Abstract
In the twenty-first century, the first pandemic novel human influenza A/H1N1virus (NIV) outbreak was reported at Mexico and USA on March and early April, 2009 respectively. The outbreak occurred among human populations due to the presence of meager or no immune response against newly emerged viruses. The success of vaccines and drugs depends on their low susceptibility to the formation of escape mutants in virus. Identification of excess, non-synonymous substitutions over synonymous ones is a main indicator of positive Darwinian selection in protein-coding genes of NIVs. The positive Darwinian selection operating on each site of proteins were inferred by computing ω, the ratio of the non-synonymous/synonymous substitutions [dN/dS (or) Ka/Ks], which was calculated by three different methods in terms of codon-based maximum likelihood, branch-site and empirical Bayesian methods under various models. Totally, nine sites from PB2, PB1, HA, M2 and NS1 are inferred as positively selected. The function for amino acid sites of NIVs proteins under positive selection are inferred by comparing the sites with experimentally determined functionally known amino acid sites. Completely 4 positively selected sites of PB1, HA and M2 are found to be involved in B-cell epitopes (BCEs). Interestingly, most of these sites are also involving in T-cell epitopes (TCEs). However, more sites under positive selection forces are involved in TCEs than those of BCEs. Amino acid sites engaged in both BCEs and TCEs should be measured as highly suitable targets, because these sites could induce the strong humoral and cellular immune responses against targets.
Similar content being viewed by others
Introduction
Humans were victims of several spells of viral outbreaks causing flu in the course of the twentieth century. The ‘Spanish flu’ caused by the H1N1 virus killed 25–50 million people worldwide in 1918, ‘Asian flu’ by the H2N2 virus killed 1–4 million people in 1957 and ‘Hong Kong flu’ by the H3N2 virus killed 0.75–2 million people in 1968. The subtypes H1N1 and H3N2 influenza viruses still continue to circulate and may cause annual epidemics that kill 0.25–0.5 million people worldwide (Suzuki 2006). A report states that there were the extensive influenza outbreaks in 1173, 1510, 1580, 1729, 1781 and 1830. However the term pandemics influenza was used to express these outbreaks since eighteenth century. Before this, in the fourteenth century, it was described as epidemics. These previous outbreaks created the awareness to the people (Potter 2001); as a result, a human infection with influenza A virus became a nationally notifiable disease in the United States (Smith et al. 2009) and worldwide, since 2007. Due to the influenza, each year in the USA, more than 200,000 patients are admitted in hospitals and approximately 36,000 influenza related deaths occurred (http://www.ncbi.nlm.nih.gov/genomes/FLU/flubiology.html).
Influenza infection in humans is characterized by the progressive infection of lung tissue, but not the simultaneous infection of the entire lung. Seasonal influenza infection never progress to complete infection of the human lung (Lazrak et al. 2009), but in the case of pandemic influenza infection, the entire lung is affected. Efficient human-to-human transmission is a prerequisite for any influenza A virus to become pandemic. Furthermore subtype H1N1 viruses are known to cause the disease less frequently than those of subtype H3N2 viruses (Ghedin et al. 2005; Zamarin et al. 2006; Obenauer et al. 2006). In March and early April 2009, novel human influenza A/H1N1 viruses (from here, I refer this virus as ‘NIVs’) emerged in Mexico and in the United States, respectively (CDC 2009) and caused worldwide pandemic outbreaks. Totally, from April to June 2009, NIVs have been widely spread over 170 countries by human-to-human transmissions, causing the World Health Organization to raise its pandemic alert level from 5 to 6 (Smith et al. 2009; Qu et al. 2011; Arunachalam et al. 2012a). The reasons for this pandemic outbreak among humans are (1) meager or no immune response, (2) lack of previous infection and (3) constant changes in the viral genome (Webby and Webster 2003).
Identifying the origin of NIVs might facilitate to know about how NIVs adapt in humans and how to control the spread of NIVs (Arunachalam et al. 2012a). Today vaccines and drugs exist for the prophylaxis and treatment of influenza virus infections. Vaccines are composed of either inactivated or live attenuated virions of subtypes H1N1 and H3N2 human influenza A viruses as well as those of influenza B viruses. Vaccines fail to protect the humans from infections while antigenicities of the wild viruses evolve continuously (Mostow et al. 1970). As a result, vaccines need to be yearly reformulated with updated seed strains. In addition, even when vaccine and wild viruses are similar, escape mutants are often generated (Suzuki 2006; Jin et al. 2005; Zharikova et al. 2005; Venkatramani et al. 2006). Recent epidemiological studies show that seasonal trivalent inactivated influenza vaccine could provide partial protection against NIVs (Garcia-Garcia et al. 2009; Echevarria-Zuno et al. 2010). However, it does not stimulate any immune responses against NIVs (Tu et al. 2010). Escape mutants are often generated for drugs like amantadine (Webster et al. 1986) and less frequently for drugs like oseltamivir (Kiso et al. 2004). In order to develop more effective vaccines and drugs that are less susceptible to the generation of escape mutants, it is important to understand the selective pressure acting on each site of the proteins encoded by NIV genome. The objective of this investigation is to determine the sites in the proteins of NIVs genome that are under the positive selection pressure, which will ultimately be important in the effective design of vaccines and drugs.
Materials and methods
Sequence selection
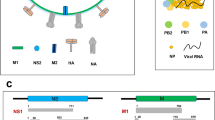
Influenza viruses are the etiological agents of ‘flu’ (Suzuki 2006; Smith et al. 1933) and are classified into types A–C, among which, type A viruses are the most pathogenic to humans (Suzuki and Nei 2002). On the basis of the antigenic properties of hemagglutinin (HA) and neuraminidase (NA), influenza A viruses are classified into subtypes H1–H16 and N1–N9, respectively (Arunachalam et al. 2012a; WHO 1980). Influenza A viruses possesses a single stranded, negative sense and 8 segmented (segments 1–8) RNA genome in an enveloped virion (Noda et al. 2006). In order to analyze all the proteins of the NIVs, the entire protein-coding regions were extracted from the NCBI Influenza Virus Resource (Bao et al. 2008). Among the eight gene segments, the five segments PB2, PA, HA, NP and NA encode a single protein whereas rest of the three segments PB1, MP and NS encode two proteins, namely, PB1 and PB1–F2, M1 and M2 and NS1 and NS2, respectively. Nucleotide positions 95–367 of PB1 and the entire region (positions 1–273) of PB1–F2, positions 1–27 of both M1 and M2 and positions 1–31 of both NS1 and NS2 are overlapped in different reading frames. Prior to selecting the sequences, the purge was made on the sequences with unclear, derived from the same strains as others, laboratory and vaccine strains, shorter sequences and sequences with redundant stop codons. In that case, 99, 95, 97, 94, 96, 97, 58, 22, 75 and 32 sequences that differed from each other were finally selected for positive Darwinian selection analysis for PB2, PB1, PA, HA, NP, NA, M1, M2, NS1 and NS2, correspondingly. The strains names and their databank accession numbers of sequences used in the present study are listed in the ST1. Due to the presence of premature stop codons in PB1–F2 at various sites such as 12, 58, 88 and 91; it has not been included for analysis.
Estimation of dN–dS
For each protein, a multiple alignment of the nucleotide sequences was performed using MUSCLE (Edgar 2004) under neighbor joining (NJ) cluster method implemented in MEGA5 (Tamura et al. 2011). Estimation of selection at each amino acid site of the proteins encoded by NIVs genomes was inferred using MEGA5 via HyPhy (Pond et al. 2005). The maximum likelihood (ML) statistical method (Dempster et al. 1977) was used under Tamura–Nei model (Tamura and Nei 1993) with syn–nonsynonymous substitution type for computing dS and dN values. The gaps and/or missing data treatment was done under ‘Use all sites’ option. Each time of this analysis, the results of HyPhy were automatically exported directly to MEGA5, which could be used to generate sequence-wide profiles for further analyses (Tamura et al. 2011). For estimating ML values, a tree topology was automatically computed. Here, the estimation of site-specific evolutionary rate may depend on the evolutionary tree used (Mayrose et al. 2005). The test statistic dN–dS was used to detect the codons that have undergone positive selection, where dS was the number of synonymous substitutions/site (s/S) and dN was the number of non-synonymous substitutions per site (n/N). Commonly, a species adapting into a new environments, changes in the synonymous base substitutions occur almost always at a much higher rate than those of non-synonymous substitutions (Kimura 1968; Duret 2008). A positive value for the test statistic indicates that an excess of non-synonymous substitutions. In this case, the probability of rejecting the null hypothesis of neutral evolution (p value) was calculated (Suzuki and Gojobori 1999; Pond and Frost 2005). The value of p < 0.05 was considered as statistically significant at a 5 % level (data not shown). Normalized dN–dS for the test statistic were also obtained using the total number of substitutions in the tree (measured in expected substitutions per site). It is useful for making comparisons across the data sets.
Positive Darwinian selection analysis
Currently, availability of large number of genome, gene and protein sequences of different organisms in the biological databases, initiated the development of numerous statistical methods to analyze these sequences. As a result, the rapid improvement in use of phylogenetics and molecular selection programmes (Arunachalam et al. 2012a; Dixit et al. 2010; Blanquer and Uriz 2007; Malickbasha et al. 2010; Arunachalam et al. 2012b; Arunachalam et al. 2012c). Thus, the type of molecular evolution operating on individual codon sites of NIVs were detected by computing ‘ω’, the ratio between non-synonymous (Ka or dN) and synonymous (Ks or dS) substitutions (Stern et al. 2007). The ratio (ω = dN/dS or Ka/Ks) has a straightforward measurement of selective pressure at each codon of protein-coding genes that ‘ω’ values of >1, 1 and <1 indicates positive/diversifying selection, random drift/neutral evolution and negative/purifying selection, respectively (Yang and Bielawski 2000; Chen and Sun 2011). I used both Datamonkey (a server running by HyPhy) (Delport et al. 2010) and Selecton programme, in which the codon-based maximum likelihood (CML), branch-site (BS) and empirical Bayesian evolutionary methods under different substitution models were used for selection analysis. Interestingly, the Datamonkey and Selecton were automatically translating the codons into amino acid sequences whenever the analysis was begin. In the beginning, gene sequences of NIVs were used as input in Datamonkey, subsequently this server was estimated the ‘ω’ values for each site that could be used to infer whether the amino acid sites under positive Darwinian or purifying selection. The selection analyses were made with state-of-the-art statistical methods include four independent CML methods namely single-likelihood ancestor counting (SLAC), fixed effects likelihood (FEL), random effects likelihood (REL) (Pond and Frost 2005), and internal fixed effects likelihood (IFEL) (Pond et al. 2006), and one BS evolutionary method namely mixed effects model of episodic selection (MEME) (Pond et al. 2011). The model selection test was carried out for each method for each gene and the best model was found (Table 1) among the 203 nucleotide substitution models. During the SLAC, FEL, MEME and IFEL analyses, the differential test significance levels were given as 0.1, whereas for REL, the empirical Bayes factors (BF) were given as 50. The ‘ω’ ratio of amino acid sites either with p values ≤0.05 or BF ≥ 20, were considered as statistically significant. The statistically reliable p values for the above mentioned four tests indicate that the presence of strong linear correlation between the approximate and the maximum likelihood parameter estimates (Pond and Frost 2005).
As like Datamonkey, the Selecton 2.4 (Stern et al. 2007) was also run with NIVs gene sequences to calculate the ‘ω’ ratio of each amino acid site using empirical Bayesian method (Yang et al. 2000; Mayrose et al. 2004) under three different evolutionary models namely mechanistic empirical combination (MEC) (Doron-Faigenboim and Pupko 2006) [under JTT empirical amino acid matrix (Jones et al. 1992)], M8 (β + w = 1) and M5 (γ) (Yang et al. 2000). For models MEC, M8 and M5, the number of categories was set as 8, 8 and 14, respectively. I did compare the models MEC and M8 (where the hypothesis allows positive selection operating on the proteins) with null model M8a (β + w = 1) (Swanson et al. 2003) (where hypothesis does not allows positive selection). The lower AICc score indicates that the better fit of the model to the data, and hence the model was considered as more justified. The statistical reliability of the ‘ω’ value was estimated by measuring the confidence of the inference, for instance, for positively selected site (ω > 1), if the corresponding lower bound of the confidence interval was >1, then the assumption of positive selection at this site was considered as reliable. The result of ‘ω’ inferences is projected on the representative primary sequence of each protein of NIVs [PB2, PB1, PA, HA, NP, NA, M1, NS1, NS2: A/Guam/NHRC0002/2009(H1N1)/2009; M2: A/Guam/NHRC0001/2009(H1N1)/2009] (SF. 3). The different types of selection are represented with seven colour scales. The yellow shade’s (colors 1 and 2) indicate that the amino acid sites with ω > 1. However, only sites (ω > 1) with the confidence interval (lower bound) larger than 1 at 95 % level are considered as statistically significant that are colored in dark yellow, whereas the sites with light-yellow color indicates that are not significant. Besides, the shades from white to magenta (colors 3–7) indicate that the different levels of ω ≤ 1 (Stern et al. 2007).
The functions for the amino acid sites under strong positive selection pressure were identified by interpreting the sites with functionally known amino acid sites involved in immune epitopes, antiviral resistance and growth in eggs. Experimentally verified amino acid sites involved in immune epitopes were composed from the immune epitope database (IEDB) and analysis resource (Vita et al. 2010).
Results and discussion
Inference of site-specific positive Darwinian selection
The results on selection pressure of individual amino acid sites of all the proteins encoded by the NIVs genome are shown in Table 1 and SF1, 2, 3. An excess of non-synonymous substitutions over synonymous ones is an important indicator of positive selection at molecular level. For PB2, 198 out of 759 codon sites are variable (26.1 %). Among the 198 variable codon sites, the overabundance of non-synonymous (dN) substitutions (positively selected values) are observed in minority of sites (53 sites; 26.8 %) and the excess of synonymous (dS) substitutions (negatively selected values) are observed in majority of sites (145 sites; 73.2 %) (Table 1; SF1, 2). In this case, the probability of rejecting the null hypothesis of neutral evolution is statistically significant at a 5 % level (data not shown). The selection profiles for all the other proteins are similar to that for PB2, with the exception of M2. In M2 gene, among 19 variable codon sites, the excess of synonymous substitutions are observed in minority of sites (8 sites; 42.1 %) and the excess of non-synonymous substitutions are observed in majority of sites (11 sites; 57.9 %). Our results indicate that the speed of accumulations of synonymous substitution rate is higher than those of non-synonymous substitutions rate in all the genes/proteins, except in M2. It suggests that the changes in the amino acid sequences (non-synonymous) are most probably to reduce the functionality of proteins than to increase it. As a result, they are likely to lower the fitness of NIVs, and they could have a lower probability of being fixed than those changes, which do not change the amino acid sequences (synonymous). These findings are in agreement with the hypothesis suggested by Kimura (1968) and Duret (2008) who proposed that when the organisms are adapting into a new environment, changes in the synonymous base substitutions occur almost always at a much higher rate than those of non-synonymous substitutions. Positively (value) selected non-synonymous substitutions might, however, be involved in the NIVs adaptation in humans (Pond et al. 2006).
Moreover, the force of natural selection in each amino acid site of NIVs proteins was inferred by calculating ‘ω’ that had accumulated over the phylogenetic tree (data not shown), because the direction and magnitude of natural selection may vary during evolution. The phylogenetic trees were inferred using the NJ method (Pond et al. 2006; Mullick et al. 2011). It is noted that the REL analysis in Datamonkey could not be performed due to the alignment size restriction for PB2, PB1, PA, HA, NP and NA genes. Totally, 71 amino acid sites in the entire proteins of NIVs that underwent positive Darwinian selection (ω > 1) are identified. Among the 71 positively selected sites, only 12 sites (highlighted with ‘^’) are identified with statistical significance level as p ≤ 0.05 for SLAC, FEL, IFEL, MEME or as BF ≥ 20 for REL (Pond et al. 2006), when both CML and BS methods are used under best fit model in Datamonkey (Tables 1, 3). In the case of positive Darwinian selection, the significance of one method alone is not sufficient to infer that the amino acid site is subjected to positive selection pressure. Hence, only sites that have been detected by more than one method, are considered as positively selected (Pond et al. 2006; Delport et al. 2010). Accordingly, absolutely 9 (highlighted with ‘^^’) out of 12 statistically reliable amino acid sites are considered as positively selected that are, position 194 (p value 0.05, method—MEME; IFEL; MEC) of PB2, positions 587 (0.03, IFEL; MEC) and 736 (0.004, MEME; 0.01, IFEL) of PB1, positions 7 (0.05, MEME; MEC; M5), 49 (0.03, IFEL; MEC, M5) and 391 (0.004, IFEL; 0.03, FEL and MEME; M5) of HA, position 10 (0.03, IFEL; M5) of M2 and positions 108 (BF 60.954, REL; M5) and 123 (64.414, REL; M5) of NS1 (Table 1; SF3). Furthermore, for positions 108 and 123 of NS1, in addition to REL and M5, model MEC also has supported that these sites are positively selected. However, model MEC is rejected by null model M8a in likelihood ratio test (LRT) analysis (discussed in the next paragraph). Moreover, for the reliable position 588 (0.04, IFEL) of PB2 and position 321 (0.004, MEME) of PA, no parallel positively selected sites are identified by other methods, whereas for position 30 (BF > 1,000; REL) of M1, the parallel positively selected sites are observed in MEC and M8 models, however, these two models are rejected by null model M8a during LRT analysis (Table 2; SF3). It should be noted that no sites are inferred as positively selected for NP, whereas among 6 and 2 positively selected sites in NA and NS2 respectively, no statistically reliable sites are inferred (Table 1). The presences of (highly variable) reliable positively selected sites in NIVs proteins indicate that these sites have undergone the amino acid fixations during the way of NIVs evolution. These positively selected amino acid sites might be interpreted as being an effect of molecular adaptation, which confers an evolutionary advantage to the NIVs. Due to increased adaptation to a new host, the selection pressure could always be high, particularly in the early stage of viral pandemics (Furuse et al. 2010). It is important to note that the sites under positive selection, specifically in PB1, M1, M2, NS1 and NS2 may remain tentative, because, the non-overlapping regions in reading frame could only be considered for the analyses.
For selection analysis, initially, the Selecton programme was allowed to perform a model tests for each protein with different evolutionary models that assume different biological assumptions. Use of different evolutionary models could permit contrasting the different hypotheses by comparing the likelihood ratio of a model, which assumes positive selection (MEC and M8; except M5), to a model, which does not allow positive selection (M8a). The LRT was performed to identify, which model fits better to the protein and, thus, the model that was considered as more reasonable. Results of LRT (presence of lower AIC scores) indicate that MEC model fits better to PB2, PB1, PA, HA and NA proteins (Table 2). The LRT results also indicate that the null model M8a is fits better to M1, NS1 and NS2 than those of models MEC/M8, MEC and MEC, respectively. The LRT for models MEC and M8 could only be performed against null model M8a, if the models MEC and/or M8 show positively selected sites. It should be noted that model MEC for NP and M2 proteins and model M8 for all the proteins, except in M1, show no positively selected sites. For M1 protein, model M8 shows single positively selected site. Although, totally 4, 11, 2, 7, 4 and 13, 13, 9, 1 positively selected sites are inferred for PB2, PB1, PA, HA, NA and HA, M2, NS1, NS2 proteins under MEC and M5 models, respectively (Table 1; SF3). However, no statistically reliable positively selected sites are identified in the NIVs proteins, when empirical Bayesian method was used under MEC, M8 and M5 models. That the lower bound of the confidence interval (95 %) could not be >1 for positively selected sites is the reason and, as a result, no dark yellow colored amino acid sites are detected (SF3) (Stern et al. 2007). It is quite obvious that the CML and BS methods are reasonably better for inferring statistically reliable positively selected sites than those of empirical Bayesian method. However, empirical Bayesian approach is inferred more reliable negatively selected sites. It validates the hypothesis that the positive selection is rare as it occurs in few amino acid sites during the short time and is hardly detected effectively when compare to a large amount of neutral and purifying selections (Duret 2008; Nei and Kumar 2000).
Inferring functions for the amino acid sites under selection
Proteins are responsible for a group of biological functions, and identification of biological functions of amino acid sites would be helpful for assigning similar functions to functionally unknown amino acid sites. Due to the lack of prior vaccination, human immune system is unable to ramp-up its responses against novel influenza viruses. As a result, unexpected pandemic outbreaks occurred. Human immune responses against viruses are highly depending on the recognition of conserved epitopes of the viruses by antibodies. A report shows that T-cells are protecting humans in an enhanced way from viral infection and death (Suzuki 2006; Cusick et al. 2009; Hughes and Nei 1988). The B-cell epitopes (BCEs) of the viruses, typically consist of 15–22 continuous/discontinuous amino acid sites that could be recognized by respective B-cells producing antibodies to neutralize the viral infectivity (Klein and Horejsi 1997). Use of 3-D structures of antigen–antibody complexes and monoclonal antibodies, totally 5 (epitopes A–E), 3 (epitopes A–C) and one BCEs were identified in HA (Wiley et al. 1981), NA (Air et al. 1985) and M2 (Zebedee and Lamb 1988), respectively. The T-cell epitopes (TCEs) are classified into CD8+ and CD4+ that are typically consist of 9 and 13–18 continuous amino acid sites, respectively (Klein and Horejsi 1997). The infected cells have CD8+ and CD4+ TCEs along with the human leukocyte antigen (HLA) class I (includes HLA-A, -B, and -C) and HLA class II (includes HLA–DQ and –DR), which are recognized by cytotoxic T-lymphocytes (CTLs) and helper T-cells (Th cells), respectively. These CTLs known to exert cytotoxicity to infected cells. There are two kinds of Th cells namely Th1 and Th2 that could activate the CTLs and B-cells, correspondingly and might also exert cytotoxicity (Suzuki 2006). Macken et al. (2001) were identified CD8+ and CD4+ TCEs in influenza A virus proteins by performing techniques like 3-D structural analysis of HLA–epitope complexes, peptide-binding and lytic assays. In addition to these natural immune responses against viral infections, the artificial acquired immunity could also be produced by vaccines. A typical method of growing influenza viruses for vaccine production is through the use of chick embryos eggs (Smith et al. 2008). Moreover, genome sequences of control virus and the isolate from allantoic cavity of chick embryos egg was compared and identified that the amino acids that facilitate the virus to adapt and grow up in eggs (Hardy et al. 1995). Apart from the natural and artifact immune responses, specific drugs are also available to effectively eliminate the viruses from infected patients. However, a few amino acids of NIVs proteins involved in the resistance to drugs like amantadine and oseltamivir, were identified by performing inhibition assays (Suzuki 2006; Hay et al. 1985; Gubareva et al. 2000). Thus, in the present study, in addition to inferring positive Darwinian amino acid sites, there is also a necessary to interpret these sites with experimentally verified functionally known amino acid sites (Table 3).
Several reports show the positive selection on influenza H1N1, H3N2 and H5N1 viruses (Matrosovich et al. 2000; Suzuki 2006; Campitelli et al. 2006; Wolf et al. 2006; Shen et al. 2009; Furuse et al. 2010; Janies et al. 2010; Chen and Sun 2011; Li et al. 2011). Here, I have interpreted some of the aforementioned previous studies with the present results. There was a report by Suzuki (2006) shows that totally 4 positively selected amino acid sites identified in the entire proteins of human influenza A/H3N2 viruses using parsimony method via ADAPTSITE. The amino acid positions 220 and 229 of HA, position 131 of NP, and position 370 of NA, were inferred as positively selected when he analyzed 284, 246 and 345 sequences, respectively. Suzuki (2006) inferred that the biological functions of amino acid positions 229 of HA and 370 of NA as BCEs epitope D and epitope A, respectively, however, no functions were inferred for 220 of HA and 131 of NP. On the other hand, Campitelli et al. (2006) reported that totally 16 positively selected amino acid sites in the entire proteins of the human and avian influenza A/H5N1 viruses using Bayesian method. The positively selected amino acid sites 17, 82, 199, 334, 336, 355 and 727 of PB2, positions 138, 140, 155, 156, 218 and 227 of HA and positions 171, 205 and 209 of NS1, were identified by evaluating 16, 192 and 31 sequences, respectively. Furthermore, they found that the positively selected sites of HA were all involved in BCEs, TCEs and growth in eggs. However, no functions were inferred for the positively selected sites of PB2 and NS1. Chen and Sun (2011) reported that 43 positively selected sites for HA of subtype human H3N2 influenza viruses using Bayes and Naive Empirical Bayes analyses under M2, M3 and M8 models by analyzing 262 sequences under seven data sets. They assumed that 42 out of 43 positively selected amino acid sites were involved in BCEs. Notably, Furuse et al. (2010) reported totally 8, 4 and 2 amino acid sites under positive selection pressure on the entire HA gene of seasonal H1N1, swine H1 and 2009 H1N1 viruses, respectively using only FEL method in HyPhy (Pond et al. 2006). Among these, positions 187, 190, 192, 225 of seasonal H1N1, positions 83, 192 of swine H1 and position 206 of 2009 H1N1 viruses are located at antigenic sites. In addition, sites 190 and 225 of seasonal H1N1 are also key determinants for effective binding to human-like receptors (Kobasa et al. 2004; Stevens et al. 2006; Tumpey et al. 2007; Furuse et al. 2010). However, the specific roles of rest of the amino acid sites of all above the viruses are unknown. Li et al. (2011) reported totally 9 and 2 amino acid sites under positive selection for HA and NA genes, respectively of pandemic influenza H1N1 viruses using SLAC and FEL methods in HyPhy. Interestingly, 7 (186, 222, 261, 411, 451, 460, 530) out of 9 sites of HA and 1 (453) out of 2 sites of NA are located within the T-cell and/or B-cell antigenic regions. It should be noted that sites 411, 451, 460, 530 of HA and 35, 453 of NA could only be identified by FEL method, but not by SLAC method. In the present study, totally 9 positively selected sites are identified and among these 9 sites, single position from PB2 (site 194) and M2 (10), two positions from PB1 (587, 736) and NS1 (108, 123) and three positions from HA (7, 49, 391) are inferred (Table 1). However, no amino acid sites are inferred as positively selected for PA, NP, NA, M1 and NS2. It should be noted that no similar positively selected sites are identified when comparing the present result with the previous observations (Suzuki 2006; Campitelli et al. 2006; Chen and Sun 2011; Furuse et al. 2010; Li et al. 2011). It could be attributed to the different sequences and different statistical methods, used in the adaptive selection analyses. Differences occur predominantly in use of different subtypes from different seasons and the different statistical methods. It is noted that variation in use of total number of sequences could not affect the sensitivity of the analysis (Chen and Sun 2011). Moreover, aforementioned previous analyses carried out by using single method indicate that the positively selected sites identified might remain tentative, because the multiple tests were not approved. Whereas, the positively selected sites identified in the present study are based on the multiple methods indicate that the amino acid sites under strong selection strength be more reliable.
These positively selected sites were interpreted with functionally known amino acid sites for Darwinian natural selection (Hughes and Nei 1988). Experimentally determined functionally known epitopes along with epitope identification codes, name of the host organisms (which was used to expose the immunogen) and the literature information interpreted in this study, were composed from the IEDB (www.immuneepitope.org) (Vita et al. 2010). The biological function of amino acid position 194 of PB2 is identified as TCEs (Babon et al. 2009) and in support of this, the adjacent site 196 is also found to be involved in TCEs (HLA-H-2–b; –Kb; –Db) (Crowe et al. 2006; Thomas et al. 2007). Interestingly, the position of 587 of PB1, 49 and 391 of HA and 10 of M2 are involved in both BCEs and TCEs. However, sites 736 of PB1 and 108 and 123 of NS1 are found to be involved merely in TCEs. The function of position 7 of HA is unknown; however, the adjacent sites 8 and 9 involved in TCEs, suggesting that site 7 might also be involved in this function (Table 3). Totally, function for 8 out of 9 amino acid sites are exactly inferred with known immune epitopes, whereas the two adjacent sites are inferred for a rest of the site 7 of HA. Among all the positively selected sites, there are single positions from PB1 and M2 and 2 positions from HA are found to be involved in BCEs. However, no sites from PB2 and NS1 are found to be involved in BCEs. Interestingly, some of the sites involved in BCEs are also involved in TCEs. Positively selected sites were positioned in the B-cell and/or T-cell antigenic regions, might indicate that positive selection from the hosts, possibly caused by vaccination and continuous use of anti-viral drugs. It might lead to parallel variations in the T-cell and/or B-cell antigenic regions of the viruses. Hence, this would reduce the efficiency of vaccines and have helped viruses to better adapt to the new hosts (Li et al. 2011). All experimentally derived epitopes available in IEDB are quantitatively categorized either as positive or negative, based on the B-cell and/or T-cell assays (ELISA/FACs/bioassay etc.) ‘Immunogen epitope relation’ reactions, where the epitopes of influenza are recognized by antisera of influenza infected host human/mouse; the immunogen that the host was exposed to was the influenza virus (Vita et al. 2010). The present results also show that few quantitatively negative epitopes (Table 3), which can also be important to verify experimentally, whether it will produce positive response over against NIVs. It would be interesting to examine the functions of these positively selected sites of NIVs, experimentally, using site directed mutagenesis (Hoffmann et al. 2000). Interestingly, no positively selected sites are found to be involved in antiviral resistance and growth in eggs.
Inference of amino acid sites involving in BCEs
In spite of more number of amino acid sites having shown excess of non-synonymous substitutions (positive values) (SF1, 2), the positive selection operating on only a few amino acid sites involved in BCEs and TCEs is supported by the ω > 1 with statistical significance (Table 1). It is observed that positively selected sites for all the BCEs (the p values for PB1, HA, M2 0.004–0.03) are generally efficient when comparing to the sites involved in TCEs (the p values 0.05 for PB2, HA and Bayesian Factors 60.95–64.41 for NS1) (Table 3). Interestingly, it is found that the sites involved in BCEs are also involved in TCEs. Human immune responses against viruses, highly depends on the recognition of conserved BCEs and TCEs by the antibodies, exclusively, T-cells are protecting humans from infection and death (Cusick et al. 2009). However, vaccines are available for influenza viruses to induce immune responses against BCEs (Cox et al. 2004). In support of this issue, in the present study, the positively selected sites are inferred for both BCEs and TCEs, and sites merely inferred for TCEs (Table 3). It indicates that the both humoral and cellular immune responses are engaged in elimination of NIVs from infected humans (Thomas et al. 2006). The present observation is consistent with the previous reports (Thomas et al. 2006; Suzuki and Gojobori 2001) where they also observed both type of immune responses against influenza A viruses and hepatitis C viruses. Moreover, a report states that all the negatively selected sites were apparently involved in the neutralization of BCEs of polio viruses, and the vaccine based on these sites was known to be extremely successful (Suzuki 2004). In my case, totally 4 [single site for PB1, M2 and two sites for HA (p 0.004–0.03)] amino acid sites under positive selection are found to be involved in BCEs (Table 3). The total numbers of positively selected sites engaged in BCEs are lower; it is distinct to the case of polio viruses, where majority of the negatively selected sites involved in BCEs (Suzuki 2004). However, the p values of positively selected sites are more reliable indicate that it is also be possible to use these 4 positively selected sites as targets. It validate the hypothesis that the amino acid sites under significant functional constraints (small p values) are suitable targets for developing vaccine and drugs with less vulnerable to the formation of escape mutants (Bush et al. 1999; Smith et al. 2004). In case, these positively selected sites will be used as targets, the significance humoral immune responses may be expressed towards the targets. As a result, generation of advantageous escape mutant’s could be bigger and outcompete the functional restrictions; however, the fitness of these mutants ought to be less if it is developed. In order to prevent the generation of escape mutants, use of mixture of several vaccines are recommended (Suzuki 2006). In the context of these findings, further experimental study should be focused on to determine, whether the total of 4 positively selected amino acid sites involving in BCEs will be effective in making successful vaccine against NIVs.
There are three different approaches viz. CML, BS and Bayesian used for determining individual amino acid site of NIVs proteins whether under positive selection. Totally, nine sites from PB2, PB1, HA, M2 and NS1, are inferred as positively selected. Functions for positively selected sites are inferred by interpreting these sites with experimentally determined functionally known amino acid sites. Totally, 4 positively selected sites of PB1, HA and M2 are found to be involved in BCEs. Besides, most of the sites involved in BCEs are also involving in TCEs. Amino acid mutations in many TCEs are expected to be advantageous, since, the haplotype of HLA is restrict T-cells to recognize TCEs, but it fails to restrict B-cells to recognize BCEs (Suzuki 2006; Berkhoff et al. 2005). In support of this issue, majority of the sites under positive selection forces are involving in TCEs than those of BCEs. Apart from the use of positively selected sites to identify the epitopes involved in the elimination of viruses from patients, there are strong functional constraints also operating on it. Generating vaccines and drugs for multiple targets would be preferable due to its less susceptibility to make escape mutants (Suzuki 2006; Jin et al. 2005; Zharikova et al. 2005; Venkatramani et al. 2006; Boon et al. 2002a; Gog et al. 2003; Kilbourne et al. 2002). In support of this hypothesis, amino acid sites engaged in both BCEs and TCEs could be measured as highly suitable targets, because these sites playing a predominant role in inducing strong humoral and cellular immune responses against targets. As explained in aforementioned previous reports and in my studies, sites involving in multiple functions might provide some valuable insights for the future design of highly protective vaccines to improve the protection against pandemic influenza infections. Moreover, further experiments are required to validate the accurate role of these sites in vaccine design and development.
References
Air GM, Els MC, Brown LE, Laver WG, Webster RG (1985) Location of antigenic sites on the three-dimensional structure of the influenza N2 virus neuraminidase. Virology 145:237–248
Arunachalam R, Paulkumar K, Annadurai G (2012a) Phylogenetic analysis of pandemic influenza A/H1N1 virus. Biologia 67(1):14–31
Arunachalam R, Paulkumar K, Annadurai G (2012b) Genetic ancestor of external antigens of pandemic influenza A/H1N1 virus. Interdiscip Sci Comput Life Sci 4:282–290
Arunachalam R, Senthilkumar B, Senbagam D, Selvamaleeswaran P, Rajasekarapandian M (2012c) Molecular phylogenetic approach for classification of Salmonella typhi. Res J Microbiol 7(1):13–22
Babon JAB, Cruz J, Orphin L, Pazoles P, Co MDT, Ennis FA, Terajima M (2009) Genome-wide screening of human T-cell epitopes in influenza A virus reveals a broad spectrum of CD4(+) T-cell responses to internal proteins, hemagglutinins, and neuraminidases. Hum Immunol 70:711–721
Bao Y, Bolotov P, Dernovoy D, Kiryutin B, Zaslavsky L, Tatusova T, Ostell J, Lipman D (2008) The influenza virus resource at the National Center for Biotechnology Information. J Virol 82(2):596–601
Berkhoff EGM, de-Wit E, Geelhoed-Mieras MM, Boon ACM, Symons J, Fouchier RA, Osterhaus AD, Rimmelzwaan GF (2005) Functional constraints of influenza A virus epitopes limit escape from cytotoxic T lymphocytes. J Virol 79:11239–11246
Blanquer A, Uriz MJ (2007) Cryptic speciation in marine sponges evidenced by mitochondrial and nuclear genes: a phylogenetic approach. Mol Phylogenet Evol 45:392–397
Boon ACM, de-Mutsert G, Graus YMF, Fouchier RAM, Sintnicolaas K, Osterhaus AD, Rimmelzwaan GF (2002a) Sequence variation in a newly identified HLA-B35-restricted epitope in the influenza A virus nucleoprotein associated with escape from cytotoxic T lymphocytes. J Virol 76:2567–2572
Boon ACM, de-Mutsert G, Graus YMF, Fouchier RAM, Sintnicolaas K, Osterhaus AD, Rimmelzwaan GF (2002b) The magnitude and specificity of influenza A virus-specific cytotoxic T-lymphocyte responses in humans is related to HLA-A and -B phenotype. J Virol 76:582–590
Boon ACM, de-Mutsert G, Fouchier RAM, Osterhaus ADM, Rimmelzwaan GF (2006) The hypervariable immunodominant NP418–426 epitope from the influenza A virus nucleoprotein is recognized by cytotoxic T lymphocytes with high functional avidity. J Virol 80:6024–6032
Bush RM, Fitch WM, Bender CA, Cox NJ (1999) Positive selection on the H3 hemagglutinin gene of human influenza virus A. Mol Biol Evol 16:1457–1465
Campitelli L, Ciccozzi M, Salemi M, Taglia F, Boros S, Donatelli I, Rezza G (2006) H5N1 influenza virus evolution: a comparison of different epidemics in birds and humans (1997–2004). J Gen Virol 87:955–960
Centers for Disease Control and Prevention (CDC) (2009) Swine influenza A(H1N1) infection in two children-Southern California, March–April 2009. Morb Mortal Wkly Rep 58:400–402
Chen J, Sun Y (2011) Variation in the analysis of positively selected sites using nonsynonymous/synonymous rate ratios: an example using influenza virus. PLoS One 6(5):e19996
Cox RJ, Brokstad KA, Ogra P (2004) Influenza virus: immunity and vaccination strategies. Comparison of the immune response to inactivated and live, attenuated influenza vaccines. Scand J Immunol 59:1–15
Crowe SR, Miller SC, Brown DM, Adams PS, Dutton RW, Harmsen AG, Lund FE, Randall TD, Swain SL, Woodland DL (2006) Uneven distribution of MHC class II epitopes within the influenza virus. Vaccine 24:457–467
Cusick MF, Wang S, Eckels DD (2009) In vitro responses to avian influenza H5 by human CD4 T cells. J Immunol 183:6432–6441
Delport W, Poon A-FY, Frost SDW, Pond SLK (2010) Datamonkey 2010: a suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics 26:2455–2457
Dempster AP, Laird NM, Rubin DB (1977) Maximum likelihood estimation from incomplete data via the EM algorithm. J R Stat Soc Ser B 39:1–38
Dixit J, Srivastava H, Sharma M, Das MK, Singh OP, Raghavendra K, Nanda N, Dash AP, Saksena DN, Das A (2010) Phylogenetic inference of Indian malaria vectors from multilocus DNA sequences. Infect Genet Evol 10:755–763
Doron-Faigenboim A, Pupko T (2006) A combined empirical and mechanistic codon model. Mol Biol Evol 24:388–397
Duret L (2008) Neutral theory: the null hypothesis of molecular evolution. Nat Educ 1(1). http://www.nature.com/scitable/topicpage/neutral-theory-the-null-hypothesis-of-molecular-839
Echevarria-Zuno S, Mejia-Arangure JM, Grajales-Muniz C, Gonzalez-Bonilla C, Borja-Aburto VH (2010) Seasonal vaccine effectiveness against pandemic A/H1N1 reply. Lancet 375:802–803
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:113
Furuse Y, Shimabukuro K, Odagiri T, Sawayama R, Okada T, Khandaker I, Suzuki A, Oshitani H (2010) Comparison of selection pressures on the HA gene of pandemic (2009) and seasonal human and swine influenza A H1 subtype viruses. Virology 405:314–321
Garcia-Garcia L, Valdespino-Gomez JL, Lazcano-Ponce E, Jimenez-Corona A, Higuera-Iglesias A, Cruz-Hervert P, Cano-Arellano B, Garcia-Anaya A, Ferreira-Guerrero E, Baez-Saldaña R, Ferreyra-Reyes L, Ponce-de-León-Rosales S, Alpuche-Aranda C, Rodriguez-López MH, Perez-Padilla R, Hernandez-Avila M (2009) Partial protection of seasonal trivalent inactivated vaccine against novel pandemic influenza A/H1N1 2009: case–control study in Mexico City. BMJ 339:b3928
Ghedin E, Sengamalay NA, Shumway M, Zaborsk J, Feldblyum T, Subbu V, Spiro DJ, Sitz J, Koo H, Bolotov P, Dernovoy D, Tatusova T, Bao Y, George KS, Taylor J, Lipman DJ, Fraser CM, Taubenberger JK, Salzberg SL (2005) Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution. Nature 437:1162–1166
Gog JR, Rimmelzwaan GF, Osterhaus AD, Grenfell BT (2003) Population dynamics of rapid fixation in cytotoxic T lymphocyte escape mutants of influenza A. Proc Natl Acad Sci USA 100:11143–11147
Gubareva LV, Kaiser L, Hayden FG (2000) Influenza virus neuraminidase inhibitors. Lancet 355:827–835
Hardy CT, Young SA, Webster RG, Naeve CW, Owens RJ (1995) Egg fluids and cells of the chorioallantoic membrane of embryonated chicken eggs can select different variants of influenza A (H3N2) viruses. Virology 211:302–306
Hay AJ, Wolstenholme AJ, Skehel JJ, Smith MH (1985) The molecular basis of the specific anti-influenza action of amantadine. EMBO J 4:3021–3024
Hoffmann E, Neumann G, Kawaoka Y, Hobom G, Webster RG (2000) A DNA transfection system for generation of influenza A virus from eight plasmids. Proc Natl Acad Sci USA 97:6108–6113
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335:167–170
Janies DA, Voronkin IO, Studer J, Hardman J, Alexandrov BB, Treseder TW, Valson C (2010) Selection for resistance to oseltamivir in seasonal and pandemic H1N1 influenza and widespread co-circulation of the lineages. Int J Health Geogr 9:13
Jin H, Zhou H, Liu H, Chan W, Adhikary L, Mahmood K, Lee MS, Kemble G (2005) Two residues in the hemagglutinin of A/Fujian/411/02-like influenza viruses are responsible for antigenic drift from A/Panama/2007/99. Virology 336:113–119
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Khurana S, Suguitan-Jr AL, Rivera Y, Simmons CP, Lanzavecchia A, Sallusto F, Manischewitz J, King LR, Subbarao K, Golding H (2009) Antigenic fingerprinting of H5N1 avian influenza using convalescent sera and monoclonal antibodies reveals potential vaccine and diagnostic targets. PLoS Med 6:e1000049
Kilbourne ED, Smith C, Brett I, Pokorny BA, Johansson B, Cox N (2002) The total influenza vaccine failure of 1947 revisited: major intrasubtypic antigenic change can explain failure of vaccine in a post-World War II epidemic. Proc Natl Acad Sci USA 99:10748–10752
Kimura M (1968) Evolutionary rate at the molecular level. Nature 217:624–626
Kiso M, Mitamura K, Sakai-Tagawa Y, Shiraishi K, Kawakami C, Kimura K, Hayden FG, Sugaya N, Kawaoka Y (2004) Resistant influenza A viruses in children treated with oseltamivir: descriptive study. Lancet 364:759–765
Klein J, Horejsi V (1997) Immunology, 2nd edn. Blackwell Science, Oxford
Kobasa D, Takada A, Shinya K, Hatta M, Halfmann P, Theriault S, Suzuki H, Nishimura H, Mitamura K, Sugaya N, Usui T, Murata T, Maeda Y, Watanabe S, Suresh M, Suzuki T, Suzuki Y, Feldmann H, Kawaoka Y (2004) Enhanced virulence of influenza A viruses with the haemagglutinin of the 1918 pandemic virus. Nature 431(7009):703–707
Kreijtz JHCM, de-Mutsert G, van-Baalen CA, Fouchier RAM, Osterhaus ADME, Rimmelzwaan GF (2008) Cross-recognition of avian H5N1 influenza virus by human cytotoxic T-lymphocyte populations directed to human influenza A virus. J Virol 82:5161–5166
Lazrak A, Iles KE, Liu G, Noah DL, Noah JW, Matalon S (2009) Influenza virus M2 protein inhibits epithelial sodium channels by increasing reactive oxygen species. FASEB J 23(11):3829–3842
Lee LY-H, Ha DLAH, Simmons C, de-Jong MD, Chau NV, Schumacher R, Peng YC, McMichael AJ, Farrar JJ, Smith GL, Townsend AR, Askonas BA, Rowland-Jones S, Dong T (2008) Memory T cells established by seasonal human influenza A infection cross-react with avian influenza A (H5N1) in healthy individuals. J Clin Invest 118:3478–3490
Li W, Shi W, Qiao H, Ho SYW, Luo A, Zhang Y, Zhu C (2011) Positive selection on hemagglutinin and neuraminidase genes of H1N1 influenza viruses. Virology J 8:183
Liu W, Chen YH (2005) High epitope density in a single protein molecule significantly enhances antigenicity as well as immunogenicity: a novel strategy for modern vaccine development and a preliminary investigation about B cell discrimination of monomeric proteins. Eur J Immunol 35:505–514
Liu W, Peng Z, Liu Z, Lu Y, Ding J, Chen YH (2004) High epitope density in a single recombinant protein molecule of the extracellular domain of influenza A virus M2 protein significantly enhances protective immunity. Vaccine 23:366–371
Macken C, Lu H, Goodman J, Boykin L (2001) The value of a database in surveillance and vaccine selection. In: Osterhaus ADME, Cox N, Hampson AW (eds) Options for the control of influenza IV. Elsevier Science, Amsterdam, pp 103–106
Malickbasha M, Arunachalam R, Senthilkumar B, Rajasekarapandian M, Annadurai G (2010) Effect of ompR gene mutation in expression of ompC and ompF of Salmonella typhi. Interdis Sci Comput Life Sci 2:157–162
Matrosovich M, Tuzikov A, Bovin N, Gambaryan A, Klimov A, Castrucci MR, Donatelli I, Kawaoka Y (2000) Early alterations of the receptor-binding properties of H1, H2, and H3 avian influenza virus hemagglutinins after their introduction into mammals. J Virol 74(18):8502–8512
Mayrose I, Graur D, Ben-Tal N, Pupko T (2004) Comparison of site–specific rate–inference methods for protein sequences: empirical Bayesian methods are superior. Mol Biol Evol 21:1781–1791
Mayrose I, Mitchell A, Pupko T (2005) Site-specific evolutionary rate inference: taking phylogenetic uncertainty into account. J Mol Evol 60:345–353
Mostow SR, Schoenbaum SC, Dowdle WR, Coleman MT, Kaye HS, Hierholzer JC (1970) Studies on inactivated influenza vaccines. II. Effect of increasing dosage on antibody response and adverse reactions in man. Am J Epidemiol 92:248–256
Mozdzanowska K, Feng JQ, Eid M, Kragol G, Cudic M, Otvos L Jr, Gerhard W (2003) Induction of influenza type A virus-specific resistance by immunization of mice with a synthetic multiple antigenic peptide vaccine that contains ectodomains of matrix protein 2. Vaccine 21:2616–2626
Mullick J, Cherian SS, Potdar VA, Chadha MS, Mishra AC (2011) Evolutionary dynamics of the influenza A pandemic (H1N1) 2009 virus with emphasis on Indian isolates: evidence for adaptive evolution in the HA gene. Infect Genet Evol 11:997–1005
Nayak JL, Richards KA, Chaves FA, Sant AJ (2010) Analyses of the specificity of CD4 T cells during the primary immune response to influenza virus reveals dramatic MHC-linked asymmetries in reactivity to individual viral proteins. Viral Immunol 23:169–180
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Noda T, Sagara H, Yen A, Takada A, Kida H, Cheng H, Kawaoka Y (2006) Architecture of ribonucleoprotein complexes in influenza A virus particles. Nature 439:490–492
Obenauer JC, Denson J, Mehta PK, Su X, Mukatira S, Finkelstein DB, Xu X, Wang J, Ma J, Fan Y, Rakestraw KM, Webster RG, Hoffmann E, Krauss S, Zheng J, Zhang Z, Naeve CW (2006) Large-scale sequence analysis of avian influenza isolates. Science 311:1576–1580
Pond SLK, Frost SDW (2005) Not so different after all: a comparison of methods for detecting amino acid sited under selection. Mol Biol Evol 22(5):1208–1222
Pond SLK, Frost SDW, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21(5):676–679
Pond SLK, Frost SDW, Grossman Z, Gravenor MB, Richman DD, Brown AJ (2006) Adaptation to different human populations by HIV-1 revealed by codon-based analyses. PLoS Comput Biol 2(6):e62
Pond SLK, Murrell B, Fourment M, Frost SDW, Delport W, Scheffler K (2011) A random effects branch-site model for detecting episodic diversifying selection. Mol Biol Evol. doi:10.1093/molbev/msr125
Potter CW (2001) A history of influenza. J Appl Microbiol 91:572–579
Qu Y, Zhang R, Cui P, Song G, Duan Z, Lei F (2011) Evolutionary genomics of the pandemic 2009 H1N1 influenza viruses (pH1N1v). Virol J 8:250
Richards KA, Chaves FA, Krafcik FR, Topham DJ, Lazarski CA, Sant AJ (2007) Direct ex vivo analyses of HLA–DR1 transgenic mice reveal an exceptionally broad pattern of immunodominance in the primary HLA–DR1-restricted CD4 T-cell response to influenza virus hemagglutinin. J Virol 81:7608–7619
Richards KA, Chaves FA, Sant AJ (2009) Infection of HLA–DR1 transgenic mice with a human isolate of influenza a virus (H1N1) primes a diverse CD4 T-cell repertoire that includes CD4 T cells with heterosubtypic cross-reactivity to avian (H5N1) influenza virus. J Virol 83:6566–6577
Roti M, Yang J, Berger D, Huston L, James EA, Kwok WW (2008) Healthy human subjects have CD4+ T cells directed against H5N1 influenza virus. J Immunol 180:1758–1768
Schanen BC, De-Groot AS, Moise L, Ardito M, McClaine E, Martin W, Wittman V, Warren WL, Drake DR (2011) Coupling sensitive in vitro and in silico techniques to assess cross-reactive CD4(+) T cells against the swine-origin H1N1 influenza virus. Vaccine 29:3299–3309
Shen J, Ma J, Wang Q (2009) Evolutionary trends of A(H1N1) influenza virus hemagglutinin since 1918. PLoS One 4(11):e7789
Smith W, Andrewes CH, Laidlaw PP (1933) A virus obtained from influenza patients. Lancet 225:66–68
Smith DJ, Lapedes AS, de-Jong JC, Bestebroer TM, Rimmelzwaan GF, Osterhaus ADME, Fouchier RAM (2004) Mapping the antigenic and genetic evolution of influenza virus. Science 305:371–376
Smith KA, Colvin CJ, Weber PSD, Spatz SJ, Coussens PM (2008) High titer growth of human and avian influenza viruses in an immortalized chick embryo cell line without the need for exogenous proteases. Vaccine 26:3778–3782
Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Ma SK, Cheung CL, Raghwani J, Bhatt S, Peiris JS, Guan Y, Rambaut A (2009) Origin and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459(7250):1122–1126
Stern A, Doron-Faigenboim A, Erez E, Martz E, Bacharach E, Pupko T (2007) Selecton 2007: advanced models for detecting positive and purifying selection using Bayesian inference approach. Nucleic Acids Res 35:W506–W511
Stevens J, Blixt O, Glaser L, Taubenberger JK, Palese P, Paulson JC, Wilson IA (2006) Glycan microarray analysis of the hemagglutinins from modern and pandemic influenza viruses reveals different receptor specificities. J Mol Biol 355(5):1143–1155
Suzuki Y (2004) Negative selection on neutralization epitopes of poliovirus surface proteins: implications for prediction of candidate epitopes for immunization. Gene 328:127–133
Suzuki Y (2006) Natural selection on the Influenza virus genome. Mol Biol Evol 23(10):1902–1911
Suzuki Y, Gojobori T (1999) A method for detecting positive selection at single amino acid sites. Mol Biol Evol 16(10):1315–1328
Suzuki Y, Gojobori T (2001) Positively selected amino acid sites in the entire coding region of hepatitis C virus subtype 1b. Gene 276:83–87
Suzuki Y, Nei M (2002) Origin and evolution of influenza virus hemagglutinin genes. Mol Biol Evol 19:501–509
Swanson WJ, Nielsen R, Yang Q (2003) Pervasive adaptive evolution in mammalian fertilization proteins. Mol Biol Evol 20:18–20
Tamura K, Nei M (1993) Estimation of the number of nucleotide substituions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 11:715–724
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetic analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Thomas PG, Keating R, Hulse-Post DJ, Doherty PC (2006) Cell mediated protection in influenza infection. Emerg Infect Dis 12:48–54
Thomas PG, Brown SA, Keating R, Yue W, Morris MY, So J, Webby RJ, Doherty PC (2007) Hidden epitopes emerge in secondary influenza virus-specific CD8+ T cell responses. J Immunol 178:3091–3098
Trojan A, Urosevic M, Hummerjohann J, Giger R, Schanz U, Stahel RA (2003) Immune reactivity against a novel HLA-A3-restricted influenza virus peptide identified by predictive algorithms and interferon-gamma quantitative PCR. J Immunol 26:41–46
Tu W, Mao H, Zheng J, Liu Y, Chiu SS, Qin G, Chan PL, Lam KT, Guan J, Zhang L, Guan Y, Yuen KY, Peiris JS, Lau YL (2010) Cytotoxic T lymphocytes established by seasonal human influenza cross-react against 2009 pandemic H1N1 influenza virus. J Virol 84(13):6527–6535
Tumpey TM, Maines TR, Van-Hoeven N, Glaser L, Solorzano A, Pappas C, Cox NJ, Swayne DE, Palese P, Katz JM, Garcia-Sastre A (2007) A two-amino acid change in the hemagglutinin of the 1918 influenza virus abolishes transmission. Science 315(5812):655–659
Venkatramani L, Bochkareva E, Lee JT, Gulati U, Laver WG, Bochkarev A, Air GM (2006) An epidemiologically significant epitope of a 1998 human influenza virus neuraminidase forms a highly hydrated interface in the NA-antibody complex. J Mol Biol 356:651–663
Vita R, Zarebski L, Greenbaum JA, Emami H, Hoof I, Salimi N, Damle R, Sette A, Peters B (2010) The immune epitope database 2.0. Nucleic Acids Res 38(Database issue):D854–D862
Webby RJ, Webster RG (2003) Are we ready for pandemic influenza? Science 302(5650):1519–1522
Webster RG, Kawaoka Y, Bean WJ (1986) Vaccination as a strategy to reduce the emergence of amantadine- and rimantadine-resistant strains of A/chick/Pennsylvania/83 (H5N2) influenza virus. J Antimicrob Chemother 18:157–164
Wiley DC, Wilson IA, Skehel JJ (1981) Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 289:373–378
Wolf YI, Viboud C, Holmes EC, Koonin EV, Lipman DJ (2006) Long intervals of stasis punctuated by bursts of positive selection in the seasonal evolution of influenza A virus. Biol Direct 1:34
World Health Organization (WHO) (1980) A revision of the system of nomenclature for influenza viruses: a WHO memorandum. Bull WHO 58:585–591
Wu F, Yuan X-Y, Li J, Chen Y-H (2009) The co-administration of CpG–ODN influenced protective activity of influenza M2e vaccine. Vaccine 27:4320–4324
Yang Z, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15(12):496–503
Yang Z, Nielsen R, Goldman N, Pedersen AM (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Yang J, Mai D, LaFond RE, Gates TJ, James EA, Malhotra U, Kwok WW (2011) H1N1 influenza, H9N2 influenza, yellow fever virus, and west nile virus specific CD4+ T cells epitopes restricted by various DR alleles. Available from http://www.immuneepitope.org/assayId/1816085
Zamarin D, Ortigozza MB, Palese P (2006) Influenza A virus PB1–F2 protein contributes to viral pathogenesis in mice. J Virol 80(16):7976–7983
Zebedee SL, Lamb RA (1988) Influenza A virus M2 protein: monoclonal antibody restriction of virus growth and detection of M2 in virions. J Virol 62:2762–2772
Zhao R, Cui S, Guo L, Wu C, Gonzalez R, Paranhos-Baccalà G, Vernet G, Wang J, Hung T (2011) Identification of a highly conserved h1 subtype-specific epitope with diagnostic potential in the hemagglutinin protein of influenza a virus. PLoS One 6:e23374
Zharikova D, Mozdzanowska K, Feng J, Zhang M, Gerhard W (2005) Influenza type A virus escape mutants emerge in vivo in the presence of antibodies to the ectodomain of matrix protein-2. J Virol 79:6644–6654
Acknowledgments
Author thanks Dr. G. Annadurai, MS University, India and Prof. K.G. Sivaramakrishnan, Madras Christian College, Chennai, India for their fruitful comments. Author also thanks Prof. Mark Johnson and Dr. Tiina Salminen for the excellent computing facilities at the Structural Bioinformatics Laboratory (Åbo Akademi University, Finland) funded by Biocenter Finland infrastructure (bioinformatics and translational activities); Sigrid Juselius; Tor, Joe, and Pentti Borg Foundations. Author was partially supported by the Finnish Government Scholarship of Centre for International Mobility (CIMO), Finland [KM-11-7352, 05.05.2011] and University Grants Commission [F.87-5/2009(IC), 28.06.2011], India. No funders had a role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Arunachalam, R. Detection of site-specific positive Darwinian selection on pandemic influenza A/H1N1 virus genome: integrative approaches. Genetica 141, 143–155 (2013). https://doi.org/10.1007/s10709-013-9713-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-013-9713-x