Abstract
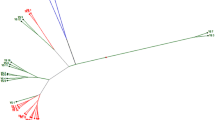
Turnip of the Brassica genus is a traditional crop that is widely cultivated in farming and farming-pastoral regions in Tibet. It is mostly utilized as animal fodder and vegetable around the Tibetan plateau. Given their high altitudes and variable climate types, different regions of Tibet plateau are home to a rich diversity of turnip. A mass of studies related to genetic diversity of highland crops have been carried out in recent years. However, the genetic diversity of Tibetan turnip remains to be elucidated to date. In this study, we performed morphological investigation and simple sequence repeat analysis, to characterize the genetic diversity and population structure of Tibetan turnips. Then, we explored the physiological response of seedlings under low-temperature stress. We also performed a field experiment, to identify clubroot-resistant cultivars. Results showed significant differences in phenotypic traits among different sources of 25 Tibetan turnip accessions. The genetic similarity dendrogram showed that the genetic distance in the 25 accessions was between 0.61 and 0.77. When the similarity coefficient is between 0.644 and 0.646, they can be divided into three subgroups, and they presented a certain relationship between geographical origin and its genetic distance. The central diffusion location suggests that the genetic diversity and population structure of these Tibetan turnip accessions are consistent with their geographical origin. In addition, the potential clubroot resistance accession is 156-13, and the chilling resistance accessions are 156-8 and 156-10. These results suggest the Tibetan turnip is a valuable resource for the genetic analysis and breeding of new clubroot-resistant cultivars.






Similar content being viewed by others
References
Agarwal M, Shrivastava N, Padh H (2008) Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep 27:617–631
Aissiou F, Laperche A, Falentin C, Lodé M, Deniot G, Boutet G, Régnier F, Trotoux G, Huteau V, Coriton O, Rousseau-Guentin M, Abrous O, Chèvre AM, Hadj-Arab H (2018) A novel Brassica rapa L. genetic diversity found in Algeria. Euphytica 214:241
Andersen NS, Poulsen G, Andersen BA, Kiær LP, D’Hertefeldt T, Wilkinson MJ, Jørgensen RB (2009) Processes affecting genetic structure and conservation: a case study of wild and cultivated Brassica rapa. Genet Resour Crop Evol 56:189–200
Ashworth EN (1992) Formation and spread of ice in plant tissues. Hortic Rev 13:215–255
Bjørnstad Å, Aastveit K (1990) Pleiotropic effects on the ml-o mildew resistance gene in barley in different genetical backgrounds. Euphytica 46:217–226
Bradshaw JE, Gemmell DJ, Wilson RN (2010) Transfer of resistance to clubroot (Plasmodiophora brassicae) to swedes (Brassica napus L. var. napobrassica Peterm) from B. rapa. Ann Appl Biol 130:337–348
Cao JS, Cao SC, Miao Y, Lu G (1997) Cldistic operational analysis and study on the evolution of Chinese cabbage groups (Brassica campestris L.). Acta Hortic Sin 24(1):35–42
Chen J, Jing J, Zhan Z, Zhang T, Zhang C, Piao Z (2013) Identification of novel QTLs for isolate-specific partial resistance to Plasmodiophora brassicae in Brassica rapa. PLoS ONE 8:e85307
Committee of Chinese herbal medicine (2002) Chinese herbal medicine—Tibetan medicine. Science & Technology Press of Shanghai, Shanghai, pp 349–350
De Barba M, Miquel C, Lobreaux S, Quenette PY, Swenson JE, Taberlet P (2017) High-throughput microsatellite genotyping in ecology: improved accuracy, efficiency, standardization and success with low-quantity and degraded DNA. Mol Ecol Resour 17:492–507
Delfina B, Alessandro T, Francesca D, Andrea V, Patrizia V, Giampiero V, Luigi C (2016) Next generation breeding. Plant Sci 242:3–13
Diederichsen E, Frauen M, Linders EGA, Hatakeyama K, Hirai M (2009) Status and perspectives of clubroot resistance breeding in crucifer crops. J Plant Growth Regul 28:265–281
Dixon GR, Dixon GR (2009) Plasmodiophora brassicae in its environment. J Plant Growth Regul 28:212–228
Donald C, Porter I (2009) Integrated control of clubroot. J Plant Growth Regul 28:289–303
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Gao JT, Li N, Xuan ZY, Yang WC (2017) Genetic diversity among “Qamgur” varieties in China revealed by SSR markers. Euphytica 213:204
Goodwin S, McPherson JD, McCombie WR (2016) Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet 17:333–351
Hammer O, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
Hasan MJ, Strelkov SE, Howard RJ, Rahman H (2012) Screening of Brassica germplasm for resistance to Plasmodiophora brassicae pathotypes prevalent in Canada for broadening diversity in clubroot resistance. Can J Plant Sci 92:501–515
Hatakeyama K, Niwa T, Kato T, Ohara T, Kakizaki T, Matsumoto S (2017) The tandem repeated organization of NB-LRR genes in the clubroot-resistant CRb locus in Brassica rapa L. Mol Genet Genom 292:397–405
Heber U (1958) Ursachen der frostresistenz bei winterweizen. Planta 52:431–446
Howard R, Strelkov S, Harding M (2010) Clubroot of cruciferous crops: new perspectives on an old disease. Can J Plant Pathol 32:43–57
Hughes MA, Dunn MA (1996) The molecular biology of plant acclimation to low temperature. J Exp Bot 47:291–305
Kalia RK, Rai MK, Kalia S, Singh R, Dhawan AK (2011) Microsatellite markers: an overview of the recent progress in plants. Euphytica 177:309–334
Kaur S, Panesar PS, Bera MB, Kaur V (2015) Simple sequence repeat markers in genetic divergence and marker-assisted selection of rice cultivars: a review. Crit Rev Food Sci Nutr 55:41–49
Kuan KC (1985) Cruciferae. In: Wu CY (ed) Flora Xizangica, vol 2. Science Press of China, Beijing, pp 323–412
Kuroda H, Sagisaka S (2008) Participation of hydrogen peroxide in the dysfunction of peroxide-scavenging systems in apple flower buds associated with freezing injury. Engei Gakkai Zasshi 67:161–165
Li JW (1981) The origins and evolution of vegetable crops in China. Sci Agric Sin 14:90–95
Li XX, Shen D (2008) Descriptors and data standard for root and stem mustard (Brassica juncea Coss.). Chinese Agricultural Press, Beijing, pp 1–82
Liu J, Cui H (1995) Electrical conductivity method for chilling-resistance evaluation in cucumber. J Northwest Sci-Tech Univ Agric For 23(4):74–77
Liu H, Jiang SP, Yang LL, Chen ZJ, Bao SF, Xu AG, Bu HT, Gao P (2012) Hypoglycemic effect of crude saponins of turnip (Brassica rapa L.) on diabetic mice. J Northwest A F Univ 40(6):23–27
Liu SR, Li WY, Dang L, Hu CG, Zhang JZ (2013) Development and characterization of genomic and expressed SSRs in citrus by genome-wide analysis. PLoS ONE 8:e75149
Manzanares-Dauleux MJ, Delourme R, Baron F, Thomas G (2000) Mapping of one major gene and of QTLs involved in resistance to clubroot in Brassica napus. Theor Appl Genet 101:885–891
Meng FJ, Liu L, Peng M, Wang ZK, Wang C, Zhao YY (2015) Genetic diversity and population structure analysis in wild strawberry (Fragaria nubicola L.) from Motuo in Tibet Plateau based on simple sequence repeats (SSRs). Biochem Syst Ecol 63:113–118
Meng Z, Song F, Liu T (2018) Genetic diversity and genetic structure analysis of maize (Zea mays) landraces in Tibet. Int J Agric Biol 20:791–798
Neik TX, Barbetti MJ, Batley J (2017) Current status and challenges in identifying disease resistance genes in Brassica napus. Front Plant Sci 8:1788
Padilla G, Cartea ME, Rodriguez VM, Ordas A (2005) Genetic diversity in a germplasm collection of Brassica rapa subsp. rapa L. from northwestern Spain. Euphytica 145:171–180
Pan X, Cao Q, Wang G (2002) Evaluation of lipid peroxidation for use in selection of cold hardiness cultivars of Almond. Acta Ecol Sin 22:1902–1911
Pang W, Shan L, Li X, Li P, Sha Y, Yong PL, Piao Z (2014) Genetic detection of clubroot resistance loci in a new population of Brassica rapa. Hortic Environ Biotechnol 55:540–547
Peng G, Lahlali R, Hwang SF, Pageau D, Hynes RK, Mcdonald MR, Gossen BD, Strelkov SE (2014) Crop rotation, cultivar resistance, and fungicides/biofungicides for managing clubroot (Plasmodiophora brassicae) on canola. Can J Plant Pathol 36:99–112
Porrashurtado L, Ruiz Y, Santos C, Phillips C, Carracedo A, Lareu MV (2013) An overview of STRUCTURE: applications, parameter settings, and supporting software. Front Genet 4:98
Prasad TK, Anderson MD, Martin BA, Stewart CR (1994) Evidence for chilling-induced oxidative stress in maize seedlings and a regulatory role for hydrogen peroxide. Plant Cell 6:65–74
Rieger M (1989) Freeze protection for horticultural crops. Hortic Rev 11:45–109
Soengas P, Cartea ME, Francisco M, Lema M, Velasco P (2011) Genetic structure and diversity of a collection of Brassica rapa subsp. rapa L. revealed by simple sequence repeat markers. J Agric Sci 149:617–624
Strelkov SE, Hwang SF, Manolii VP, Cao T, Feindel D (2016) Emergence of new virulence phenotypes of Plasmodiophora brassicae on canola (Brassica napus) in Alberta, Canada. Eur J Plant Pathol 145:1–13
Takahashi Y, Yokoi S, Takahata Y (2016) Genetic divergence of turnip (Brassica rapa L. em. Metzg. subsp. rapa) inferred from simple sequence repeats in chloroplast and nuclear genomes and morphology. Genet Resour Crop Evol 63:869–879
Tang WM, Chu BQ, Gao WL, Wu CJ, Gong LX, Dai XL, Zhang Y (2015) Comparative studies of chemical compositions and antioxidant capacity of low-polarity components from Tibetan turnip (Brassica rapa L.) and Maca (Lepidium meyenii Walp). Nat Prod Res Dev 27:674–680
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23:48–55
Wang JL, Dan B, Hu SY, Tu JX, Luan YF, Meng X, Zhuo G, Nima ZM, Tang L (2002) RAPD analysis for the genetic diversity of Brassica rapa in Tibet. Acta Genet Sin 29:1021–1027
Wang JL, Luan YF, Daci ZG, Chang TJ (2006) Geographical distribution and biological characters of wild rapeseed in Tibet. Chin J Oil Crop Sci 28(2):134–137
Wang JL, Dan B, Cheng HH, Fang HL, Chang TJ, Li P, Luan YF (2008) AFLP analysis on genetic diversity of wild and cultivated rapeseed (B. campestris and B. juncea) in Tibet. Chin J Oil Crop Sci 30:10–16
Williams PH (1996) A system for the determination of races of Plasmodiophora brassicae that infect cabbage and rutabaga. Phytopathology 56:624–626
Wise RR, Naylor AW (1987) Chilling-enhanced photooxidation: evidence for the role of singlet oxygen and superoxide in the breakdown of pigments and endogenous antioxidants. Plant Physiol 83:278–282
Wit F, Weg MVD (1964) Clubroot-resistance in turnips (Brassica campestris L.). Euphytica 13:9–18
Wu CY, Lu AM, Tang YC (2003) The families and genera of angiosperms in China. Science Press of China, Beijing, pp 504–521
Xu ZX, Gong TL, Zhao FF (2006) Analysis of climate change in Tibetan Plateau over the past 40 years. J Subtrop Resour Environ 1(1):24–32
Yada B, Alajo A, Ssemakula GN, Brown-Guedira G, Otema MA, Stevenson PC, Mwanga ROM, Yencho GC (2017) Identification of simple sequence repeat markers for sweetpotato weevil resistance. Euphytica 213:129
Yildirim E, Yildirim N, Ercisli S, Agar G, Karlidag H (2010) Genetic relationships among turnip (Brassica rapa var. rapa) genotypes. Genet Mol Res 9:987
Yu FQ, Zhang XG, Huang Z, Chu MG, Song T, Falk KC, Deora A, Chen QL, Zhang Y, McGregor L, Gossen BD, McDonald MR, Peng G (2016) Identification of genome-wide variants and discovery of variants associated with Brassica rapa clubroot resistance gene Rcr1 through bulked segregant RNA sequencing. PLoS ONE 11(4):e0153218
Yuan H, Zeng X, Xu Q, Wang Y, Zha S, Nyima T (2018) Genetic diversity of germplasm resource and core collection development in hulless barley. J Triticeae Crops 38:922–928
Zhan L, Paterson IG, Fraser BA, Watson B, Bradbury IR, Ravindran PN, Reznick D, Beiko RG, Bentzen P (2017) MEGASAT: automated inference of microsatellite genotypes from sequence data. Mol Ecol Resour 17:247–256
Zhang Y, Tang WM, Ma NI, Li JJ, Zhu MD, Gong LX, Chen XJ, Liu YF, Bian B, Wu XQ (2014) Effects of Tibetan turnip and its processed products on human tolerance to hypoxia. Food Sci 35:178–182
Zhu H, Song P, Koo DH, Guo L, Li Y, Sun S, Weng Y, Yang L (2016) Genome wide characterization of simple sequence repeats in watermelon genome and their application in comparative mapping and genetic diversity analysis. BMC Genom 17:557
Acknowledgements
The authors gratefully thank those students in College of Plant Science, Tibet Agricultural and Animal Husbandry University for collecting the seeds of turnips from the different counties in Tibet. This work was partially supported by National Natural Science Foundation of China (31460521), the Breeding Project of the Sci-tech Foundation of Zhejiang Province (2016C02051-6-1), the Project of SRTP in Zhejiang University (2018R401080), the Project of Sci-tech Foundation of Jiangsu Province (BE2018310), and the Project of Application on Public Welfare Technology in Zhejiang Province (LGN18C150003). The authors thank Dr. Zhenning Liu for his critical reading the manuscript, also thank the anonymous reviewers for their critical comments and suggestions on manuscript revision.
Author information
Authors and Affiliations
Contributions
XY and WG proposed and designed the research. YG, LZ, YZ and YG performed the experiments. YG, RL, JL, KL and KZ performed the statistical analyses, interpretation of experimental results. YG and XY wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10722_2019_824_MOESM1_ESM.jpg
Figure S1. Results of malondialdehyde content under the different low-temperatures stresses (5 °C, 15 °C, and 25 °C). Bands in blue, orange, and green present 5 °C, 15 °C, and 25 °C respectively. Error bar represent standard error (three repetitions) (JPEG 894 kb)
10722_2019_824_MOESM2_ESM.jpg
Figure S2. Results of soluble sugar content under different low-temperatures stress (5 °C, 15 °C and 25 °C). Bands in blue, orange, and green present 5 °C, 15 °C, and 25 °C respectively. Error bar represent standard error (three repetitions). (JPEG 905 kb)
Rights and permissions
About this article
Cite this article
Gao, Y., Gong, W., Li, R. et al. Genetic diversity analysis of Tibetan turnip(Brassica rapa L. ssp. rapifera Matzg) revealed by morphological, physiological, and molecular marker. Genet Resour Crop Evol 67, 209–223 (2020). https://doi.org/10.1007/s10722-019-00824-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-019-00824-3