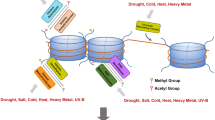
Abstract
Plants, unlike animals, cannot move from one place to another and have to face different climatic disturbances wherever they are growing. So, they have innumerable built-in mechanisms to adapt to various abiotic stressful conditions like drought, heat, cold, and salinity. The changing environmental conditions influence the expression patterns of genes. Epigenetics involves heritable changes in DNA bases or histone proteins, which ultimately create different conformational states of chromatin. The regulatory enzymes of epigenetic modifications are grouped as writers, readers and erasers, which add, recognize and remove the epigenetic marks, respectively. Here, we provide a comprehensive overview of the mechanism of DNA methylation by the RdDM pathway, its maintenance and removal, and different histone modification categories like acetylation, methylation, phosphorylation and ubiquitination. This review further discusses in detail the crucial role these modifications play in adapting to major abiotic stresses and how plants preserve these experiences as stress memory to respond to recurring stresses. It emphasizes the role of epigenetic modifications as a crucial mechanism for building plant’s tolerance and how it can be an important research priority to improve plant growth and development under abiotic stress conditions.



Similar content being viewed by others
Abbreviations
- CMT:
-
Chromomethylase
- AGO:
-
Argonoute
- DRM2:
-
Domains rearranged methyltransferase 2
- MET1:
-
Methyltransferase 1
- RdDM:
-
RNA-directed DNA methylation
- HAT:
-
Histone acetyltransferase
- HDAC:
-
Histone deactylase
References
Antunez-Sanchez J, Naish M, Ramirez-Prado JS, Ohno S, Huang Y, Dawson A, Opassathian K, Manza-Mianza D, Ariel F, Raynaud C, Wibowo A (2020) A new role for histone demethylases in the maintenance of plant genome integrity. Elife 9:e58533. https://doi.org/10.7554/eLife.58533
Aranda S, Mas G, Di Croce L (2015) Regulation of gene transcription by polycomb proteins. Sci Adv 1:e1500737. https://doi.org/10.1126/sciadv.1500737
Ashapkin VV, Kutueva LI, Aleksandrushkina NI, Vanyushin BF (2020) Epigenetic mechanisms of plant adaptation to biotic and abiotic stresses. Int J Mol Sci 21:7457. https://doi.org/10.3390/ijms21207457
Baek D, Jiang J, Chung J-S, Wang B, Chen J, Xin Z, Shi H (2011) Regulated AtHKT1 gene expression by a distal enhancer element and DNA methylation in the promoter plays an important role in salt tolerance. Plant Cell Physiol 52:149–161. https://doi.org/10.1093/pcp/pcq182
Bäurle I, Trindade I (2020) Chromatin regulation of somatic abiotic stress memory. J Exp Bot 71:5269–5279. https://doi.org/10.1093/jxb/eraa098
Bobde RC, Kumar A, Vasudevan D (2022) Plant-specific HDT family histone deacetylases are nucleoplasmins. Plant Cell 34:4760–4777. https://doi.org/10.1093/plcell/koac275
Buszewicz D, Archacki R, Palusiński A, Kotliński M, Fogtman A, Iwanicka-Nowicka R, Sosnowska K, Kuciński J, Pupel P, Olędzki J, Dadlez M (2016) HD2C histone deacetylase and a SWI/SNF chromatin remodelling complex interact and both are involved in mediating the heat stress response in Arabidopsis. Plant Cell Environ 39:2108–2122. https://doi.org/10.1111/pce.12756
Carter B, Bishop B, Ho KK, Huang R, Jia W, Zhang H, Pascuzzi PE, Deal RB, Ogas J (2018) The chromatin remodelers PKL and PIE1 act in an epigenetic pathway that determines H3K27me3 homeostasis in Arabidopsis. Plant Cell 30:1337–1352. https://doi.org/10.1105/tpc.17.00867
Chandrika NNP, Sundaravelpandian K, Yu S, Schmidt W (2013) ALFIN-LIKE 6 is involved in root hair elongation during phosphate deficiency in Arabidopsis. New Phytol 198:709–720. https://doi.org/10.1111/nph.12194
Chen X, Schönberger B, Menz J, Ludewig U (2018) Plasticity of DNA methylation and gene expression under zinc deficiency in Arabidopsis roots. Plant Cell Physiol 59:1790–1802. https://doi.org/10.1093/pcp/pcy100
Chinnusamy V, Gong Z, Zhu J (2008) Abscisic acid-mediated epigenetic processes in plant development and stress responses. J Integ Plant Biol 50:1187–1195. https://doi.org/10.1111/j.1744-7909.2008.00727.x
Cong W, Miao Y, Xu L, Zhang Y, Yuan C, Wang J, Zhuang T, Lin X, Jiang L, Wang N (2019) Transgenerational memory of gene expression changes induced by heavy metal stress in rice (Oryza sativa L.). BMC Plant Biol 19:1–14. https://doi.org/10.1186/s12870-019-1887-7
Cuerda-Gil D, Slotkin RK (2016) Non-canonical RNA-directed DNA methylation. Nat Plants 2:1–8
Ding Y, Fromm M, Avramova Z (2012) Multiple exposures to drought ‘train’ transcriptional responses in Arabidopsis. Nat Commun 3:740. https://doi.org/10.1038/ncomms1732
Ding Y, Wang W, Zhuang Q, Luo Y (2020) Adaptation of paddy rice in China to climate change: the effects of shifting sowing date on yield and irrigation water requirement. Agric Water Manag 228:105890. https://doi.org/10.1016/j.agwat.2019.105890
Du J, Zhong X, Bernatavichute YV, Stroud H, Feng S, Caro E, Vashisht AA, Terragni J, Chin HG, Tu A (2012) Dual binding of chromomethylase domains to H3K9me2-containing nucleosomes directs DNA methylation in plants. Cell 151:167–180
Duan CG, Zhu JK, Cao X (2018) Retrospective and perspective of plant epigenetics in China. J Genet Genom 45:621–638. https://doi.org/10.1016/j.jgg.2018.09.004
Earley KW, Shook MS, Brower-Toland B, Hicks L, Pikaard CS (2007) In vitro specificities of Arabidopsis co-activator histone acetyltransferases: implications for histone hyperacetylation in gene activation. Plant J 52:615–626. https://doi.org/10.1111/j.1365-313X.2007.03264.x
Fan H, Zhang Z, Wang N, Cui Y, Sun H, Liu Y, Wu H, Zheng S, Bao S, Ling H (2014) SKB1/PRMT 5-mediated histone H 4 R 3 dimethylation of I b subgroup bHLH genes negatively regulates iron homeostasis in Arabidopsis thaliana. Plant J 77:209–221. https://doi.org/10.1111/tpj.12380
Fina JP, Masotti F, Rius SP, Crevacuore F, Casati P (2017) HAC1 and HAF1 histone acetyltransferases have different roles in UV-B responses in Arabidopsis. Front Plant Sci 8:1179. https://doi.org/10.3389/fpls.2017.01179
Finnegan EJ, Peacock WJ, Dennis ES (1996) Reduced DNA methylation in Arabidopsis thaliana results in abnormal plant development. Proc Natl Acad Sci 93:8449–8454. https://doi.org/10.1073/pnas.93.16.8449
Friedrich T, Faivre L, Bäurle I, Schubert D (2019) Chromatin-based mechanisms of temperature memory in plants. Plant Cell Environ 42:762–770. https://doi.org/10.1111/pce.13373
Goldberg AD, Allis CD, Bernstein E (2007) Epigenetics: a landscape takes shape. Cell 128:635–638. https://doi.org/10.1016/j.cell.2007.02.006
Gu T, Han Y, Huang R, McAvoy RJ, Li Y (2016) Identification and characterization of histone lysine methylation modifiers in Fragaria vesca. Sci Rep 6:23581. https://doi.org/10.1038/srep23581
Hammond CM, Strømme CB, Huang H, Patel DJ, Groth A (2017) Histone chaperone networks shaping chromatin function. Nat Rev Molecul Cell Biol 18:141–158. https://doi.org/10.1038/nrm.2016.159
Hsieh TF, Ibarra CA, Silva P, Zemach A, Eshed-Williams L, Fischer RL, Zilberman D (2009) Genome-wide demethylation of Arabidopsis endosperm. Science 324:1451–1454. https://doi.org/10.1126/science.1172417
Kim J, Kim JH, Richards EJ, Chung KM, Woo HR (2014) Arabidopsis VIM proteins regulate epigenetic silencing by modulating DNA methylation and histone modification in cooperation with MET1. Mol Plant 7:1470–1485
Kim JM, Sasaki T, Ueda M, Sako K, Seki M (2015) Chromatin changes in response to drought, salinity, heat, and cold stresses in plants. Front Plant Sci 6:114. https://doi.org/10.3389/fpls.2015.00114
Kourani M, Mohareb F, Rezwan FI, Anastasiadi M, Hammond JP (2022) Genetic and physiological responses to heat stress in Brassica napus. Front Plant Sci 13:832147. https://doi.org/10.3389/fpls.2022.832147
Lämke J, Bäurle I (2017) Epigenetic and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants. Genom Biol 18:1–11. https://doi.org/10.1186/s13059-017-1263-6
Lämke J, Brzezinka K, Altmann S, Bäurle I (2016) A hit-and-run heat shock factor governs sustained histone methylation and transcriptional stress memory. The EMBO J. https://doi.org/10.15252/embj.201592593
Lang-Mladek C, Popova O, Kiok K, Berlinger M, Rakic B, Aufsatz W, Jonak C, Hauser MT, Luschnig C (2010) Transgenerational inheritance and resetting of stress-induced loss of epigenetic gene silencing in Arabidopsis. Mol plant 3(3):594–602
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220. https://doi.org/10.1038/nrg2719
Liu J, He Z (2020) Small DNA methylation, big player in plant abiotic stress responses and memory. Front Plant Sci 11:595603. https://doi.org/10.3389/fpls.2020.595603
Liu R, Lang Z (2020) The mechanism and function of active DNA demethylation in plants. J Integr Plant Biol 62:148–159. https://doi.org/10.1111/jipb.12879
Liu P, Zhang S, Zhou B, Luo X, Zhou XF, Cai B, Jin YH, Niu D, Lin J, Cao X, Jin JB (2019) The histone H3K4 demethylase JMJ16 represses leaf senescence in Arabidopsis. Plant Cell 31:430–443. https://doi.org/10.3389/fpls.2020.595603
Mehdi S, Derkacheva M, Ramström M, Kralemann L, Bergquist J, Hennig L (2016) The WD40 domain protein MSI1 functions in a histone deacetylase complex to fine-tune abscisic acid signaling. Plant Cell 28:42–54. https://doi.org/10.1105/tpc.15.00763
Moglia A, Gianoglio S, Acquadro A, Valentino D, Milani AM, Lanteri S, Comino C (2019) Identification of DNA methyltransferases and demethylases in Solanum melongena L., and their transcription dynamics during fruit development and after salt and drought stresses. PLoS ONE 14:e0223581. https://doi.org/10.1371/journal.pone.0223581
Negrão S, Schmöckel SM, Tester M (2017) Evaluating physiological responses of plants to salinity stress. Ann Bot 119:1–11. https://doi.org/10.1093/aob/mcw191
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K (2017) Transcriptional regulatory network of plant heat stress response. Trends Plant Sci 22:53–65. https://doi.org/10.1016/j.tplants.2016.08.015
Ortega-Galisteo AP, Morales-Ruiz T, Ariza RR, Roldán-Arjona T (2008) Arabidopsis DEMETER-LIKE proteins DML2 and DML3 are required for appropriate distribution of DNA methylation marks. Plant Mol Biol 67:671–681. https://doi.org/10.1007/s11103-008-9346-0
Pandey N, Pandey RS (2015) Deciphering UV-B-induced variation in DNA methylation pattern and its influence on regulation of DBR2 expression in Artemisia annua L. Planta 242:869–879. https://doi.org/10.1007/s00425-015-2323-3
Pandey R, MuÈller A, Napoli CA, Selinger DA, Pikaard CS, Richards EJ, Bender J, Mount DW, Jorgensen RA (2002) Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 30:5036–5055. https://doi.org/10.1093/nar/gkf660
Park J, Lim CJ, Shen M, Park HJ, Cha JY, Iniesto E, Rubio V, Mengiste T, Zhu JK, Bressan RA (2018) Epigenetic switch from repressive to permissive chromatin in response to cold stress. Proc Natl Acad Sci 115:E5400–E5409. https://doi.org/10.1073/pnas.172124111
Popova OV, Dinh HQ, Aufsatz W, Jonak C (2013) The RdDM pathway is required for basal heat tolerance in Arabidopsis. Mol Plant 6:396–410. https://doi.org/10.1093/mp/sst023
Ricketts MD, Frederick B, Hoff H, Tang Y, Schultz DC, Singh Rai T, Grazia Vizioli M, Adams PD, Marmorstein R (2015) Ubinuclein-1 confers histone H3. 3-specific-binding by the HIRA histone chaperone complex. Nat Commun 6:7711. https://doi.org/10.1038/ncomms8711
Sani E, Herzyk P, Perrella G, Colot V, Amtmann A (2013) Hyperosmotic priming of Arabidopsis seedlings establishes a long-term somatic memory accompanied by specific changes of the epigenome. Genome Biol 14:1–24. https://doi.org/10.1186/gb-2013-14-6-r59
Secco D, Wang C, Shou H, Schultz MD, Chiarenza S, Nussaume L, Ecker JR, Whelan J, Lister R (2015) Stress induced gene expression drives transient DNA methylation changes at adjacent repetitive elements. Elife 4:e09343. https://doi.org/10.7554/eLife.09343
Secco D, Whelan J, Rouached H, Lister R (2017) Nutrient stress-induced chromatin changes in plants. Curr Plant Biol 39:1–7. https://doi.org/10.1016/j.pbi.2017.04.001
Seleiman MF, Al-Suhaibani N, Ali N, Akmal M, Alotaibi M, Refay Y, Dindaroglu T, Abdul-Wajid HH, Battaglia ML (2021) Drought stress impacts on plants and different approaches to alleviate its adverse effects. Plants 10:259. https://doi.org/10.3390/plants10020259
Shen Y, De Jonge J, Forsberg SK, Pettersson ME, Sheng Z, Hennig L, Carlborg Ö (2014) Natural CMT2 variation is associated with genome-wide methylation changes and temperature seasonality. PLoS Genet 10:e1004842. https://doi.org/10.1371/journal.pgen.1004842
Singh RK, Jaishankar J, Muthamilarasan M, Shweta S, Dangi A, Prasad M (2016) Genome-wide analysis of heat shock proteins in C4 model, foxtail millet identifies potential candidates for crop improvement under abiotic stress. Sci Rep 6:32641. https://doi.org/10.1038/srep32641
Singh J, Mishra V, Wang F, Huang HY, Pikaard CS (2019) Reaction mechanisms of Pol IV, RDR2, and DCL3 drive RNA channeling in the siRNA-directed DNA methylation pathway. Mol Cell 75:576–589. https://doi.org/10.1016/j.molcel.2019.07.008
Song X, Zhang Y, Zhong Q, Zhan K, Bi J, Tang J, Xie J, Li B (2020) Identification and functional characterization of methyl-CpG binding domain protein from Tribolium castaneum. Genomics 112:2223–2232. https://doi.org/10.1016/j.ygeno.2019.12.018
Srikant T, Yuan W, Berendzen KW, Contreras-Garrido A, Drost HG, Schwab R, Weigel D (2022) Canalization of genome-wide transcriptional activity in Arabidopsis thaliana accessions by MET1-dependent CG methylation. Genome Biol 23:263. https://doi.org/10.1186/s13059-022-02833-5
Tang Z, Kang Y, Wang P (2016) Zhao FJ (2016) Phytotoxicity and detoxification mechanism differ among inorganic and methylated arsenic species in Arabidopsis thaliana. Plant Soil 401(243–257):243–257. https://doi.org/10.1007/s11104-015-2739-3
Trujillo M (2018) News from the PUB: plant U-box type E3 ubiquitin ligases. J Exp Bot 69:371–384. https://doi.org/10.1093/jxb/erx411
Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, Zhang Y (2006) Histone demethylation by a family of JmjC domain-containing proteins. Nature 43:811–816. https://doi.org/10.1038/nature04433
Tu YT, Chen CY, Huang YS, Chang CH, Yen MR, Hsieh JWA, Chen PY, Wu K (2022) HISTONE DEACETYLASE 15 and MOS4-associated complex subunits 3A/3B coregulate intron retention of ABA-responsive genes. Plant Physiol 19:882–897. https://doi.org/10.1093/plphys/kiac271
Ueda M, Seki M (2020) Histone modifications form epigenetic regulatory networks to regulate abiotic stress response. Plant Physiol 182:15–26. https://doi.org/10.1104/pp.19.00988
Van Dijk K, Ding Y, Malkaram S, Riethoven JJ, Liu R, Yang J, Laczko P, Chen H, Xia Y, Ladunga I, Avramova Z (2010) Dynamic changes in genome-wide histone H3 lysine 4 methylation patterns in response to dehydration stress in Arabidopsis thaliana. BMC Plant Biol 10:1–2. https://doi.org/10.1186/1471-2229-10-238
Virlouvet L, Ding Y, Fujii H, Avramova Z, Fromm M (2014) ABA signaling is necessary but not sufficient for RD 29 B transcriptional memory during successive dehydration stresses in Arabidopsis thaliana. The Plant J 79:150–161
Wang X, Vignjevic M, Jiang D, Jacobsen S, Wollenweber B (2014) Improved tolerance to drought stress after anthesis due to priming before anthesis in wheat (Triticum aestivum L.) var. Vinjett. J Exp Bot 65:6441–6456. https://doi.org/10.1093/jxb/eru362
Wang Z, Casas-Mollano JA, Xu J, Riethoven JJ, Zhang C, Cerutti H (2015) Osmotic stress induces phosphorylation of histone H3 at threonine 3 in pericentromeric regions of Arabidopsis thaliana. Proc Natl Acad Sci 112:8487–8492. https://doi.org/10.1073/pnas.1423325112
Wang Q, Liu P, Jing H, Zhou XF, Zhao B, Li Y, Jin JB (2021) JMJ27-mediated histone H3K9 demethylation positively regulates drought-stress responses in Arabidopsis. New Phytol 232:221–236. https://doi.org/10.1111/tpj.12548
Waqas MA, Kaya C, Riaz A, Farooq M, Nawaz I, Wilkes A, Li Y (2019) Potential mechanisms of abiotic stress tolerance in crop plants induced by thiourea. Front Plant Sci 10:1336. https://doi.org/10.3389/fpls.2019.01336
Widiez T, El Kafafi ES, Girin T, Berr A, Ruffel S, Krouk G, Vayssières A, Shen W-H, Coruzzi GM, Gojon A (2011) High nitrogen insensitive 9 (HNI9)-mediated systemic repression of root NO3− uptake is associated with changes in histone methylation. Proc Natl Acad Sci 108:13329–13334. https://doi.org/10.1073/pnas.101786310
Xiao J, Zhang H, Xing L, Xu S, Liu H, Chong K, Xu Y (2013) Requirement of histone acetyltransferases HAM1 and HAM2 for epigenetic modification of FLC in regulating flowering in Arabidopsis. J Plant Physiol 170:444–451. https://doi.org/10.1016/j.jplph.2012.11.007
Xie H, Sun Y, Cheng B, Xue S, Cheng D, Liu L, Meng L, Qiang S (2019) Variation in ICE1 methylation primarily determines phenotypic variation in freezing tolerance in Arabidopsis thaliana. Plant Cell Physiol 60:152–165. https://doi.org/10.1093/pcp/pcy197
Xing J, Wang T, Liu Z, Xu J, Yao Y, Hu Z, Peng H, Xin M, Yu F, Zhou D (2015) GENERAL CONTROL NONREPRESSED PROTEIN5-mediated histone acetylation of FERRIC REDUCTASE DEFECTIVE3 contributes to iron homeostasis in Arabidopsis. Plant Physiol 168:1309–1320. https://doi.org/10.1104/pp.15.00397
Xu Y, Zhang S, Lin S, Guo Y, Deng W, Zhang Y, Xue Y (2017) WERAM: a database of writers, erasers and readers of histone acetylation and methylation in eukaryotes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw1011
Xu J, Zhou S, Gong X, Song Y, van Nocker S, Ma F, Guan Q (2018) Single-base methylome analysis reveals dynamic epigenomic differences associated with water deficit in apple. Plant Biotechnol J 16:672–687. https://doi.org/10.1111/pbi.12820
Yang R, Zheng Z, Chen Q, Yang L, Huang H, Miki D, Wu W, Zeng L, Liu J, Zhou JX (2017) The developmental regulator PKL is required to maintain correct DNA methylation patterns at RNA-directed DNA methylation loci. Genome Biol 18:1–18. https://doi.org/10.1186/s13059-017-1226-y
Yang DL, Zhang G, Wang L, Li J, Xu D, Di C, Tang K, Yang L, Zeng L, Miki D, Duan CG (2018) Four putative SWI2/SNF2 chromatin remodelers have dual roles in regulating DNA methylation in Arabidopsis. Cell Discov 4:55. https://doi.org/10.1038/s41421-018-0056-8
Yang R, Hong Y, Ren Z, Tang K, Zhang H, Zhu JK, Zhao C (2019) A role for PICKLE in the regulation of cold and salt stress tolerance in Arabidopsis. Front Plant Sci 10:900
Yolcu S, Ozdemir F, Güler A, Bor M (2016) Histone acetylation influences the transcriptional activation of POX in Beta vulgaris L. and Beta maritima L. under salt stress. Plant Physiol Biochem 100:37–46. https://doi.org/10.1016/j.plaphy.2015.12.019
Zhang H, Lang Z, Zhu JK (2018) Dynamics and function of DNA methylation in plants. Nat Rev Mol Cell Biol 19:489–506. https://doi.org/10.1038/s41580-018-0016-z
Zhang W, Wang N, Yang J, Guo H, Liu Z, Zheng X, Li S, Xiang F (2020) The salt-induced transcription factor GmMYB84 confers salinity tolerance in soybean. Plant Sci 291:110326
Zhao F, Zhang H, Zhao T, Li Z, Jiang D (2021) The histone variant H3. 3 promotes the active chromatin state to repress flowering in Arabidopsis. Plant Physiol 186:2051–2063. https://doi.org/10.1093/plphys/kiab224
Zheng X, Chen L, Xia H, Wei H, Lou Q, Li M, Li T, Luo L (2017) Transgenerational epimutations induced by multi-generation drought imposition mediate rice plant’s adaptation to drought condition. Sci Rep 7:39843. https://doi.org/10.1038/srep39843
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zubko E, Gentry M, Kunova A, Meyer P (2012) De novo DNA methylation activity of METHYLTRANSFERASE 1 (MET1) partially restores body methylation in Arabidopsis thaliana. Plant J 71:1029–1037. https://doi.org/10.1111/j.1365-313X.2012.05051.x
Acknowledgements
This work was supported by the core grant of National Institute of Plant Genome Research and research grant from Department of Biotechnology (BT/PR36694/NNT/281722/2020) to AP. HC and JN acknowledge University Grants Commission and Council of Scientific and Industrial Research, Government of India for Junior and Senior Research Fellowships, respectively. PKT acknowledges Science and Engineering Research Board, New Delhi for JC Bose National Fellowship (JCB/2021/000036). The authors are thankful to DBT-eLibrary Consortium (DeLCON) for providing access to e-resources. CSIR-CIMAP Publication Number: CIMAP/PUB/ 2024/58.
Funding
Funding was provided by CIMAP (Grant No. JCB/2021/000036).
Author information
Authors and Affiliations
Contributions
AP and PKT conceived the idea. HC, JN and AP wrote the first draft. AP and PKT finalized the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chhatwal, H., Naik, J., Pandey, A. et al. Broadening the epigenetic horizon of abiotic stress response in plants. Plant Growth Regul (2024). https://doi.org/10.1007/s10725-024-01152-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10725-024-01152-y