2010) Fatty acid activation in cyanobacteria mediated by acyl-acyl carrier protein synthetase enables fatty acid recycling. Plant Physiol 52:1598–1610
Kumar R, Biswas K, Singh PK, Singh PK, Elumalai S, Shukla P, Pabbi S (2017) Lipid production and molecular dynamics simulation for regulation of accD gene in cyanobacteria under different N and P regimes. Biotech Biofuels 10:94
Li SJ, Cronan JE Jr (1992) The gene encoding the biotin carboxylase subunit of Escherichia coli acetyl-CoA carboxylase. J Biol Chem 267:855–863
Merrick M (2014) Post-translational modification of PII signals transduction proteins. Front Microbiol 5:763
Nayeem A, Sitkoff D, Krystek S Jr (2006) A comparative study of available software for high accuracy homology modeling: from sequence alignments to structural models. Protein Sci 15:808–824
Pardo MA, Llama MJ, Serra JL (1999) Purification, properties and enhanced expression under nitrogen starvation of the NADP-isocitrate dehydrogenase from the cyanobacterium Phormidium laminosum. Biochim Biophys Acta 1431:87–96
Płociński P, Laubitz D, Cysewski D, Stoduś K, Kowalska K, Dziembowski A (2014) Identification of protein partners in mycobacteria using a single-step affinity purification method. PLoS One 9:e91380
Podkowiński J, Tworak A (2011) Acetyl-coenzyme a carboxylase – an attractive enzyme for biotechnology. BioTechnologia 92:321–335
Rhee EC, Cronan JE (2003) The biotin carboxylase-biotin carboxyl carrier protein complex of Escherichia coli acetyl-CoA carboxylase. J Biol Chem 278:30806–30812
Rippka R, Deruelles J, Waterbury JB, Herdman MR, Stanier RY (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol 111:1–61
Rodrigues TE, Gerhardt EC, Oliveira MA, Chubatsu LS (2014) Search for novel targets of the PII signal transduction protein in bacteria identifies the BCCP component of acetyl-CoA carboxylase as a PII binding partner. Mol Microbiol 91:751–761
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor, NY
Sumper M, Träuble H (1973) Membranes as acceptors for palmitoyl CoA in fatty acid biosynthesis. FEBS Lett 30:29–34
Thelen JJ, Mekhedov S, Ohlrogge JB (2001) Brassicaceae express multiple isoforms of biotin carboxyl carrier protein in a tissue-specific manner. Plant Physiol 125:2016–2028
Tichy M, Vermaas W (1999) In vivo role of catalase-peroxidase in Synechocystis sp. strain PCC6803. J Bacteriol 181:1875–1882
Verma E, Chakraborty S, Tiwari B, Mishra AK (2018a) Transcriptional regulation of acetyl CoA and lipid synthesis by PII protein in Synechococcus PCC 7942. J Basic Microbiol 58:187–197
Verma E, Chakraborty S, Tiwari B, Singh S, Mishra AK (2018b) Alleviation of NaCl toxicity in the cyanobacterium Synechococcus sp. PCC 7942 by exogenous calcium supplementation. J Appl Phycol 30:1465
Wagner MA, Eschenbrenner M, Horn TA, Kraycer JA, Mujer CV, Hagius S, Elzer P, DelVecchio VG (2002) Global analysis of the Brucella melitensis proteome: identification of proteins expressed in laboratory- grown culture. Proteomics 2:1047–1060
Watanabe M, Ikeuchi M (2013) Phycobilisome: architecture of a light-harvesting supercomplex. Photosynth Res 116:265–276
Willis LB, Omar WSW, Sambanthamurthi R, Sinskey AJ (2008) Non radioactive assay for acetyl-a carboxylase activity. J Oil Palm Res 2:30–36
Yilancioglu K, Cokol M, Pastirmaci I, Erman B, Cetiner S (2014) Oxidative stress is a mediator for increased lipid accumulation in a newly isolated Dunaliella salina strain. PLoS One 9:e91957
Acknowledgments
We are thankful to the Head, Department of Botany, Banaras Hindu University, Varanasi, India, for providing laboratory facilities. We thank Prof. Karl Forchhammer, Department of Organismic Interactions (Microbiology), Interfaculty Institute of Microbiology and Infection, Auf der Morgenstelle, 2872076, University of Tübingen, Germany, for providing Synechococcus sp. PCC 7942 wild and mutant strains. Ekta Verma is thankful to the UGC, New Delhi, for financial support in the form of SRF.
Author information
Authors and Affiliations
Contributions
EV and AKM designed the experiments. EV, SC, and SK performed the experiments. EV, SC, BT, and SSS were involved in analyzing the data. EV, SC, SSS, and AKM wrote the manuscript. AKM critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare there are no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Fig. S1
Three dimensional structures of (A) PII protein and (B) Biotin carboxyl carrier protein of Synechococcus sp. PCC 7942. (PNG 299 kb)
Supplementary Fig. S2
Ramachandran plot of biotin carboxyl carrier protein showing the residues in favoured, allowed and outlier region. (PNG 326 kb)
Supplementary Fig. S3
Three dimensional docked structure of PII and biotin carboxyl carrier protein using the Hex programme (S3A). Blue colour structure represents the PII protein whereas brown color represents the BCCP protein. Interacting residues of both proteins are presented in Fig. S3B. Fig. S3C represents the distance between interacting residues and Fig. S3D represents the polar interactions. (PNG 1340 kb)
Supplementary Fig. S4
Residues of PII protein and biotin carboxyl carrier protein involve in protein-protein interaction. Red, yellow, green and blue colour represents the sequences of PII protein, sequences of biotin carboxyl carrier protein, residues of PII protein interacted with biotin carboxyl carrier protein and residues of biotin carboxyl carrier protein interacted with PII protein respectively. (PNG 271 kb)
Supplementary Fig. S5
SDS-PAGE images of (A) expression of cloned PII protein and (B) purification of PII protein using Ni-NTA column. (PNG 1003 kb)
Supplementary Fig. S6
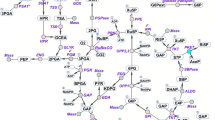
Transcript analysis in terms of reverse transcriptase PCR of Synpcc7942_1379 (Biotin carboxylase), Synpcc7942_2564 (Biotin carboxyl carrier protein), Synpcc7942_0918 (Acyl-ACP synthetase), Synpcc7942_0801 (SOD), Synpcc7942_1656 (CAT), Synpcc7942_1426 (Rubisco) and Synpcc7942_2518 (Glycogen synthase) (A). Transcript abundance of some genes of TCA cycle was also analysed in terms of reverse transcriptase PCR of Synpcc7942_0143 (Pyruvate dehydrogenase), Synpcc7942_1719 (Isocitrate dehydrogenase), Synpcc7942_0098 (Pyruvate kinase), Synpcc7942_1435 (2-oxoglutarate decarboxylase), Synpcc7942_2160 (Alanine aminotransferase) and Synpcc7942_2310 (Glutamate decarboxylase) (B). Apart from these transcripts abundance of glnB gene encoding PII protein was also estimated (C). Wt (wild); Wt-N (wild-N); MP2 (mutant); MP2-N (mutant-N). (PNG 305 kb)
Rights and permissions
About this article
Cite this article
Verma, E., Chakraborty, S., Kharwar, S. et al. Nitrogen starvation–induced oxidative stress relieves PII-mediated inhibition of acetyl-CoA carboxylase (ACCase) activity and signals enhanced lipid synthesis in Synechococcus PCC 7942. J Appl Phycol 33, 313–329 (2021). https://doi.org/10.1007/s10811-020-02316-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-020-02316-9