Abstract
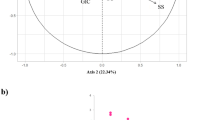
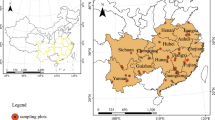
This study examines the relationship between spike morphology and natural habitat for 84 accessions of four Aegilops species, belongs to section Sitopsis, Ae. bicornis, Ae. longissima, Ae. searsii, and Ae. sharonensis in genus Aegilops, section Sitopsis, wild relatives of Triticum aestivum L. These species are considered valuable genetic resources for future cultivation and breeding of domesticated wheat. The goals of the study were to: (1) document variation in spike morphology among these four species; (2) examine the relationship between spike morphology and native habitat; (3) document geographical distribution of distinct spike morphology; and (4) examine the relationship between spike morphology and heading time and value for these four species. The results reveal significant differences in spike morphology among species of section Sitopsis. The most noteworthy variation involved the absence/presence of lateral awn, such that species with lateral awn were restricted in coastal, though species without lateral awn were mainly distributed in inland. This suggests that local climate may be a determinant of variation in lateral awn, and that this trait may be subject to convergent evolution. Differences in heading time in sympatric area were also observed. The differences may enhance species divergence and could represent a lead speciation event. The results of this study will facilitate identification of populations or accessions of wild wheat with favorable traits and/or novel adaptive genes.





Similar content being viewed by others
References
Alonso-Blanco C, Aarts MG, Bentsink L, Keurentjes JJ, Reymond M, Vreugdenhil D, Koornneef M (2009) What has natural variation taught us about plant development, physiology, and adaptation? Plant Cell 21(7):1877–1896
Ankori H, Zohary D (1962) Natural hybridization between Aegilops sharonensis and Ae. longissima: a morphological and cytological study. Cytologia 27(3):314–324
Bouyioukos C, Moscou MJ, Champouret N, Hernández-Pinzón I, Ward ER, Wulff BB (2013) Characterisation and analysis of the Aegilops sharonensis transcriptome, a wild relative of wheat in the Sitopsis Section. PLoS ONE 8(8):e72782
Brody T (1983) Patterns of trait variation in the diploid wheats (Triticum, Aegilops) and the tetraploid species Triticum dicoccoides. Plant Syst Evol 143(4):257–275
Dan J, Koyumdjisky H (1963) The soils of Israel and their distribution. J Soil Sci 14(1):12–20
Danin A (1988) Flora and vegetation of Israel and adjacent areas. In: Yom-Tov Y, Tchernov E (eds) The zoogeography of Israel, Dr. W. Junk Publishers, Dordrecht, pp 129–158
Eig A (1929) Monographisch-kritische Ubersicht der Gattung Aegilops. Rep Spec Nov Regni Veg Beih 55:1–228
Eilam T, Anikster Y, Millet E, Manisterski J, Sagi-Assif O, Feldman M (2007) Genome size and genome evolution in diploid Triticeae species. Genome 50(11):1029–1037
Feldman M, Kislev M (1977) Aegilops searsii, a new species of section Sitopsis (Platystachys). Israel J Bot 26(4):190–201
Furuta Y, Nishikawa K, Yamaguchi S (1986) Nuclear DNA content in diploid wheat and its relatives in relation to the phylogeny of tetraploid wheat. Jpn J Genet 61(2):97–105
Giorgi D, D’Ovidio R, Tanzarella OA, Porceddu E (2002) RFLP analysis of Aegilops species belonging to the Sitopsis section. Genet Resour Crop Evol 49(2):145–151
Goldreich Y (1994) The spatial distribution of annual rainfall in Israel—a review. Theor Appl Climatol 50(1–2):45–59
Goryunova SV, Chikida NN, Kochieva EZ (2008) Molecular analysis of the phylogenetic relationships among the diploid Aegilops species of the section Sitopsis. Russ J Genet 44(1):115–118
Grundbacher FJ (1963) The physiological function of the cereal awn. Bot Rev 29(3):366–381
Hammer K (1980) Zur Taxonomie und Nomenklatur der Gattung Aegilops L. Feddes Rep 91(4):225–258
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25(15):1965–1978
Hijmans RJ, Guarino L, Bussink C, Mathur P, Cruz M, Barrentes I, Rojas E (2012) DIVA-GIS 7.5. A geographic information system for the analysis of species distribution data. http://www.diva-gis.org
Hillel J, Simchen G, Feldman MW (1973) Mating systems and population structure in two closely related species of the wheat group. II. Environmental factors and population structure. Heredity 30:73–83
Johnson RR, Willmer CM, Moss DN (1975) Role of awns in photosynthesis, respiration, and transpiration of barley spikes. Crop Sci 15(2):217–221
Kato K, Miura H, Akiyama M, Kuroshima M, Sawada S (1998a) RFLP mapping of the three major genes, Vrn1, Q and B1, on the long arm of chromosome 5A of wheat. Euphytica 101(1):91–95
Kato K, Tanizoe C, Beiles A, Nevo E (1998b) Geographical variation in heading traits in wild Emmer wheat, Triticum dicoccoides. II. Variation in heading date and adaptation to diverse eco-geographical conditions. Hereditas 128(1):33–39
Kihara H (1954) Considerations on the evolution and distribution of Aegilops species based on the analyser-method. Cytologia 19(4):336–357
Kilian B, Özkan H, Deusch O, Effgen S, Brandolini A, Kohl J, Salamini F (2007) Independent wheat B and G genome origins in outcrossing Aegilops progenitor haplotypes. Mol Biol Evol 24(1):217–227
Kilian B, Mammen K, Millet E, Sharma R, Graner A, Salamini F, Hammer K, Özkan H (2011) Aegilops. In: Wild crop relatives: genomic and breeding resources. Springer Berlin, pp 1–76
Kooyers NJ (2015) The evolution of drought escape and avoidance in natural herbaceous populations. Plant Sci 234:155–162
New M, Lister D, Hulme M, Makin I (2002) A high-resolution data set of surface climate over global land areas. Clim Res 21(1):1–25
Olivera PD, Steffenson BJ (2009) Aegilops sharonensis: origin, genetics, diversity, and potential for wheat improvement. Botany 87(8):740–756
Olivera PD, Anikster Y, Steffenson BJ (2010) Genetic diversity and population structure in Aegilops sharonensis. Crop Sci 50(2):636–648
Rafi MM, Ehdaie B, Waines JG (1992) Quality traits, carbon isotope discrimination and yield components in wild wheats. Ann Bot 69(5):467–474
Roy RP (1959) Genome analysis of Aegilops sharonensis. Genetica 29(1):331–357
Sasanuma T, Miyashita NT, Tsunewaki K (1996) Wheat phylogeny determined by RFLP analysis of nuclear DNA. 3. Intra-and interspecific variations of five Aegilops Sitopsis species. Theor Appl Genet 92(8):928–934
Sourdille P, Cadalen T, Gay G, Gill B, Bernard M (2002) Molecular and physical mapping of genes affecting awning in wheat. Plant Breed 121(4):320–324
Tanaka M (1955) Chromosome pairing in hybrids between Aegilops sharonensis and some species of Aegilops and Triticum. Wheat Inf Serv 2:7–8
Team RC (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna (2012)
Teare ID, Sij JW, Waldren RP, Goltz SM (1972) Comparative data on the rate of photosynthesis, respiration, and transpiration of different organs in awned and awnless isogenic lines of wheat. Can J Plant Sci 52(6):965–971
The International Wheat Genome Sequencing Consortium (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345(6194):1251788
van Slageren MW (1994) Wild wheats: a monograph of Aegilops L. and Amblyopyrum (Jaub. & Spach) Eig (Poaceae). Wageningen Agricultural University Papers 94-7
Volis S (2007) Correlated patterns of variation in phenology and seed production in populations of two annual grasses along an aridity gradient. Evol Ecol 21(3):381–393
Waines JG, Rafi MM, Ehdaie B (1993) Yield components and transpiration efficiency in wild wheats. In: Damania AB (ed) Biodiversity and wheat improvement, Wiley, Chichester, pp 173–186
Yamane K, Kawahara T (2005) Intra and interspecific phylogenetic relationships among diploid Triticum-Aegilops species (Poaceae) based on base-pair substitutions, indels, and microsatellites in chloroplast noncoding sequences. Am J Bot 92(11):1887–1898
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100(10):6263–6268
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 303(5664):1640–1644
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Yasuda S, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci USA 103(51):19581–19586
Acknowledgments
We thank H. Yamaguchi, Y. Matsuoka, Y. Nakayama and K. Tanno for guidance and suggestions. We also thank Y. Yasui and the staff of the Laboratory of Crop Evolution for help with this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ohta, A., Yamane, K. & Kawahara, T. Relationship between spike morphology and habitat of four Aegilops species of section Sitopsis. Genet Resour Crop Evol 64, 889–899 (2017). https://doi.org/10.1007/s10722-016-0408-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-016-0408-x