Abstract
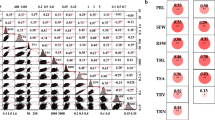
Soil salinization has become one of the important factors affecting the sustainable development of agriculture. Among them, the soda-alkali soil composed of sodium carbonate (Na2CO3) has a more serious impact on crops. In this study, 175 Brassica napus accessions from different sources were used as materials, combined with Brassica 60 K SNP chip data, and the root length traits of Brassica napus during germination under Na2CO3 stress (0.15%) and control conditions were analyzed for genome-wide association (GWAS). GWAS analysis detected that 5 SNPs were significantly related to root length traits under stress conditions; a single SNP could explain 10.22–12.01% of the phenotypic variation. A total of 15 candidate genes related to Na2CO3 stress resistance were identified upstream and downstream of significant SNPs, including cation exchange protein genes (CAX1), members of the zinc finger protein family (ZFHD1), peroxidase family proteins (POD), and transcription factors (MYB family and WRKY family), etc. The expression analysis of 5 candidate genes in extreme phenotypic materials showed that BnaA04g21850D (CAX1) and BnaA06g24040D (ACX5) were induced by Na2CO3 stress in both materials; BnaA06g31200D (ZFHD1) and BnaC02g37590D (MYB60) were up-regulated expression in sensitive materials; BnaA04g21990D (POD) was up-regulated expression in alkali-tolerant materials, indicating that these candidate genes may be involved in the process of rapeseed response to Na2CO3 stress. This study can provide a reference for in-depth analysis of the molecular mechanism of Na2CO3 stress resistance in rapeseed.





Similar content being viewed by others
References
Ashraf M, Foolad MR (2005) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216. https://doi.org/10.1016/j.envexpbot.2005.12.006
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Chalhoub B, Denoeud F, Liu S et al (2014) Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953. https://doi.org/10.1126/science.1253435
Chen L, Wan H, Qian J, Guo J, Sun C, Jing W, Yi B, Ma C, Tu J, Song L, Fu T, Shen J (2018) Genome-wide association study of cadmium accumulation at the seedling stage in rapeseed (Brassica napus L.). Front Plant Sci 9:375. https://doi.org/10.3389/fpls.2018.00375
Chuamnakthong S, Mampei M, Ueda A (2019) Characterization of Na+ exclusion mechanism in rice under saline-alkaline stress conditions. Plant Sci 287:110171. https://doi.org/10.1016/j.plantsci.2019.110171
Crawford R (1990) Importance of root to shoot communication in the responses to environmental stress. Phytochemistry 30:3499
Funck D, Baumgarten L, Stift M, Wirén NV, Schnemann L (2020) Differential contribution of P5CS isoforms to stress tolerance in Arabidopsis. Front Plant Sci 11:565134. https://doi.org/10.3389/fpls.2020.565134
Gong B, Wang X, Wei M, Yang F, Li Y, Shi Q (2016) Over-expression of S-adenosylmethionine synthetase 1 enhances tomato callus tolerance to alkali stress through polyamine and hydrogen peroxide cross-linked networks. Plant Cell Tiss Org Cult 124:377–391. https://doi.org/10.1007/s11240-015-0901-5
He Y, Wu D, You J, Qian W (2017) Genome-wide association analysis of salt tolerance related traits in Brassica napus and candidate gene prediction. Sci Agric Sin 50:1189–1201. https://doi.org/10.3864/j.issn.0578-1752.2017.07.002
He Y, Yang X, Xu C, Guo D, Niu L, Wang Y, Li J, Yan F, Wang Q (2018) Overexpression of a novel transcriptional repressor GmMYB3a negatively regulates salt-alkali tolerance and stress-related genes in soybean. Biochem BiophRes Co. https://doi.org/10.1016/j.bbrc.2018.03.026
Jin H, Kim HR, Plaha P, Liu SK, Park JY, Piao YZ, Yang ZJ, Jiang GB, Kwak SS, An G, Son M, Jin YH, Sohn JH, Lim YP (2008) Expression profiling of the genes induced by Na2CO3 and NaCl stresses in leaves and roots of Leymus chinensis. Plant Sci 175:784–792. https://doi.org/10.1016/j.plantsci.2008.07.016
Jülke S, Ludwig-Müller J (2016) Response of Arabidopsis thaliana roots with altered lipid transfer protein (LTP) gene expression to the clubroot disease and salt stress. Plants 5:2. https://doi.org/10.3390/plants5010002
Jung HW, Kim KD, Hwang BK (2005) Identification of pathogen-responsive regions in the promoter of a pepper lipid transfer protein gene (caltpi) and the enhanced resistance of the caltpi transgenic arabidopsis against pathogen and environmental stresses. Planta 221:361–373
Li F, Chen B, Xu K, Wu J, Song W, Ian B, Harper A, Martin T, Liu S, Gao G, Wang N, Yan G, Qiao J, Li J, Li H, Xiao X, Zhang T, Wu X (2014) Genome-wide association study dissects the genetic architecture of seed weight and seed quality in rapeseed (Brassica napus L.). DNA Res 2014:355–367. https://doi.org/10.1093/dnares/dsu002
Lindorff-Larsen K, Winther JR (2001) Surprisingly high stability of barley lipid transfer protein, LTP1, towards denaturant, heat and proteases. Febs Lett. https://doi.org/10.1016/S0014-5793(00)02424-8
Liu S, Fan C, Li J, Cai G, Yang Q, Wu J, Yi X, Zhang C, Zhou Y (2016) A genome-wide association study reveals novel elite allelic variations in seed oil content of Brassica napus. Theor Appl Genet 129:1203–1215. https://doi.org/10.1007/s00122-016-2697-z
Ma H, Meng C, Zhang K, Wang K, Fan H, Li Y (2020) Study on physiological mechanism of using cottonseed meal to improve salt-alkali tolerance of cotton. J Plant Growth Regul 488:145–148. https://doi.org/10.1007/s00344-020-10083-7
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Ann Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Navarro-León E, López-Moreno FJ, Torre-González ADL, Ruiz JM, Blasco B (2020) Study of salt-stress tolerance and defensive mechanisms in Brassica rapa CAX1a TILLING mutants. Environ Exp Bot 175:104061. https://doi.org/10.1016/j.envexpbot.2020.104061
Schilmiller AL, Howe K (2007) Functional diversification of Acyl-Coenzyme A oxidases in jasmonic acid biosynthesis and action. Plant Physiol 143(2):812–824
Seifert GJ, Xue H, Tuba A (2014) The Arabidopsis thaliana FASCICLIN LIKE ARABINOGALACTAN PROTEIN 4 gene acts synergistically with abscisic acid signalling to control root growth. Ann Bot 6:1125–1133. https://doi.org/10.1093/aob/mcu010
Song J, Guan Z, Hu J, Guo C, Yang Z, Wang S, Liu D, Wang B, Lu S, Zhou R, Xie W, Cheng Y, Zhang Y, Liu K, Yang Q, Chen L, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6:1–12. https://doi.org/10.1038/s41477-019-0577-7
Susanne B (2002) Jasmonate-related mutants of Arabidopsis as tools for studying stress signaling. Planta 214:497–504. https://doi.org/10.1007/s00425-001-0688-y
Tang S, Zhao H, Lu S, Yu L, Zhang G, Zhang Y, Yang Q, Zhou Y, Wang X, Ma W, Xie W, Guo L (2021) Genome-and transcriptome-wide association studies provide insights into the genetic basis of natural variation of seed oil content in Brassica napus. Mol Plant 14:470–487. https://doi.org/10.1016/j.molp.2020.12.003
Tran LSP, Nakashima K, Sakuma Y, Maruyama K, Shinozaki K, Yamaguchi-Shinozaki K (2005) Isolation and functional analysis of Arabidopsis stress-inducible zinc finger homeodomain transcription factor ZFHD1: role of the ZFHD1 and NAC transcription factors in drought-inducible expression of the early responsive to dehydration stress 1 gene. Plant Cell Physiol. https://doi.org/10.14841/jspp.2005.0.599.0
Turner SD (2014) qqman: an R package for visualizing GWAS results using QQ and manhattan plots. BioRxiv 1:005165. https://doi.org/10.1101/005165
Vestin J, Nambu K, Hees P, Bylund D (2006) The influence of alkaline and non-alkaline parent material on soil chemistry. Geoderma 135:97–106
Wan H, Chen L, Guo J, Li Q, Wen J, Yi B, Ma C, Tu J, Fu T, Shen J (2017) Genome-wide association study reveals the genetic architecture underlying salt tolerance-related traits in rapeseed (Brassica napus L.). Front Plant Sci 8:593. https://doi.org/10.1101/005165
Wan H, Wei Y, Qian J, Gao Y, Guo J, Li Q, Wen J, Yi B, Ma C, Tu J, Fu T, Shen J (2018) Association mapping of salt tolerance traits at germination stage of rapeseed (Brassica napus L.). Euphytica 214:190. https://doi.org/10.1007/s10681-018-2272-6
Wang L, Fang C, Wang K (2015) Physiological responses of to long-term salt, alkali and mixed salt-alkali stresses. J Plant Nutr 38:526–540. https://doi.org/10.1080/01904167.2014.937874
Xiong L, Gong Z, Rock CD, Subramanian S, Guo Y, Xu W, Galbraith D, Zhu JK (2001) Modulation of abscisic acid signal transduction and biosynthesis by an Sm-like protein in Arabidopsis. Dev Cell 1:771–781. https://doi.org/10.1016/S1534-5807(01)00087-9
Xu L, Hu K, Zhang Z, Guan C, Chen S, Wei H, Li J, Wen J, Yi B, Ma C, Tu J, Fu T (2016) Genome-wide association study reveals the genetic architecture of flowering time in rapeseed (Brassica napus L.). DNA Res 23:43–52. https://doi.org/10.1093/dnares/dsv035
Yong HY, Wang C, Bancroft I, Li F, Wu X, Kitashiba H, Nishio T (2015) Identification of a gene controlling variation in the salt tolerance of rapeseed (Brassica napus L.). Planta 242:313–326. https://doi.org/10.1007/s00425-015-2310-8
Zeng C, Wan H, Wu X, Dai X, Chen J, Ji Q, Qian F (2021) Genome-wide association study of low nitrogen tolerance traits at the seedling stage of rapeseed. Bio Plant 65:10
Zhang M, Smith JAC, Harberd NP, Jiang C (2016) The regulatory roles of ethylene and reactive oxygen species (ROS) in plant salt stress responses. Plant Mol Biol 9:651–659. https://doi.org/10.1007/s11103-016-0488-1
Zhang R et al (2017) Genome-wide association study of root length and hypocotyl length at germination stage under saline conditions in Brassica napus. Sci Agric Sin 50(1):15–27. https://doi.org/10.3864/j.issn.0578-1752.2017.01.002
Zhao C, Zhang H, Song C, Zhu JK, Shabala S (2020) Mechanisms of plant responses and adaptation to soil salinity. Innovation 1:100017. https://doi.org/10.1016/j.xinn.2020.100017
Zhu J, Liu J, Xiong L (1998) Genetic analysis of salt tolerance in Arabidopsis. Evidence for a critical role of potassium nutrition. Plant Cell 10:1181–1192. https://doi.org/10.2307/3870720
Funding
The study was supported by the China Agriculture Research System (CARS-12) and the Open Fund Project of Engineering Technology Research Center for Protection, Development and Utilization of Characteristic Biological Resources in the Hanjiang River Basin of Hubei Province (06450003).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: HW, CZ, JS and JW. Performed the experiments and analyzed the data: JC, HZ and JT. Data acquisition:JC, HZ, CL, JR, JT, and JL. Wrote the paper: JC. Contributed to writing the manuscript: HZ, CL, JT and HW. Revised the manuscript: HW, CZ and JS.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The experiments shown in the manuscripts submitted for publication comply with the current laws of the country in which they were performed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, J., Zhang, H., Tong, J. et al. Genome-wide association analysis of root length traits in Brassica napus at germination stage under sodium carbonate stress. Euphytica 217, 197 (2021). https://doi.org/10.1007/s10681-021-02928-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-021-02928-3