Abstract
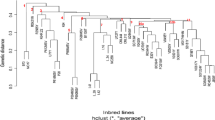
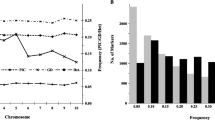
Establishing detailed information on genetic diversity and relationships among maize (Zea mays L.) inbred lines in a breeding programme help ensure effective utilization of germplasm. Introgression of temperate maize germplasm into 12 tropical elite maize inbred lines using a common donor inbred line and different selection environments might disrupt the heterotic grouping system. Therefore, the objective of the study was to determine the effect of introgression on the clustering pattern of the newly introgressed inbred lines and to identify unique genotypes for breeding. A total of 76 introgressed inbred lines derived through pedigree crosses of introgression of 12 elite tropical inbred lines from three major heterotic groups N3, SC and P with a common donor temperate maize inbred line (08CED6_7_B) were used in the study. In addition to the derived 76 introgressed inbred lines, 26 temperate parental inbred lines and 21 elite tropical parental inbred lines with known heterotic group classification were also included in the study as references samples for heterotic groupings to give a total of 123 inbred lines for the study. Introgressed inbred lines derived from four generations of pedigree selection in three distinct environments in Zimbabwe and South Africa were characterised using 20 SSR markers. The 20 SSR markers proved very effective in discriminating the introgressed inbred lines according to genetic distance and clustering. A total of 83 alleles were detected with an average of 4.15 alleles per locus, and had an allelic diversity of 0.53 and PIC of 0.47. Introgression of temperate maize germplasm into tropical elite inbred lines did not disrupt heterotic groupings, because introgressed inbred lines remained genetically inclined towards the original heterotic groups from which they were derived. However, there were some introgressed inbred lines (14%), which did not show any inclination to any of the heterotic groups. Marked genetic difference noted within and across heterotic groupings for the introgressed inbred lines provides germplasm variation that can be exploited for inbred line development and hybrid formation, respectively.




Similar content being viewed by others
References
Abadassi J, Herve Y (2000) Introgression of temperate germplasm to improve an elite tropical maize population. Euphytica 113:125–133
http://www.dnalandmarks.ca. Accessed 18 Sept 2013
http://www.genemapperV40.com. Accessed 18 Sept 2013
Ahmad SQ, Khan S, Ghaffar M, Ahmad F (2011) Gene action for some agronomic traits in maize (Zea mays L.). Crop Breed J 1:133–141
Ali F, Shah IA, Rahman H ur, Noor M, Durrishahwar M, Khan Y, Ullah I, Yan J (2012) Heterosis for yield and agronomic attributes in diverse maize germplasm. Aust J Crop Sci 6:455–462
Bordes J, Charmet G, de Vaulx RD, Lapierre A, Pollacsek M et al (2007) Doubled-haploid versus single-seed descent and S1-family variation for testcross performance in a maize population. Euphytica 154:41–51
Choukan R, Hossainzadeh A, Ghannadha MR, Warburton ML, Talei AR et al (2006) Use of SSR data to determine relationships and potential heterotic groupings within medium to late maturing Iranian maize inbred lines. Field Crops Res 95:212–222
George MLC, Regalado E, Li W, Cao M, Dahlan M et al (2004) Molecular characterization of Asian maize inbred lines by multiple laboratories. Theor Appl Genet 109:80–91
Goodman MM (1999) Broadening the genetic diversity in maize breeding by use of exotic germplasm. Madison, USA
Hallauer AR, Miranda JB (1988) Quantitative genetics in maize breeding. Iowa State University Press, Ames
Laborda PR, Oliveira KM, Garcia AAF, Paterniani MEAGZ, de Souza AP (2005) Tropical maize germplasm: What can we say about its genetic diversity in the light of molecular markers? Theor Appl Genet 111:1288–1299
Liu J (2004) Powermaker V.3.25. http://www.powermarker.net. Accessed 18 Sept 2013
McClosky B, Tanksley SD (2013) The impact of recombination on short-term selection gain in plant breeding experiments. Theor Appl Genet 126:2299–2312
Nei M (1983) Relative efficiencies of different tree-making methods for molecular data. In: Miyamoto MM, Cracraft J (eds) Phylogenetic analysis of DNA sequence. Oxford University Press, New York, pp 90–128
Nelson PT, Goodman MM (2008) Evaluation of elite exotic maize inbreds for use in temperate breeding. Crop Sci 48:85–92
Nelson PT, Jones MP, Goodman MM (2006) Selecting among available, elite tropical maize inbreds for use in long term temperate breeding. Maydica 51:255–262
Pabendon MB, Dahlan M, Sutrisno E, Regalado E, George ML (2004) Preliminary study of genetic diversity of Indonesian maize inbreds collection. In: Srinivasan G, Zaidi PH, Prasanna BM, Gonzalez F, Lesnick K (eds) Proceedings of the eighth Asian regional maize workshop: new technologies for the new millennium Bangkok, Thailand 5–8 August 2002. CIMMYT, Mexico
Patto MCV, Satovic Z, Pego S, Fevereiro P (2004) Assessing the genetic diversity of Portuguese maize germplasm using microsatellite markers. Euphytica 137:63–72
Prasanna BM (2012) Diversity in global maize germplasm: characterization and utilization. J Biol Sci 37:843–855
Prasanna BM, Mohammadi SA, Sudan C, Nair SK, Garg A et al (2004) Application of molecular marker technologies for maize improvement in India—present status and prospects. In: Srinivasan G, Zaidi PH, Prasanna BM, Gonzalez F, Lesnick K (eds) Proceedings of the eighth Asian regional maize workshop: New technologies for the new millennium. Bangkok, Thailand 5–8 August 2002. CIMMYT, Mexico
Reif JC, Fischer S, Schrag TA, Lamkey KR, Klein D et al (2010) Broadening the genetic base of European maize heterotic pools with US Cornbelt germplasm using field and molecular marker data. Theor Appl Genet 120:301–310
Sharopova N, McMullen MD, Schultz L, Schroeder S, Sanchez-Villeda H et al (2002) Development and mapping of SSR markers for maize. Plant Mol Biol 48:463–481
Singh BD (2006) Plant breeding principles and methods. Darya Ganj, New Delhi
Tallury SP, Goodman MM (1999) Experimental evaluation of the potential of tropical germplasm for temperate maize improvement. Theor Appl Genet 98:54–61
Tarter JA, Goodman MM, Holland JB (2004) Recovery of exotic alleles in semi exotic maize inbreds derived from crosses between Latin American accessions and a temperate line. Theor Appl Genet 109:609–617
Wang CL, Cheng FF, Sun ZH, Tang JH, Wu LC et al (2008) Genetic analysis of photoperiod sensitivity in a tropical by temperate maize recombinant inbred population using molecular markers. Theor Appl Genet 117:1129–1139
Wende A, Shimelis H, Derera J, Mosisa W, Danson et al (2012) Genetic interrelationships among medium to late maturing tropical maize inbred lines using selected SSR markers. Euphytica 191:269–277
Wijnker E, de Jong H (2008) Managing meiotic recombination in plant breeding. Trends Plant Sci 13:640–646
Xiao J, Li J, McCouch SR, Tanksley SD (1996) Genetic diversity and its relationship to hybrid performance and heterosis in rice as revealed by PCR-based markers. Theor Appl Genet J 92:637–643
Yao Q, Yang K, Pan G, Rong T (2007) Genetic diversity of maize (Zea mays L.) landraces from South west China based on SSR data. J Genet Genom 34:851–860
Zhang Q, Zhou ZO, Yang GP, Xu CG, Liu KD et al (1996) Molecular marker heterozygosity and hybrid performance in India and Japonica rice. Theor Appl Genet J 93:1218–1224
Zhang S, Li X, Yuan L, Li M, Peng Z (2004) Heterotic groups and exploitation of heterosis-methodology, strategy, and use in hybrid maize breeding in China. In: Srinivasan G, Zaidi PH, Prasanna BM, Gonzalez F, Lesnick K (eds) Proceedings of the eighth Asian regional maize workshop: new technologies for the new millennium. Bangkok, Thailand 5–8 August 2002. CIMMYT, Mexico
Zhi-zhai L, Rong-hua G, Jiu-ran Z, Yi-lin C, Feng-ge W et al (2010) Analysis of Genetic Diversity and population structure of maize landraces from the south maize region of China. Agric Sci China 9:1251–1262
Acknowledgements
This work was fully supported and funded by Seed Co Pvt Ltd. We thank all research personnel at various sites for conducting the trials.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare there to be no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Musundire, L., Derera, J., Dari, S. et al. Molecular characterisation of maize introgressed inbred lines bred in different environments. Euphytica 215, 46 (2019). https://doi.org/10.1007/s10681-019-2367-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2367-8