Abstract
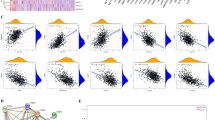
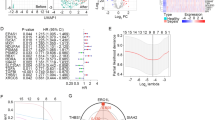
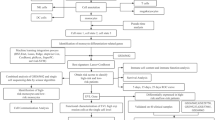
Sepsis is a disease with a very high mortality rate, mainly involving an immune-dysregulated response due to bacterial infection. Most studies are currently limited to the whole blood transcriptome level; however, at the single cell level, there is still a great deal unknown about specific cell subsets and disease markers. We obtained 29 peripheral blood single-cell sequencing data, including 66,283 cells from 10 confirmed samples of sepsis infection and 19 healthy samples. Cells related to the sepsis phenotype were identified and characterized by the “scissor” method. The regulatory relationships of sepsis-related phenotype cells in the cellular communication network were clarified using the “cell chat” method. The least absolute shrinkage and selection operator (LASSO), support vector machine (SVM), and random forest (RF) were used to identify sepsis signature genes of diagnostic value. External validation was performed using multiple datasets from the GEO database (GSE28750, GSE185263, GSE57065) and 40 clinical samples. Bayesian algorithm was used to calculate the regulatory network of LILRA5 co-expressed genes. The stability of atenolol-targeting LILRA5 was determined by molecular docking techniques. Ultimately, action trajectory and survival analyses demonstrate the effectiveness of atenolol-targeted LILRA5 in treating patients with sepsis. We successfully identified 1215 healthy phenotypic cells and 462 sepsis phenotypic cells. We focused on 447 monocytes of the sepsis phenotype. Among the cellular communications, there were a large number of differences between these cells and other immune cells showing a significant inflammatory phenotype compared to the healthy phenotypic cells. Together, the three machine learning algorithms identified the LILRA5 marker gene in sepsis patients, and validation results from multiple external datasets as well as real-world clinical samples demonstrated the robust diagnostic performance of LILRA5. The AUC values of LILRA5 in the external datasets GSE28750, GSE185263, and GSE57065 could reach 0.875, 0.940, and 0.980, in that order. Bayesian networks identified a large number of unknown regulatory relationships for LILRA5 co-expression. Molecular docking results demonstrated the possibility of atenolol targeting LILRA5 for the treatment of sepsis. Behavioral trajectory analysis and survival analysis demonstrate that atenolol has a desirable therapeutic effect. LILRA5 is a marker gene in sepsis patients, and atenolol can stably target LILRA5.







Similar content being viewed by others
AVAILABILITY OF DATA AND MATERIALS
The datasets supporting the conclusions of this article are available in the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP548/an-immune-cell-signature-of-bacterial-sepsis-patient-pbmcs#study-summary) and the GEO database (The dataset(s) supporting the conclusions of this article is(are) available). The GEO registration numbers are GSE69063, GSE28750, GSE185263, GSE57065).
Abbreviations
- PCA:
-
Principal component analysis
- PCs:
-
Principal components
- GEO:
-
Gene expression omnibus
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the curve
- LASSO:
-
Least absolute shrinkage and selection operator
- SVM:
-
Support vector machine
- RF:
-
Random forest
References
Arina, Pietro, and Mervyn Singer. 2021. Pathophysiology of sepsis. Current Opinion in Anaesthesiology 34: 77–84. https://doi.org/10.1097/ACO.0000000000000963.
Downes, Kevin J., Julie C. Fitzgerald, and Scott L. Weiss. 2020. Utility of procalcitonin as a biomarker for sepsis in children. Edited by Colleen Suzanne Kraft. Journal of Clinical Microbiology 58: e01851-e1919. https://doi.org/10.1128/JCM.01851-19.
Stocker, Martin, Wendy van Herk, Salhab el Helou, Sourabh Dutta, Frank A B A. Schuerman, Rita K. van den Tooren-de, Jantien W. Groot, Wieringa, et al. 2021. C-Reactive protein, procalcitonin, and white blood count to rule out neonatal early-onset sepsis within 36 hours: A secondary analysis of the neonatal procalcitonin intervention study. Clinical Infectious Diseases 73: e383–e390. https://doi.org/10.1093/cid/ciaa876.
Chakraborty, Rebanta K., and Bracken Burns. 2022. Systemic inflammatory response syndrome. In StatPearls. Treasure Island (FL): StatPearls Publishing.
Yende, Sachin, John A. Kellum, Victor B. Talisa, Octavia M. Peck Palmer, Chung-Chou H. Chang, Michael R. Filbin, Nathan I. Shapiro, et al. 2019. Long-term host immune response trajectories among hospitalized patients with sepsis. JAMA network open 2: e198686. https://doi.org/10.1001/jamanetworkopen.2019.8686.
Barichello, Tatiana, Jaqueline S. Generoso, Diogo Dominguini, Emily Córneo, Vijayasree V. Giridharan, Taha A. Sahrapour, Lutiana R. Simões, et al. 2021. Postmortem evidence of brain inflammatory markers and injury in septic patients: A systematic review. Critical Care Medicine Publish Ahead of Print. https://doi.org/10.1097/CCM.0000000000005307.
Kumar, Sushil, Harshad Ingle, Durbaka Vijaya Raghava. Prasad, and Himanshu Kumar. 2013. Recognition of bacterial infection by innate immune sensors. Critical Reviews in Microbiology 39: 229–246. https://doi.org/10.3109/1040841X.2012.706249.
Heckenberg, Sebastiaan G. B., Matthijs C. Brouwer, and Diederik van de Beek. 2014. Bacterial meningitis. Handbook of Clinical Neurology 121: 1361–1375. https://doi.org/10.1016/B978-0-7020-4088-7.00093-6.
Mook-Kanamori, Barry B., Madelijn Geldhoff, Tom van der Poll, and Diederik van de Beek. 2011. Pathogenesis and pathophysiology of pneumococcal meningitis. Clinical Microbiology Reviews 24: 557–591. https://doi.org/10.1128/CMR.00008-11.
Sellner, Johann, Martin G. Täuber, and Stephen L. Leib. 2010. Pathogenesis and pathophysiology of bacterial CNS infections. Handbook of Clinical Neurology 96: 1–16. https://doi.org/10.1016/S0072-9752(09)96001-8.
Iwasaki, Akiko, and Ruslan Medzhitov. 2010. Regulation of adaptive immunity by the innate immune system. Science (New York, N.Y.) 327: 291–295. https://doi.org/10.1126/science.1183021.
Kataoka, Hiroshi, Hajime Kono, Zubin Patel, Yoshitaka Kimura, and Kenneth L. Rock. 2014. Evaluation of the contribution of multiple DAMPs and DAMP receptors in cell death-induced sterile inflammatory responses. PloS One 9: e104741. https://doi.org/10.1371/journal.pone.0104741.
Kigerl, Kristina A., Juan Pablo, W. de Rivero Vaccari, Dalton Dietrich, Phillip G. Popovich, and Robert W. Keane. 2014. Pattern recognition receptors and central nervous system repair. Experimental Neurology 258: 5–16. https://doi.org/10.1016/j.expneurol.2014.01.001.
Wiersinga, Willem Joost, Stije J. Leopold, Duncan R. Cranendonk, and Tom van der Poll. 2014. Host innate immune responses to sepsis. Virulence 5: 36–44. https://doi.org/10.4161/viru.25436.
Generoso, Jaqueline S., Vijayasree V. Giridharan, Juneyoung Lee, Danielle Macedo, and Tatiana Barichello. 2021. The role of the microbiota-gut-brain axis in neuropsychiatric disorders. Revista Brasileira De Psiquiatria (Sao Paulo, Brazil: 1999) 43: 293–305. https://doi.org/10.1590/1516-4446-2020-0987.
Barichello, Tatiana, Pavani Sayana, Vijayasree V. Giridharan, Anithachristy S. Arumanayagam, Boomadevi Narendran, Amanda Della Giustina, Fabricia Petronilho, João. Quevedo, and Felipe Dal-Pizzol. 2019. Long-term cognitive outcomes after sepsis: A translational systematic review. Molecular Neurobiology 56: 186–251. https://doi.org/10.1007/s12035-018-1048-2.
Korsunsky, Ilya, Nghia Millard, Jean Fan, Kamil Slowikowski, Fan Zhang, Kevin Wei, Yuriy Baglaenko, Michael Brenner, Po-ru Loh, and Soumya Raychaudhuri. 2019. Fast, sensitive and accurate integration of single-cell data with Harmony. Nature Methods 16: 1289–1296. https://doi.org/10.1038/s41592-019-0619-0.
Sun, Duanchen, Xiangnan Guan, Amy E. Moran, Wu. Ling-Yun, David Z. Qian, Pepper Schedin, Mu-Shui. Dai, et al. 2022. Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data. Nature Biotechnology 40: 527–538. https://doi.org/10.1038/s41587-021-01091-3.
Jin, Suoqin, Christian F. Guerrero-Juarez, Lihua Zhang, Ivan Chang, Raul Ramos, Chen-Hsiang. Kuan, Peggy Myung, Maksim V. Plikus, and Qing Nie. 2021. Inference and analysis of cell-cell communication using Cell Chat. Nature Communications 12: 1088. https://doi.org/10.1038/s41467-021-21246-9.
Wu, Yingcheng, Shuaixi Yang, Jiaqiang Ma, Zechuan Chen, Guohe Song, Dongning Rao, Yifei Cheng, et al. 2022. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discovery 12: 134–153. https://doi.org/10.1158/2159-8290.CD-21-0316.
Friedman, Jerome, Trevor Hastie, and Robert Tibshirani. 2010. Regularization paths for generalized linear models via coordinate descent. Journal of Statistical Software 33. https://doi.org/10.18637/jss.v033.i01.
Robin, Xavier, Natacha Turck, Alexandre Hainard, Natalia Tiberti, Frédérique. Lisacek, Jean-Charles. Sanchez, and Markus Müller. 2011. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12: 77. https://doi.org/10.1186/1471-2105-12-77.
Sato, Noriaki, Yoshinori Tamada, Yu. Guangchuang, and Yasushi Okuno. 2022. CBNplot : Bayesian network plots for enrichment analysis. Edited by Zhiyong Lu. Bioinformatics 38: 2959–2960. https://doi.org/10.1093/bioinformatics/btac175.
Singer, Mervyn, Clifford S. Deutschman, Christopher Warren Seymour, Manu Shankar-Hari, Djillali Annane, Michael Bauer, Rinaldo Bellomo, et al. 2016. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA 315: 801. https://doi.org/10.1001/jama.2016.0287.
Reyes, Miguel, Michael R. Filbin, Roby P. Bhattacharyya, Kianna Billman, Thomas Eisenhaure, Deborah T. Hung, Bruce D. Levy, et al. 2020. An immune-cell signature of bacterial sepsis. Nature Medicine 26: 333–340. https://doi.org/10.1038/s41591-020-0752-4.
Young, Matthew D., Thomas J. Mitchell, Felipe A. Vieira, Maxine G. B. Braga, Benjamin J. Tran, John R. Stewart, Grace Collord Ferdinand, et al. 2018. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361: 594–599. https://doi.org/10.1126/science.aat1699.
Wang, Bo., Junjie Zhu, Emma Pierson, Daniele Ramazzotti, and Serafim Batzoglou. 2017. Visualization and analysis of single-cell RNA-seq data by kernel-based similarity learning. Nature Methods 14: 414–416. https://doi.org/10.1038/nmeth.4207.
Sinha, Debajyoti, Akhilesh Kumar, Himanshu Kumar, Sanghamitra Bandyopadhyay, and Debarka Sengupta. 2018. dropClust: Efficient clustering of ultra-large scRNA-seq data. Nucleic Acids Research 46: e36–e36. https://doi.org/10.1093/nar/gky007.
Villani, Alexandra-Chloé, Rahul Satija, Gary Reynolds, Siranush Sarkizova, Karthik Shekhar, James Fletcher, Morgane Griesbeck, et al. 2017. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356: eaah4573. https://doi.org/10.1126/science.aah4573.
Perretti, Mauro, and Fulvio D’Acquisto. 2009. Annexin A1 and glucocorticoids as effectors of the resolution of inflammation. Nature Reviews Immunology 9: 62–70. https://doi.org/10.1038/nri2470.
Ness, Traci L., Kristin J. Carpenter, Jillian L. Ewing, Craig J. Gerard, Cory M. Hogaboam, and Steven L. Kunkel. 2004. CCR1 and CC chemokine ligand 5 interactions exacerbate innate immune responses during sepsis. The Journal of Immunology 173: 6938–6948. https://doi.org/10.4049/jimmunol.173.11.6938.
He, Min, Richard Horuk, Shabbir M. Moochhala, and Madhav Bhatia. 2007. Treatment with BX471, a CC chemokine receptor 1 antagonist, attenuates systemic inflammatory response during sepsis. American Journal of Physiology-Gastrointestinal and Liver Physiology 292: G1173–G1180. https://doi.org/10.1152/ajpgi.00420.2006.
Parker, Margaret M., James H. Shelhamer, Charles Natanson, David W. Alling, and Joseph E. Parrillo. 1987. Serial cardiovascular variables in survivors and nonsurvivors of human septic shock: Heart rate as an early predictor of prognosis. Critical Care Medicine 15: 923–929. https://doi.org/10.1097/00003246-198710000-00006.
Kuo, Ming-Jen, Ruey-Hsing Chou, Ya-Wen Lu, Jiun-Yu Guo, Yi-Lin Tsai, Cheng-Hsueh Wu, Po-Hsun Huang, and Shing-Jong Lin. 2021. Premorbid β1-selective (but not non-selective) β-blocker exposure reduces intensive care unit mortality among septic patients. Journal of Intensive Care 9: 40. https://doi.org/10.1186/s40560-021-00553-9.
Barichello, Tatiana, Jaqueline S. Generoso, Mervyn Singer, and Felipe Dal-Pizzol. 2022. Biomarkers for sepsis: More than just fever and leukocytosis-a narrative review. Critical Care (London, England) 26: 14. https://doi.org/10.1186/s13054-021-03862-5.
Fu, Guang, Meihong Deng, Matthew D. Neal, Timothy R. Billiar, and Melanie J. Scott. 2021. Platelet–monocyte aggregates: Understanding mechanisms and functions in sepsis. Shock 55: 156–166. https://doi.org/10.1097/SHK.0000000000001619.
Vogel, Sebastian, Rebecca Bodenstein, Qiwei Chen, Susanne Feil, Robert Feil, Johannes Rheinlaender, Tilman E. Schäffer, et al. 2015. Platelet-derived HMGB1 is a critical mediator of thrombosis. The Journal of Clinical Investigation 125: 4638–4654. https://doi.org/10.1172/JCI81660.
Vogel, Sebastian, Dominik Rath, Oliver Borst, Andreas Mack, Patricia Loughran, Michael T. Lotze, Matthew D. Neal, Timothy R. Billiar, and Meinrad Gawaz. 2016. Platelet-derived high-mobility group box 1 promotes recruitment and suppresses apoptosis of monocytes. Biochemical and Biophysical Research Communications 478: 143–148. https://doi.org/10.1016/j.bbrc.2016.07.078.
Zhou, Hui, Meihong Deng, Yingjie Liu, Chenxuan Yang, Rosemary Hoffman, Jingjiao Zhou, Patricia A. Loughran, Melanie J. Scott, Matthew D. Neal, and Timothy R. Billiar. 2018. Platelet HMGB1 is required for efficient bacterial clearance in intra-abdominal bacterial sepsis in mice. Blood Advances 2: 638–648. https://doi.org/10.1182/bloodadvances.2017011817.
Morrell, Craig N., Angela A. Aggrey, Lesley M. Chapman, and Kristina L. Modjeski. 2014. Emerging roles for platelets as immune and inflammatory cells. Blood 123: 2759–2767. https://doi.org/10.1182/blood-2013-11-462432.
Kral, Julia Barbara, Waltraud Cornelia Schrottmaier, Manuel Salzmann, and Alice Assinger. 2016. Platelet Interaction with innate immune cells. Transfusion Medicine and Hemotherapy: Offizielles Organ Der Deutschen Gesellschaft Fur Transfusionsmedizin Und Immunhamatologie 43: 78–88. https://doi.org/10.1159/000444807.
Semple, John W., Joseph E. Italiano, and John Freedman. 2011. Platelets and the immune continuum. Nature Reviews. Immunology 11: 264–274. https://doi.org/10.1038/nri2956.
Weyrich, Andrew S., and Guy A. Zimmerman. 2004. Platelets: Signaling cells in the immune continuum. Trends in Immunology 25: 489–495. https://doi.org/10.1016/j.it.2004.07.003.
Goasguen, Jean E., John M. Bennett, Barbara J. Bain, Teresa Vallespi, Richard Brunning, Ghulam J. Mufti, and International Working Group on Morphology of Myelodysplastic Syndrome. 2009. Morphological evaluation of monocytes and their precursors. Haematologica 94: 994–997. https://doi.org/10.3324/haematol.2008.005421.
Bomans, Katharina, Judith Schenz, Isabella Sztwiertnia, Dominik Schaack, Markus Alexander Weigand, and Florian Uhle. 2018. Sepsis induces a long-lasting state of trained immunity in bone marrow monocytes. Frontiers in Immunology 9: 2685. https://doi.org/10.3389/fimmu.2018.02685.
Shen, Tian, Xingjian Cao, Junwei Shi, Yu Yu, Yihua Zhu, and Delin Gu. 2018. The morphological changes of monocytes in peripheral blood as a potential indicator for predicting active pulmonary tuberculosis. Clinica Chimica Acta; International Journal of Clinical Chemistry 481: 189–192. https://doi.org/10.1016/j.cca.2018.03.015.
Shi, Chao, and Eric G. Pamer. 2011. Monocyte recruitment during infection and inflammation. Nature Reviews. Immunology 11: 762–774. https://doi.org/10.1038/nri3070.
Fingerle, G., A. Pforte, B. Passlick, M. Blumenstein, M. Ströbel, and H.W. Ziegler-Heitbrock. 1993. The novel subset of CD14+/CD16+ blood monocytes is expanded in sepsis patients. Blood 82: 3170–3176.
Wong, Kok Loon, Wei Hseun Yeap, June Jing Yi. Tai, Siew Min Ong, Truong Minh Dang, and Siew Cheng Wong. 2012. The three human monocyte subsets: Implications for health and disease. Immunologic Research 53: 41–57. https://doi.org/10.1007/s12026-012-8297-3.
Margaria, Jean Piero, Lucia Moretta, Jose Carlos Alves-Filho, and Emilio Hirsch. 2022. PI3K Signaling in mechanisms and treatments of pulmonary fibrosis following sepsis and acute lung injury. Biomedicines 10: 756. https://doi.org/10.3390/biomedicines10040756.
Qiu, Peng, Yang Liu, and Jin Zhang. 2019. Review: The role and mechanisms of macrophage autophagy in sepsis. Inflammation 42: 6–19. https://doi.org/10.1007/s10753-018-0890-8.
Esmon, C.T. 2001. Role of coagulation inhibitors in inflammation. Thrombosis and Haemostasis 86: 51–56.
Aykac, Asli, Ahmet Özer Şehirli, and M. Zafer Gören. 2020. Evaluation of the effect of prazosin treatment on α-2c adrenoceptor and apoptosis protein levels in the predator scent-induced rat model of post-traumatic stress disorder. Journal of Molecular Neuroscience 70: 1120–1129. https://doi.org/10.1007/s12031-020-01518-7.
Ackland, Gareth L., Song T. Yao, Alain Rudiger, Alex Dyson, Ray Stidwill, Dmitry Poputnikov, Mervyn Singer, and Alexander V. Gourine. 2010. Cardioprotection, attenuated systemic inflammation, and survival benefit of beta1-adrenoceptor blockade in severe sepsis in rats. Critical Care Medicine 38: 388–394. https://doi.org/10.1097/CCM.0b013e3181c03dfa.
Calzavacca, Paolo, Yugeesh R. Lankadeva, Simon R. Bailey, Michael Bailey, Rinaldo Bellomo, and Clive N. May. 2014. Effects of selective β1-adrenoceptor blockade on cardiovascular and renal function and circulating cytokines in ovine hyperdynamic sepsis. Critical Care (London, England) 18: 610. https://doi.org/10.1186/s13054-014-0610-1.
Bedet, Alexandre, Guillaume Voiriot, Julien Ternacle, Elisabeth Marcos, Serge Adnot, Geneviève Derumeaux, and Armand Mekontso Dessap. 2020. Heart rate control during experimental sepsis in mice: Comparison of ivabradine and β-blockers. Anesthesiology 132: 321–329. https://doi.org/10.1097/ALN.0000000000003045.
Funding
This work was supported by grants from the National Natural Science Foundation of China (81971474), the Natural Science Foundation of Hebei Province (No. C2021206011), the Hebei Key R&D Program Project Special Project for the Construction of Beijing-Tianjin-Hebei Collaborative Innovation Community (No. 22347702D), the Hebei Provincial Department of Education Grants for Cultivating Innovation Ability of Graduate Students at the Provincial Level (CXZZBS2023107), and the Hebei Medical University Innovation Grant Program (XCXZZB202301).
Author information
Authors and Affiliations
Contributions
Jingyuan Ning, Xiaoqing Fan, Keran Sun, Xuan Wang, and Cuiqing Ma conceived and wrote the paper. Jingyuan Ning, Cuiqing Ma, Xiaoqing Fan, Keran Sun, Xuan Wang, Keqi Jia, and Hongru Li analyzed the materials and drafted the manuscript. Cuiqing Ma revised the whole paper. All authors have reviewed the final version of the manuscript and approved it to submit to your journal.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
All methods were carried out in accordance with relevant guidelines and regulations.
Consent for Publication
Not applicable.
Competing Interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ning, J., Fan, X., Sun, K. et al. Single-cell Sequence Analysis Combined with Multiple Machine Learning to Identify Markers in Sepsis Patients: LILRA5. Inflammation 46, 1236–1254 (2023). https://doi.org/10.1007/s10753-023-01803-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-023-01803-8