Abstract
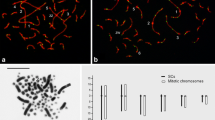
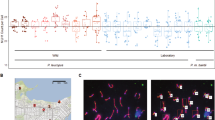
Meiotic recombination in female ducks was directly studied by immunolocalization of MLH1 protein, a mismatch repair protein of mature recombination nodules. In total, 6820 crossovers were scored along the autosomal synaptonemal complexes in 122 meiotic nuclei. From this analysis we predict that the female map length of the duck is 2845 cM, with a genome wide recombination rate of 2 cM/Mb. MLH1-focus mapping along the six largest bivalents shows regional variations of recombination frequencies that can be linked to differences in chromosome morphology. From this MLH1 mapping it can be inferred that distally located markers will appear more separated in genetic maps than physically equidistant markers located near the centromeres on bivalents 1 and 2. Instead, markers at interstitial positions on the acrocentric bivalents 3–6 will appear more tightly linked than expected on the basis of their physical distance because recombination is comparatively lower at the mid region of these chromosomes. The present results provide useful information to complement linkage mapping in ducks and extend previous knowledge about the variation of recombination rates among domestic Galloanserae.



Similar content being viewed by others
References
Anderson LK et al (2014) Combined fluorescent and electron microscopic imaging unveils the specific properties of two classes of meiotic crossovers. Proc Natl Acad Sci USA 111:13415–13420. doi:10.1073/pnas.1406846111
Backström N et al (2010) The recombination landscape of the zebra finch Taeniopygia guttata genome. Genome Res. doi:10.1101/gr.101410.109
Borodin PM, Karamysheva TV, Belonogova NM, Torgasheva AA, Rubtsov NB, Searle JB (2008) Recombination map of the common shrew, Sorex araneus (Eulipotyphla, Mammalia). Genetics 178:621–632
Borodin PM, Basheva EA, Zhelezova AI (2009) Immunocytological analysis of meiotic recombination in the American mink (Mustela vison). Anim Genet 40:235–238. doi:10.1111/j.1365-2052.2008.01808.x
Calderón PL, Pigozzi MI (2006) MLH1-focus mapping in birds shows equal recombination between sexes and diversity of crossover patterns. Chromosome Res 14:605–612
del Priore L, Pigozzi MI (2012) Chromosomal axis formation and meiotic progression in chicken oocytes: a quantitative analysis. Cytogenet Genome Res 137:15–21. doi:10.1159/000339133
del Priore L, Pigozzi MI (2015) Sex-specific recombination maps for individual macrochromosomes in the Japanese quail (Coturnix japonica). Chromosome Res 23:199–210. doi:10.1007/s10577-014-9448-2
Dolezel J, Bartos J, Voglmayr H, Greilhuber J (2003) Nuclear DNA content and genome size of trout and human. Cytom A 51:127–128. doi:10.1002/cyto.a.10013 (author reply 129)
Fillon V, Vignoles M, Crooijmans RP, Groenen MA, Zoorob R, Vignal A (2007) FISH mapping of 57 BAC clones reveals strong conservation of synteny between Galliformes and Anseriformes. Anim Genet 38:303–307. doi:10.1111/j.1365-2052.2007.01578.x
Froenicke L, Anderson LK, Wienberg J, Ashley T (2002) Male mouse recombination maps for each autosome identified by chromosome painting. Am J Hum Genet 71:1353–1368
Frohlich J, Vozdova M, Kubickova S, Cernohorska H, Sebestova H, Rubes J (2015) Variation of meiotic recombination rates and MLH1 foci distribution in spermatocytes of cattle, sheep and goats. Cytogenet Genome Res. doi:10.1159/000439452
Gregory TR (2016) Animal Genome Size Database. http://www.genomesize.com
Groenen MA et al (2009) A high-density SNP-based linkage map of the chicken genome reveals sequence features correlated with recombination rate. Genome Res 19:510–519. doi:10.1101/gr.086538.108
Huang Y, Zhao Y, Haley CS, Hu S, Hao J, Wu C, Li N (2006) A genetic and cytogenetic map for the duck (Anas platyrhynchos). Genetics 173:287–296. doi:10.1534/genetics.105.053256
Huang CW, Cheng YS, Rouvier R, Yang KT, Wu CP, Huang HL, Huang MC (2009) Duck (Anas platyrhynchos) linkage mapping by AFLP fingerprinting. Genet Sel Evol 41:28. doi:10.1186/1297-9686-41-28
Huang Y et al (2013) The duck genome and transcriptome provide insight into an avian influenza virus reservoir species. Nat Genet 45:776–783. doi:10.1038/ng.2657
Hultén M, Tease C (2006) Genetic mapping: comparison of direct and indirect approaches. Encycl Life Sci. doi:10.1002/9780470015902.a0006268
Kocsis E, Trus BL, Steer CJ, Bisher ME, Steven AC (1991) Image averaging of flexible fibrous macromolecules: the clathrin triskelion has an elastic proximal segment. J Struct Biol 107:6–14
Lisachov AP, Zadesenets KS, Rubtsov NB, Borodin PM (2015) Sex chromosome synapsis and recombination in male guppies. Zebrafish 12:174–180. doi:10.1089/zeb.2014.1000
Mary N et al (2014) Meiotic recombination analyses of individual chromosomes in male domestic pigs (Sus scrofa domestica). PLoS ONE 9:e99123. doi:10.1371/journal.pone.0099123
Mezard C, Jahns MT, Grelon M (2015) Where to cross? New insights into the location of meiotic crossovers. Trends Genet 31:393–401. doi:10.1016/j.tig.2015.03.008
Pigozzi MI (2001) Distribution of MLH1 foci on the synaptonemal complexes of chicken oocytes. Cytogenet Cell Genet 95:129–133
Pigozzi MI, Solari AJ (1999) Recombination nodule mapping and chiasma distribution in spermatocytes of the pigeon, Columba livia. Genome 42:308–314
Rahn MI, Solari AJ (1986) Recombination nodules in the oocytes of the chicken, Gallus domesticus. Cytogenet Cell Genet 43:187–193
Rodionov AV, Chechik MS (2002) Lampbrush chromosomes in the Japanese quail Coturnix coturnix japonica: cytological maps of macro chromosomes and meiotic crossover frequency in females. Genetika 38:1246–1251
Rodionov AV, Lukina NA, Galkina SA, Solovei I, Saccone S (2002) Crossing over in chicken oogenesis: cytological and chiasma-based genetic maps of the chicken lampbrush chromosome 1. J Hered 93:125–129
Sherman JD, Stack SM (1995) Two-dimensional spreads of synaptonemal complexes from solanaceous plants. VI. High-resolution recombination nodule map for tomato (Lycopersicon esculentum). Genetics 141:683–708
Skinner BM et al (2009) Comparative genomics in chicken and Pekin duck using FISH mapping and microarray analysis. BMC Genom 10:357. doi:10.1186/1471-2164-10-357
Solari AJ, Pigozzi MI (1993) Recombination nodules and axial equalization in the ZW pairs of the Peking duck and the guinea fowl. Cytogenet Cell Genet 64:268–272
Stapley J, Birkhead TR, Burke T, Slate J (2008) A linkage map of the zebra finch Taeniopygia guttata provides new insights into avian genome evolution. Genetics 179:651–667. doi:10.1534/genetics.107.086264
Sun F et al (2006) Variation in MLH1 distribution in recombination maps for individual chromosomes from human males. Hum Mol Genet 15:2376–2391. doi:10.1093/hmg/ddl162
Vozdova M, Sebestova H, Kubícková S, Cernohorska H, Vahala J, Rubes J (2013) A comparative study of meiotic recombination in cattle (Bos taurus) and three wildebeest species (Connochaetes gnou, C. taurinus taurinus and C. t. albojubatus). Cytogenet Genome Res 140:36–45. doi:10.1159/000350444
Zhang L, Liang Z, Hutchinson J, Kleckner N (2014) Crossover patterning by the beam-film model: analysis and implications. PLoS Genet 10:e1004042. doi:10.1371/journal.pgen.1004042
Acknowledgments
Research in our laboratory is funded by grants PIP 003 from Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET) and PICT 0059 from Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT). Support from the University of Buenos Aires is gratefully acknowledged. M.I.P. is a member of CONICET and L.d.P. is the recipient of a doctoral fellowship from CONICET.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s10709-016-9922-1.
Rights and permissions
About this article
Cite this article
Pigozzi, M.I., del Priore, L. Meiotic recombination analysis in female ducks (Anas platyrhynchos). Genetica 144, 307–312 (2016). https://doi.org/10.1007/s10709-016-9899-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-016-9899-9