Abstract
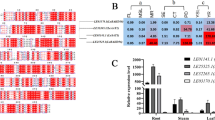
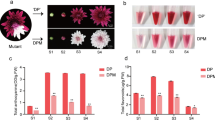
Despite the nutritional and economic importance of carotenoids and their products, regulation of carotenoid content in leaves remains to be fully elucidated. Recent findings indicate that carotenoid content are determined, at least in part, by the activity of carotenoid cleavage dioxygenase 1 (CCD1). This study examined whether NtCCD1 affects leaf carotenoids in the model plant Nicotiana tabacum. Three NtCCD1s, NtCCD1a, NtCCD1b, and NtCCD1c, were investigated including their phylogenetic relationships, conserved motifs, and exon-intron architecture. Of the three transcripts, NtCCD1c exhibited the highest expression level in various tissue as determined by real-time PCR. Confocal microscopy indicated that all NtCCD1s are located in the cytoplasm. The enzymatic activity of the NtCCD1c protein was also studied by co-expressing NtCCD1c in Escherichia coli engineered to accumulate β-carotene or lycopene. SPME-GC-MS indicated that NtCCD1c cleaves β-carotene and lycopene specifically at the 9–10/9′-10′ site to produce β-ionone and pseudoionone, respectively. Virus-induced gene silencing (VIGS) of NbCCD1 increased carotenoid contents including β-carotene, α-carotene, zeaxanthin, phytoene, lutein, violaxanthin, neoxanthin, and β-cryptoxanthin, along with a decline in reactive oxygen species (ROS). RNA-seq of silenced plants suppressed NbCCD1 but enhanced expression of essential genes encoding antioxidant enzymes including NtGST, NtCuZnSOD, NtAAO, and NtPOD, along with a decrease in expression of genes related to carbon fixation such as NtRCA1, NtRCA2, and NtRBCS. These observations suggest that NtCCD1 is a negative regulator of carotenoid content and plays an essential role in the regulation of ROS levels in tobacco leaves.









Similar content being viewed by others
Data availability
The datasets generated or analyzed during the current study are included in this article and its supplementary information files.
References
Allison B, Combs S, Deluca S, Lemmon G, Mizoue L, Meiler J (2014) Computational design of protein-small molecule interfaces. J Struct Biol 185(2):193–202. https://doi.org/10.1016/j.jsb.2013.08.003
Auldridge ME, Block A, Vogel JT, Dabney-Smith C, Mila I, Bouzayen M, Magallanes-Lundback M et al (2006a) Characterization of three members of the Arabidopsis carotenoid cleavage dioxygenase family demonstrates the divergent roles of this multifunctional enzyme family. Plant J 45(6):982–993. https://doi.org/10.1111/j.1365-313X.2006.02666.x
Auldridge ME, Mccarty DR, Klee HJ (2006b) Plant carotenoid cleavage oxygenases and their apocarotenoid products. Curr Opin Plant Biol 9(3):315–321. https://doi.org/10.1016/j.pbi.2006.03.005
Bouvier F, Isner JC, Dogbo O, Camara B (2005) Oxidative tailoring of carotenoids: a prospect towards novel functions in plants. Trends Plant Sci 10(4):187–194. https://doi.org/10.1016/j.tplants.2005.02.007
Bouvier F, Suire C, Mutterer J, Camara B (2003) Oxidative remodeling of chromoplast carotenoids: identification of the carotenoid dioxygenase CsCCD and CsZCD genes involved in Crocus secondary metabolite biogenesis. Plant Cell 15(1):47–62. https://doi.org/10.1105/tpc.006536
Brandi F, Bar E, Mourgues F, Horvath G, Turcsi E, Giuliano G, Liverani A et al (2011) Study of ‘redhaven’ peach and its white-fleshed mutant suggests a key role of CCD4 carotenoid dioxygenase in carotenoid and norisoprenoid volatile metabolism. BMC Plant Biol 11:24. https://doi.org/10.1186/1471-2229-11-24
Campbell R, Ducreux LJ, Morris WL, Morris JA, Suttle JC, Ramsay G, Bryan GJ et al (2010) The metabolic and developmental roles of carotenoid cleavage dioxygenase4 from potato. Plant Physiol 154(2):656–664. https://doi.org/10.1104/pp.110.158733
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen S, Zhou Y, Chen Y, Gu J (2018) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Cheng GT, Li YS, Qi SM, Wang J, Zhao P, Lou QQ, Wang YF et al (2021) SlCCD1A enhances the aroma quality of tomato fruits by promoting the synthesis of carotenoid-derived volatiles. Foods 10(11). https://doi.org/10.3390/foods10112678
Chiou CY, Pan HA, Chuang YN, Yeh KW (2010) Differential expression of carotenoid-related genes determines diversified carotenoid coloration in floral tissues of Oncidium cultivars. Planta 232(4):937–948. https://doi.org/10.1007/s00425-010-1222-x
Cunningham FJ, Pogson B, Sun Z, Mcdonald KA, Dellapenna D, Gantt E (1996) Functional analysis of the beta and epsilon lycopene cyclase enzymes of Arabidopsis reveals a mechanism for control of cyclic carotenoid formation. Plant Cell 8(9):1613–1626. https://doi.org/10.1105/tpc.8.9.1613
Dhar MK, Mishra S, Bhat A, Chib S, Kaul S (2020) Plant carotenoid cleavage oxygenases: structure-function relationships and role in development and metabolism. Brief Funct Genomics 19(1):1–9. https://doi.org/10.1093/bfgp/elz037
Domonkos I, Kis M, Gombos Z, Ughy B (2013) Carotenoids, versatile components of oxygenic photosynthesis. Prog Lipid Res 52(4):539–561. https://doi.org/10.1016/j.plipres.2013.07.001
Dong C, Qu G, Guo J, Wei F, Gao S, Sun Z, Jin L et al (2022a) Rational design of geranylgeranyl diphosphate synthase enhances carotenoid production and improves photosynthetic efficiency in Nicotiana tabacum. Sci Bull 67(3):315–327. https://doi.org/10.1016/j.scib.2021.07.003
Dong C, Zhang M, Wei F, Guo Y, Qin L, Wei P, Wang Y et al (2022b) Inhibition of red chlorophyll catabolite reductase improved chlorophyll and carotenoid synthesis in tobacco. Plant Cell Tissue Organ Cult 148(3):687–698. https://doi.org/10.1007/s11240-022-02228-1
Floss DS, Walter MH (2009) Role of carotenoid cleavage dioxygenase 1 (CCD1) in apocarotenoid biogenesis revisited. Plant Signal Behav 4(3):172–175. https://doi.org/10.4161/psb.4.3.7840
Frusciante S, Diretto G, Bruno M, Ferrante P, Pietrella M, Prado-Cabrero A, Rubio-Moraga A et al (2014) Novel carotenoid cleavage dioxygenase catalyzes the first dedicated step in saffron crocin biosynthesis. Proc Natl Acad Sci U S A 111(33):12246–12251. https://doi.org/10.1073/pnas.1404629111
Gao J, Yang S, Tang K, Li G, Gao X, Liu B, Wang S et al (2021) GmCCD4 controls carotenoid content in soybeans. Plant Biotechnol J 19(4):801–813. https://doi.org/10.1111/pbi.13506
Gonzalez-Jorge S, Ha SH, Magallanes-Lundback M, Gilliland LU, Zhou A, Lipka AE, Nguyen YN et al (2013) Carotenoid cleavage dioxygenase4 is a negative regulator of beta-carotene content in Arabidopsis seeds. Plant Cell 25(12):4812–4826. https://doi.org/10.1105/tpc.113.119677
Havaux M (2014) Carotenoid oxidation products as stress signals in plants. Plant J 79(4):597–606. https://doi.org/10.1111/tpj.12386
Hou X, Rivers J, Leon P, Mcquinn RP, Pogson BJ (2016) Synthesis and function of apocarotenoid signals in plants. Trends Plant Sci 21(9):792–803. https://doi.org/10.1016/j.tplants.2016.06.001
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297. https://doi.org/10.1016/j.tplants.2016.06.001
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12(4):357–360. https://doi.org/10.1038/nmeth.3317
Ko MR, Song MH, Kim JK, Baek SA, You MK, Lim SH, Ha SH (2018) RNAi-mediated suppression of three carotenoid-cleavage dioxygenase genes, OsCCD1, 4a, and 4b, increases carotenoid content in rice. J Exp Bot 69(21):5105–5116. https://doi.org/10.1093/jxb/ery300
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Lu S, Li L (2008) Carotenoid metabolism: biosynthesis, regulation, and beyond. J Integr Plant Biol 50(7):778–785. https://doi.org/10.1111/j.1744-7909.2008.00708.x
Messing SA, Gabelli SB, Echeverria I, Vogel JT, Guan JC, Tan BC, Klee HJ et al (2010) Structural insights into maize viviparous14, a key enzyme in the biosynthesis of the phytohormone abscisic acid. Plant Cell 22(9):2970–2980. https://doi.org/10.1105/tpc.110.074815
Moise AR, von Lintig J, Palczewski K (2005) Related enzymes solve evolutionarily recurrent problems in the metabolism of carotenoids. Trends Plant Sci 10(4):178–186. https://doi.org/10.1016/j.tplants.2005.02.006
Nguyen T, Masuda J, Miyajima I, Thien NQ, Mojtahedi N, Hiramatsu M, Kim JH et al (2012) Involvement of carotenoid cleavage dioxygenase 4 gene in tepal color change in Lilium brownii var. Colchesteri. J Jpn Soci Hortic 81(4):366–373. https://doi.org/10.2503/JJSHS1.81.366
Niyogi KK, Truong TB (2013) Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis. Curr Opin Plant Biol 16(3):307–314. https://doi.org/10.1016/j.pbi.2013.03.011
Ohmiya A, Kishimoto S, Aida R, Yoshioka S, Sumitomo K (2006) Carotenoid cleavage dioxygenase (CmCCD4a) contributes to white color formation in chrysanthemum petals. Plant Physiol 142(3):1193–1201. https://doi.org/10.1104/pp.106.087130
Paparella A, Shaltiel-Harpaza L, Ibdah M (2021) Beta-ionone: its occurrence and biological function and metabolic engineering. Plants (Basel) 10(4). https://doi.org/10.3390/plants10040754
Park H, Bradley P, Greisen PJ, Liu Y, Mulligan VK, Kim DE, Baker D et al (2016) Simultaneous optimization of biomolecular energy functions on features from small molecules and macromolecules. J Chem Theory Comput 12(12):6201–6212. https://doi.org/10.1021/acs.jctc.6b00819
Priya R, Siva R (2014) Phylogenetic analysis and evolutionary studies of plant carotenoid cleavage dioxygenase gene. Gene 548(2):223–233. https://doi.org/10.1016/j.gene.2014.07.037
Qi Z, Fan X, Zhu C, Chang D, Pei J, Zhao L (2022) Overexpression and characterization of a novel plant carotenoid cleavage dioxygenase 1 from Morus notabilis. Chem Biodivers 19(2):e202100735. https://doi.org/10.1002/cbdv.202100735
Schwartz SH, Qin X, Zeevaart JA (2001) Characterization of a novel carotenoid cleavage dioxygenase from plants. J Biol Chem 276(27):25208–25211. https://doi.org/10.1074/jbc.M102146200
Simkin AJ (2021) Carotenoids and apocarotenoids in planta: their role in plant development, contribution to the flavour and aroma of fruits and flowers, and their nutraceutical benefits. Plants (Basel) 10(11). https://doi.org/10.3390/plants10112321
Simkin AJ, Schwartz SH, Auldridge M, Taylor MG, Klee HJ (2004) The tomato carotenoid cleavage dioxygenase 1 genes contribute to the formation of the flavor volatiles beta-ionone, pseudoionone, and geranylacetone. Plant J 40(6):882–892. https://doi.org/10.1111/j.1365-313X.2004.02263.x
Sun Z, Hans J, Walter MH, Matusova R, Beekwilder J, Verstappen FW, Ming Z et al (2008) Cloning and characterisation of a maize carotenoid cleavage dioxygenase (ZmCCD1) and its involvement in the biosynthesis of apocarotenoids with various roles in mutualistic and parasitic interactions. Planta 228(5):789–801. https://doi.org/10.1007/s00425-008-0781-6
Tan BC, Joseph LM, Deng WT, Liu L, Li QB, Cline K, Mccarty DR (2003) Molecular characterization of the Arabidopsis 9-cis epoxycarotenoid dioxygenase gene family. Plant J 35(1):44–56. https://doi.org/10.1046/j.1365-313x.2003.01786.x
Tian L (2015) Recent advances in understanding carotenoid-derived signaling molecules in regulating plant growth and development. Front Plant Sci 6:790. https://doi.org/10.3389/fpls.2015.00790
Vogel JT, Tan BC, Mccarty DR, Klee HJ (2008) The carotenoid cleavage dioxygenase 1 enzyme has broad substrate specificity, cleaving multiple carotenoids at two different bond positions. J Biol Chem 283(17):11364–11373. https://doi.org/10.1074/jbc.M710106200
Walter MH, Strack D (2011) Carotenoids and their cleavage products: biosynthesis and functions. Nat Prod Rep 28(4):663–692. https://doi.org/10.1039/c0np00036a
Yahyaa M, Bar E, Dubey NK, Meir A, Davidovich-Rikanati R, Hirschberg J, Aly R et al (2013) Formation of norisoprenoid flavor compounds in carrot (Daucus carota L.) roots: characterization of a cyclic-specific carotenoid cleavage dioxygenase 1 gene. J Agric Food Chem 61(50):12244–12252. https://doi.org/10.1021/jf404085k
Zhou Q, Li Q, Li P, Zhang S, Liu C, Jin J, Cao P et al (2019) Carotenoid cleavage dioxygenases: identification, expression, and evolutionary analysis of this gene family in tobacco. Int J Mol Sci 20(22). https://doi.org/10.3390/ijms20225796
Funding
The research leading to these results has received funding from the Cultivation Program for Young Backbone Teachers in Henan University of Technology, Science Project 110202101042 (JY 19)/ 2022530000241007, and 110202102033.
Author information
Authors and Affiliations
Contributions
Chen Dong and Yulong Gao designed the study and prepared the manuscript. Fei Du, Zongyu Hu and Lili Qin performed the experiments and modified the manuscript. Chen Zhang, Zhiwen Wang and Yongchun Shi analyzed the data. Xiaoran Wang and Ran Wang supervised the study. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Paul Holford.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Fei Du, Zongyu Hu and Lili Qin these authors contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Du, F., Hu, Z., Qin, L. et al. Suppression of carotenoid cleavage dioxygenase 1 (NtCCD1) increases carotenoid contents and attenuates reactive oxygen species (ROS) in Tobacco Leaves. Plant Growth Regul 100, 667–679 (2023). https://doi.org/10.1007/s10725-023-00961-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-00961-x