Abstract
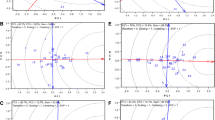
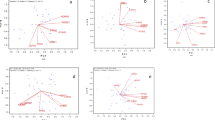
Genotype × environment (GE) interaction can difficult soybean breeding programs to achieve the aim of more productive cultivars. Environment stratification is a way to circumvent this problem. However, multiyear data studies are difficult to work with, because they are, usually, unbalanced. GGE Biplot is an efficient method to find mega-environments, however, it allows for, at most, 30% of the unbalanced data. Thinking about how we could resolve this problem we came up with the idea of test a method that englobes GGE Biplot graphs, environment coincidence matrices and networks of environment. Wherefore, this work aimed to gather GGE Biplot graphs of a network of trials unbalance multiyear soybean via matrices of coincidence and networks of environment to optimize environmental stratification. Data from an experimental network of 43 trials was used, these experiments were implanted in 23 municipalities during the crop seasons of 2011/2012, 2012/2013, 2013/2014 and 2015/2016 in Brazil. The trials were implanted under an experimental block design with randomized treatments and approximately 30 genotypes were evaluated, of which most of these genotypes were not repeated between the trials evaluated. The GE interaction were statistically significant for all 43 trials. The step by step of our analyses was: GGE Biplots graphs were obtained; the environment coincidence matrices were calculated; the values of matrices were used to obtain the networks of environmental similarity. The study demonstrated that by the method was possible to identify, using unbalanced multiyear data, four mega-environments. The region under study can be represented by the municipalities of Palotina, Maracaju, Bela Vista do Paraíso and Rolândia. Therefore, integrating GGE Biplot graphs and networks of environmental similarity is an efficient method to optimize a soybean program by environment stratification.





Similar content being viewed by others
References
Choudhary M, Kumar B, Kumar P, Guleria SK, Singh NK, Khulbe R, Kamboj MC, Vyas M, Srivastava RK, Puttaramanaik SD, Mahajan V, Rakshit S (2019) GGE biplot analysis of genotype × environment interaction and identification of mega-environment for baby corn hybrids evaluation in India. Indian J Genet 79(4):658–669. https://doi.org/10.31742/IJGPB.79.4.3
Cruz CD (2013) GENES–GENES—a software package for analysis in experimental statistics and quantitative genetics. Acta Scientiarum—Agronomy 35(3):271–276. https://doi.org/10.4025/actasciagron.v35i3.21251
Cruz CD, Regazzi AJ (2007) Modelos Biométricos Aplicados ao Melhoramento Genético, 4th ed. Editora UFV
Dalló SC, Zdziarski AD, Woyann LG, Milioli AS, Zanella R, Conte J, Benin G (2019) Across year and year-by-year GGE biplot analysis to evaluate soybean performance and stability in multi-environment trials. Euphytica 215(6):1–12. https://doi.org/10.1007/s10681-019-2438-x
da Silva KJ, Teodoro PE, da Silva MJ, Teodoro LPR, Cardoso MJ, Godinho VDPC, Mota JH, Simon GA, Tardin FD, da Silva AR, Guedes FL, de Menezes CB (2021) Identification of mega-environments for grain sorghum in Brazil using GGE biplot methodology. Agron J 113(4):3019–3030. https://doi.org/10.1002/agj2.20707
Hühn M, Truberg B (2002) Contributions to the analysis of genotype x environment interactions: theoretical results of the application and comparison of clustering techniques for the stratification of field test sites. J Agron Crop Sci 188(2):65–72. https://doi.org/10.1046/j.1439-037X.2002.00549.x
Kaster M (2012) Regionalização dos testes de Valor de Cultivo e Uso e da indicação de cultivares de soja - terceira Aproximação pp 231–235
Marcolin F, Vezzetti E (2017) Novel descriptors for geometrical 3D face analysis. Multimedia Tools Appl 76(12):13805–13834. https://doi.org/10.1007/s11042-016-3741-3
Oliveira, AMdaS de AMdaS, Hamawaki OT, Neto JOdeO, Penna JCV, Juliatti FC, de Souza SA, Hühn M, Truberg B, Cruz CD, R AJ, Egazzi (2007) Modelos Biométrocos Aplicados ao Melhoramento Genético. In J Agron Crop Sci 4th ed., vol 188, Issue 2. Editora UFV. https://doi.org/10.1046/j.1439-037X.2002.00549.x
Pelúzio JM, Fidelis RR, Giongo P, Silva JCD, Cappellari D, Barros HB (2008) Adaptabilidade e estabilidade de cultivares de soja em quatro épocas de semeadura no sul do Estado do Tocantins
Ramburan S, Zhou M, Labuschagne MT (2012) Investigating test site similarity, trait relations and causes of genotype× environment interactions of sugarcane in the Midlands region of South Africa. Field Crop Res 129:71–80. https://doi.org/10.1016/j.fcr.2012.01.017
R (CoreTeam) (2017) R: a language and environment for statistical computing 2, 1–12. https://www.r-project.org/
Silva RR, Benin G (2012) Análises Biplot: conceitos, interpretações e aplicações. Ciência Rural 42(8):1404–1412. https://doi.org/10.1590/s0103-84782012000800012
Silva GN, Júnior ACS, Sant’Anna IC, Cruz CD, Nascimento M, Soares PC (2020) Similarity networks for the classification of rice as to adaptability and stability. Scielo. https://doi.org/10.1590/S1678-3921.pab2020.v55.01017
Sousa MBE, Damasceno-Silva KJ, Rocha MDM, Menezes Júnior JÂND, Lima LRL (2018) Genotype by environment interaction in cowpea lines using GGE biplot method. Revista Caatinga 31:64–71. https://doi.org/10.1590/1983-21252018v31n108rc
Woyann LG, Benin G, Storck L, Trevizan DM, Meneguzzi C, Marchioro VS, Tonatto M, Madureira A (2017) Estimation of missing values affects important aspects of GGE biplot analysis. Crop Sci 57(1):40–52. https://doi.org/10.2135/cropsci2016.02.0100
Yan W (2001) GGEbiplot—a windows application for graphical analysis of multienvironment trial data and other types of two-way data. Agron J 93(5):1111–1118. https://doi.org/10.2134/agronj2001.9351111x
Yan W (2015) Mega-environment analysis and test location evaluation based on unbalanced multiyear data. Crop Sci 55(1):113–122. https://doi.org/10.2135/cropsci2014.03.0203
Yan W (2011) GGE biplot versus AMMI graphs for genotype-by-environment data analysis. J Indian Soc Agric Stat 95:181–193. https://www.cabdirect.org/cabdirect/abstract/20113313412
Yan W, Frégeau-Reid J (2018) Genotype by Yield∗Trait (GYT) Biplot: a novel approach for genotype selection based on multiple traits. Sci Rep 8(1):1–10. https://doi.org/10.1038/s41598-018-26688-8
Yan W, Hunt LA, Sheng Q, Szlavnics Z (2000) Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Sci 40(3):597–605. https://doi.org/10.2135/cropsci2000.403597x
Yan W, Kang MS, Ma B, Woods S, Cornelius PL (2007) GGE biplot versus AMMI analysis of genotype-by-environment data. Crop Sci 47(2):643–655. https://doi.org/10.2135/cropsci2006.06.0374
Yan W, Tinker NA (2006) Biplot analysis of multi-environment trial data: principles and applications. Can J Plant Sci 86(3):623–645. https://doi.org/10.4141/P05-169
Acknowledgements
The authors thank GDM Seeds (GDM Genética do Brasil S.A.) for providing the multiyear data. The authors also thank the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the financial suport.
Funding
This work was supported the Fundação de Amparo à Pesquisa do Estado de Minas Gerais—FAPEMIG, the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES and the Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rodrigues, F.C., Silva, F.C.S., Carneiro, P.C.S. et al. Environmental stratification in trials of unbalanced multiyear soybean (Glycine max (l.) Merril) via the integration of GGE Biplot graphs and networks of environmental similarity. Euphytica 218, 71 (2022). https://doi.org/10.1007/s10681-022-02994-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-022-02994-1