Abstract
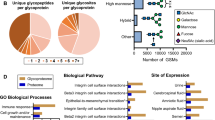
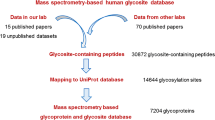
Glycosylation is a very important post-translational modification involved in various cellular processes, such as cell adhesion, signal transduction and immune response. Urine is a rich source of glycoproteins and attractive biological fluid for biomarker discovery, owing to its availability, ease of collection, and correlation with pathophysiology of diseases. Although the urinary proteomics have been explored previously, the urinary glycoproteome characterization remains challenging requiring the development and optimization of analytical and bioinformatics methods for protein glycoprofiling. This study describes the high confident identification of 472 unique N-glycosylation sites covering 256 urinary glycoproteins. Besides, 202 unique N-glycosylation sites were identified in low molecular weight endogenous glycopeptides, which belong to 90 glycoproteins. Global site-specific characterization of the N-linked glycan heterogeneity was achieved by intact glycopeptide analysis, revealing 303 unique glycopeptides most of them displaying complex/hybrid glycans composed by sialic acid and fucose. These datasets consist in a valuable resource of glycoproteins and N-glycosylation sites found in healthy human urine that can be further explored in different disorders, in which the N-linked glycosylation may be aberrant.




Similar content being viewed by others
References
Apweiler R., Hermjakob H., Sharon N.: On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta. 1473(1), 4–8 (1999)
Ferris S.P., Kodali V.K., Kaufman R.J.: Glycoprotein folding and quality-control mechanisms in protein-folding diseases. Dis. Model. Mech. 7(3), 331–341 (2014). doi:10.1242/dmm.014589
Marth J.D., Grewal P.K.: Mammalian glycosylation in immunity. Nat. Rev. Immunol. 8(11), 874–887 (2008). doi:10.1038/nri2417
Cummings R.D., Pierce J.M.: The challenge and promise of glycomics. Chem. Biol. 21(1), 1–15 (2014). doi:10.1016/j.chembiol.2013.12.010
Crocker P.R., Feizi T.: Carbohydrate recognition systems: functional triads in cell-cell interactions. Curr. Opin. Struct. Biol. 6(5), 679–691 (1996)
Pinho S.S., Reis C.A.: Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer. 15(9), 540–555 (2015). doi:10.1038/nrc3982
Fuster M.M., Esko J.D.: The sweet and sour of cancer: glycans as novel therapeutic targets. Nat. Rev. Cancer. 5(7), 526–542 (2005). doi:10.1038/nrc1649
Dube D.H., Bertozzi C.R.: Glycans in cancer and inflammation–potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 4(6), 477–488 (2005). doi:10.1038/nrd1751
Stowell S.R., Ju T., Cummings R.D.: Protein glycosylation in cancer. Annu. Rev. Pathol. 10, 473–510 (2015). doi:10.1146/annurev-pathol-012414-040438
Gilgunn S., Conroy P.J., Saldova R., Rudd P.M., O'Kennedy R.J.: Aberrant PSA glycosylation–a sweet predictor of prostate cancer. Nat. Rev. Urol. 10(2), 99–107 (2013). doi:10.1038/nrurol.2012.258
Hauselmann I., Borsig L.: Altered tumor-cell glycosylation promotes metastasis. Frontiers in oncology. 4, 28 (2014). doi:10.3389/fonc.2014.00028
Zhang Y., Jiao J., Yang P., Lu H.: Mass spectrometry-based N-glycoproteomics for cancer biomarker discovery. Clin. Proteomics. 11(1), 18 (2014). doi:10.1186/1559-0275-11-18
Thaysen-Andersen M., Packer N.H.: Advances in LC-MS/MS-based glycoproteomics: getting closer to system-wide site-specific mapping of the N- and O-glycoproteome. Biochim. Biophys. Acta. 1844(9), 1437–1452 (2014). doi:10.1016/j.bbapap.2014.05.002
Kaji H., Saito H., Yamauchi Y., Shinkawa T., Taoka M., Hirabayashi J., Kasai K., Takahashi N., Isobe T.: Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nat. Biotechnol. 21(6), 667–672 (2003). doi:10.1038/nbt829
Morelle W., Faid V., Chirat F., Michalski J.C.: Analysis of N- and O-linked glycans from glycoproteins using MALDI-TOF mass spectrometry. Methods Mol. Biol. 534, 5–21 (2009). doi:10.1007/978-1-59745-022-5_1
Jensen P.H., Karlsson N.G., Kolarich D., Packer N.H.: Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 7(7), 1299–1310 (2012). doi:10.1038/nprot.2012.063
Woo C.M., Iavarone A.T., Spiciarich D.R., Palaniappan K.K., Bertozzi C.R.: Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat. Methods. 12(6), 561–567 (2015). doi:10.1038/nmeth.3366
Sun S., Shah P., Eshghi S.T., Yang W., Trikannad N., Yang S., Chen L., Aiyetan P., Hoti N., Zhang Z., Chan D.W., Zhang H.: Comprehensive analysis of protein glycosylation by solid-phase extraction of N-linked glycans and glycosite-containing peptides. Nat. Biotechnol. 34(1), 84–88 (2016). doi:10.1038/nbt.3403
Medzihradszky K.F., Kaasik K., Chalkley R.J.: Tissue-specific glycosylation at the glycopeptide level. Mol. Cell. Proteomics: MCP. 14(8), 2103–2110 (2015). doi:10.1074/mcp.M115.050393
Hoffmann M., Marx K., Reichl U., Wuhrer M., Rapp E.: Site-specific O-glycosylation analysis of human blood plasma proteins. Mol. Cell. Proteomics: MCP. 15(2), 624–641 (2016). doi:10.1074/mcp.M115.053546
Parker B.L., Thaysen-Andersen M., Solis N., Scott N.E., Larsen M.R., Graham M.E., Packer N.H., Cordwell S.J.: Site-specific glycan-peptide analysis for determination of N-glycoproteome heterogeneity. J. Proteome Res. 12(12), 5791–5800 (2013). doi:10.1021/pr400783j
Stavenhagen K., Hinneburg H., Thaysen-Andersen M., Hartmann L., Varon Silva D., Fuchser J., Kaspar S., Rapp E., Seeberger P.H., Kolarich D.: Quantitative mapping of glycoprotein micro-heterogeneity and macro-heterogeneity: an evaluation of mass spectrometry signal strengths using synthetic peptides and glycopeptides. J. Mass Spectrom.: JMS. 48(6), 627–639 (2013). doi:10.1002/jms.3210
Hagglund P., Bunkenborg J., Elortza F., Jensen O.N., Roepstorff P.: A new strategy for identification of N-glycosylated proteins and unambiguous assignment of their glycosylation sites using HILIC enrichment and partial deglycosylation. J. Proteome Res. 3(3), 556–566 (2004)
Mysling S., Palmisano G., Hojrup P., Thaysen-Andersen M.: Utilizing ion-pairing hydrophilic interaction chromatography solid phase extraction for efficient glycopeptide enrichment in glycoproteomics. Anal. Chem. 82(13), 5598–5609 (2010). doi:10.1021/ac100530w
Li, X., Jiang, J., Zhao, X., Wang, J., Han, H., Zhao, Y., Peng, B., Zhong, R., Ying, W., Qian, X.: N-glycoproteome analysis of the secretome of human metastatic hepatocellular carcinoma cell lines combining hydrazide chemistry, HILIC enrichment and mass spectrometry. PloS one 8(12), e81921 (2013). doi:10.1371/journal.pone.0081921
Melo-Braga M.N., Schulz M., Liu Q., Swistowski A., Palmisano G., Engholm-Keller K., Jakobsen L., Zeng X., Larsen M.R.: Comprehensive quantitative comparison of the membrane proteome, phosphoproteome, and sialiome of human embryonic and neural stem cells. Mol. Cell. Proteomics: MCP. 13(1), 311–328 (2014). doi:10.1074/mcp.M112.026898
Pompach P., Chandler K.B., Lan R., Edwards N., Goldman R.: Semi-automated identification of N-glycopeptides by hydrophilic interaction chromatography, nano-reverse-phase LC-MS/MS, and glycan database search. J. Proteome Res. 11(3), 1728–1740 (2012). doi:10.1021/pr201183w
Cheng K., Chen R., Seebun D., Ye M., Figeys D., Zou H.: Large-scale characterization of intact N-glycopeptides using an automated glycoproteomic method. J. Proteome. 110, 145–154 (2014). doi:10.1016/j.jprot.2014.08.006
Wuhrer M., de Boer A.R., Deelder A.M.: Structural glycomics using hydrophilic interaction chromatography (HILIC) with mass spectrometry. Mass Spectrom. Rev. 28(2), 192–206 (2009). doi:10.1002/mas.20195
Yu Y.Q., Gilar M., Kaska J., Gebler J.C.: A rapid sample preparation method for mass spectrometric characterization of N-linked glycans. Rapid Commun. Mass Spectrom.: RCM. 19(16), 2331–2336 (2005). doi:10.1002/rcm.2067
Shimwell N.J., Bryan R.T., Wei W., James N.D., Cheng K.K., Zeegers M.P., Johnson P.J., Martin A., Ward D.G.: Combined proteome and transcriptome analyses for the discovery of urinary biomarkers for urothelial carcinoma. Br. J. Cancer. 108(9), 1854–1861 (2013). doi:10.1038/bjc.2013.157
Zhang H., Cao J., Li L., Liu Y., Zhao H., Li N., Li B., Zhang A., Huang H., Chen S., Dong M., Yu L., Zhang J., Chen L.: Identification of urine protein biomarkers with the potential for early detection of lung cancer. Sci. Rep. 5, 11805 (2015). doi:10.1038/srep11805
Wu J., Chen Y.D., Gu W.: Urinary proteomics as a novel tool for biomarker discovery in kidney diseases. J. Zhejiang Univ. Sci. B. 11(4), 227–237 (2010). doi:10.1631/jzus.B0900327
Thomas, C.E., Sexton, W., Benson, K., Sutphen, R., Koomen, J.: Urine collection and processing for protein biomarker discovery and quantification. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 19(4), 953–959 (2010). doi:10.1158/1055-9965.EPI-10-0069
Overbye A., Skotland T., Koehler C.J., Thiede B., Seierstad T., Berge V., Sandvig K., Llorente A.: Identification of prostate cancer biomarkers in urinary exosomes. Oncotarget. 6(30), 30357–30376 (2015). doi:10.18632/oncotarget.4851
Haj-Ahmad T.A., Abdalla M.A., Haj-Ahmad Y.: Potential urinary protein biomarker candidates for the accurate detection of prostate cancer among benign prostatic hyperplasia patients. J. Cancer. 5(2), 103–114 (2014). doi:10.7150/jca.6890
Jedinak A., Curatolo A., Zurakowski D., Dillon S., Bhasin M.K., Libermann T.A., Roy R., Sachdev M., Loughlin K.R., Moses M.A.: Novel non-invasive biomarkers that distinguish between benign prostate hyperplasia and prostate cancer. BMC Cancer. 15, 259 (2015). doi:10.1186/s12885-015-1284-z
Wang L., Li F., Sun W., Wu S., Wang X., Zhang L., Zheng D., Wang J., Gao Y.: Concanavalin A-captured glycoproteins in healthy human urine. Mol. Cell. Proteomics: MCP. 5(3), 560–562 (2006). doi:10.1074/mcp.D500013-MCP200
Yang N., Feng S., Shedden K., Xie X., Liu Y., Rosser C.J., Lubman D.M., Goodison S.: Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification. Clinical cancer research: an official journal of the American Association for Cancer Research. 17(10), 3349–3359 (2011). doi:10.1158/1078-0432.CCR-10-3121
Saraswat M., Joenvaara S., Musante L., Peltoniemi H., Holthofer H., Renkonen R.: N-linked (N-) glycoproteomics of urinary exosomes. [Corrected]. Mol. Cell. Proteomics: MCP. 14(2), 263–276 (2015). doi:10.1074/mcp.M114.040345
Halim A., Nilsson J., Ruetschi U., Hesse C., Larson G.: : Human urinary glycoproteomics; attachment site specific analysis of N- and O-linked glycosylations by CID and ECD. Mol. Cell. Proteomics: MCP. 11(4), M111 013649 (2012). doi:10.1074/mcp.M111.013649
Bern, M., Kil, Y.J., Becker, C.: Byonic: advanced peptide and protein identification software. Current protocols in bioinformatics / editoral board, Andreas D. Baxevanis. .. [et al.] Chapter 13, Unit13 20 (2012). doi:10.1002/0471250953.bi1320s40
Bern M., Cai Y., Goldberg D.: Lookup peaks: a hybrid of de novo sequencing and database search for protein identification by tandem mass spectrometry. Anal. Chem. 79(4), 1393–1400 (2007). doi:10.1021/ac0617013
Bern M.W., Kil Y.J.: Two-dimensional target decoy strategy for shotgun proteomics. J. Proteome Res. 10(12), 5296–5301 (2011). doi:10.1021/pr200780j
Cox J., Mann M.: MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26(12), 1367–1372 (2008). doi:10.1038/nbt.1511
Cox J., Neuhauser N., Michalski A., Scheltema R.A., Olsen J.V., Mann M.: Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10(4), 1794–1805 (2011). doi:10.1021/pr101065j
Palmisano G., Melo-Braga M.N., Engholm-Keller K., Parker B.L., Larsen M.R.: Chemical deamidation: a common pitfall in large-scale N-linked glycoproteomic mass spectrometry-based analyses. J. Proteome Res. 11(3), 1949–1957 (2012). doi:10.1021/pr2011268
Breuer K., Foroushani A.K., Laird M.R., Chen C., Sribnaia A., Lo R., Winsor G.L., Hancock R.E., Brinkman F.S., Lynn D.J.: InnateDB: systems biology of innate immunity and beyond–recent updates and continuing curation. Nucleic Acids Res. 41(Database issue), D1228–D1233 (2013). doi:10.1093/nar/gks1147
Chen E.Y., Tan C.M., Kou Y., Duan Q., Wang Z., Meirelles G.V., Clark N.R., Ma'ayan A.: Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013). doi:10.1186/1471-2105-14-128
Su A.I., Wiltshire T., Batalov S., Lapp H., Ching K.A., Block D., Zhang J., Soden R., Hayakawa M., Kreiman G., Cooke M.P., Walker J.R., Hogenesch J.B.: A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. U. S. A. 101(16), 6062–6067 (2004). doi:10.1073/pnas.0400782101
Kim M.S., Pinto S.M., Getnet D., Nirujogi R.S., Manda S.S., Chaerkady R., Madugundu A.K., Kelkar D.S., Isserlin R., Jain S., Thomas J.K., Muthusamy B., Leal-Rojas P., Kumar P., Sahasrabuddhe N.A., Balakrishnan L., Advani J., George B., Renuse S., Selvan L.D., Patil A.H., Nanjappa V., Radhakrishnan A., Prasad S., Subbannayya T., Raju R., Kumar M., Sreenivasamurthy S.K., Marimuthu A., Sathe G.J., Chavan S., Datta K.K., Subbannayya Y., Sahu A., Yelamanchi S.D., Jayaram S., Rajagopalan P., Sharma J., Murthy K.R., Syed N., Goel R., Khan A.A., Ahmad S., Dey G., Mudgal K., Chatterjee A., Huang T.C., Zhong J., Wu X., Shaw P.G., Freed D., Zahari M.S., Mukherjee K.K., Shankar S., Mahadevan A., Lam H., Mitchell C.J., Shankar S.K., Satishchandra P., Schroeder J.T., Sirdeshmukh R., Maitra A., Leach S.D., Drake C.G., Halushka M.K., Prasad T.S., Hruban R.H., Kerr C.L., Bader G.D., Iacobuzio-Donahue C.A., Gowda H., Pandey A.: A draft map of the human proteome. Nature. 509(7502), 575–581 (2014). doi:10.1038/nature13302
Shah P., Wang X., Yang W., Toghi Eshghi S., Sun S., Hoti N., Chen L., Yang S., Pasay J., Rubin A., Zhang H.: Integrated proteomic and glycoproteomic analyses of prostate cancer cells reveal glycoprotein alteration in protein abundance and glycosylation. Mol. Cell. Proteomics: MCP. 14(10), 2753–2763 (2015). doi:10.1074/mcp.M115.047928
Gao W.N., Yau L.F., Liu L., Zeng X., Chen D.C., Jiang M., Liu J., Wang J.R., Jiang Z.H.: Microfluidic Chip-LC/MS-based Glycomic analysis revealed distinct N-glycan profile of rat serum. Sci. Rep. 5, 12844 (2015). doi:10.1038/srep12844
Campbell M.P., Peterson R., Mariethoz J., Gasteiger E., Akune Y., Aoki-Kinoshita K.F., Lisacek F., Packer N.H.: UniCarbKB: building a knowledge platform for glycoproteomics. Nucleic Acids Res. 42(Database issue), D215–D221 (2014). doi:10.1093/nar/gkt1128
Eckhard U., Huesgen P.F., Schilling O., Bellac C.L., Butler G.S., Cox J.H., Dufour A., Goebeler V., Kappelhoff R., Keller U.A., Klein T., Lange P.F., Marino G., Morrison C.J., Prudova A., Rodriguez D., Starr A.E., Wang Y., Overall C.M.: Active site specificity profiling of the matrix metalloproteinase family: Proteomic identification of 4300 cleavage sites by nine MMPs explored with structural and synthetic peptide cleavage analyses. Matrix Biol.: journal of the International Society for Matrix Biology. 49, 37–60 (2016). doi:10.1016/j.matbio.2015.09.003
Tucher J., Linke D., Koudelka T., Cassidy L., Tredup C., Wichert R., Pietrzik C., Becker-Pauly C., Tholey A.: LC-MS based cleavage site profiling of the proteases ADAM10 and ADAM17 using proteome-derived peptide libraries. J. Proteome Res. 13(4), 2205–2214 (2014). doi:10.1021/pr401135u
Prudova A., auf dem Keller U., Butler G.S., Overall C.M.: Multiplex N-terminome analysis of MMP-2 and MMP-9 substrate degradomes by iTRAQ-TAILS quantitative proteomics. Mol. Cell. Proteomics: MCP. 9(5), 894–911 (2010). doi:10.1074/mcp.M000050-MCP201
Schilling O., Overall C.M.: Proteome-derived, database-searchable peptide libraries for identifying protease cleavage sites. Nat. Biotechnol. 26(6), 685–694 (2008). doi:10.1038/nbt1408
Acknowledgments
CNPq (GP 441878/2014-8) and FAPESP (GP: 2014/06863-3), Rebeca Kawahara is supported the Capes, PNPD, and FAPESP (2015/02866-0). Joyce Saad is supported by the “Programa Unificado de Bolsas de Estudo”. The facility Biomass at CEFAP-USP is acknowledged for the analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
Supplemental Figure 1
Tissue localization enrichment analysis in the list of identified urinary glycoproteins (PDF 36 kb)
Supplemental Figure 2
Manual verification of the Y1 + 16 Da in the MS/MS spectra from intact glycopeptides identified with NeuGc. The Y1 ion +16 Da is indicated by the red arrow. (PDF 248 kb)
Supplemental Figure 3
Charge state distribution of intact glycopeptides identified in each glycan composition class. The percentage was calculated based on the number of glycopeptides with charge +2, +3, +4 and >4 in each class to the total number of glycopeptides identified in each class. (PDF 15 kb)
Supplemental Figure 4
Overlap between the number of peptide sequences identified in the formerly N-linked peptides generated after PNGase F treatment and intact glycopeptides (with 0 % FDR) identified using Byonic software. (PDF 12 kb)
Supplemental Figure 5
Examples of annotated spectra for N-linked glycopeptides identified in urinary glycoproteins (PDF 62 kb)
Supplemental Figure 6
Sequence recognition motif in endogenous N-glycopeptides. N-glycosylation consensus sequence as derived using MotifX. A) MotifX analysis centered on the NxT glycosylation motif. B) MotifX analysis centered on the NxS glycosylation motif. C) Sequence2logo analysis obtained using the endogenous peptide sequences with six aminoacids length. (PDF 93 kb)
ESM 7
(XLSX 1076 kb)
Rights and permissions
About this article
Cite this article
Kawahara, R., Saad, J., Angeli, C.B. et al. Site-specific characterization of N-linked glycosylation in human urinary glycoproteins and endogenous glycopeptides. Glycoconj J 33, 937–951 (2016). https://doi.org/10.1007/s10719-016-9677-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-016-9677-z