Abstract
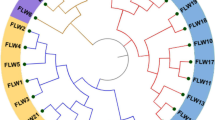
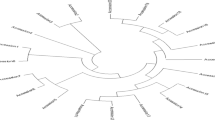
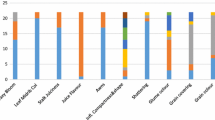
To evaluate genetic diversity in relation to rust and anthracnose disease response, ninety-six accessions were randomly selected from the core collection database of the Germplasm Research Information Network (GRIN) and characterized by a set of 40 SSR markers. The mean value of polymorphism information content (PIC) was 0.8228. Two dendrograms were generated from the molecular genetic data and field morphological data, respectively. The genetic dendrogram demonstrates that the accessions can be classified into three main clades and nine subgroups. The branched subgroups correlated very well with the locations where the accessions were collected. Geographical origin of accessions had significant influences on genetic similarity of sorghum germplasm. Out of 96 accessions, only eight accessions were highly resistant to both rust and anthracnose. All the accessions from South Africa and Mali were highly resistant to anthracnose. The information from genetic classification would be useful for choosing parents to make crosses in sorghum breeding programs and classifying sorghum accessions in germplasm management.
Similar content being viewed by others
References
Anderson, J.A., G.A. Churchill, J.E. Autrique, S.D. Tanksley & M.E. Sorrells, 1993. Optimizing parental selection for genetic linkage maps. Genome 36: 181–186.
Bhattramakki, D., J. Dong, A.K. Chhabra & G.E. Hart, 2000. An integrated SSR and RFLP linkage map of Sorghum bicolor (L.) Moench. Genome 43: 988–1002.
Boora, K.S., R. Frederiksen & C. Magill, 1998. DNA-based markers for a recessive gene conferring anthracnose resistance in sorghum. Crop Sci. 38: 1708–1709.
Botstein, D., R. White, M. Skolnick & R. Davis, 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32: 314–331.
Brueggeman, R., N. Rostoks, D. Kudrna, A. Kilian, F. Han, J. Chen, A. Druka & B. Steffenson, 2002. The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc Natl Acad Sci USA 99: 9328–9333.
Casela, C.R., R.A. Frederiksen & A.S. Fereira, 1993. Evidence for dilatory resistance to anthracnose in sorghum. Plant Dis 77: 908–911.
Collins, N., J. Drake, M. Ayliffe, Q. Sun, J. Ellis, S. Hulbert & T. Pryor, 1999. Molecular characterization of the maize Rp1-D rust resistance haplotype and its mutants. Plant Cell 11: 1365–1376.
Collins, N.C., C.A. Webb, S. Seah, J.G. Ellis, S.H. Hulbert & A. Pryor, 1998. The isolation and mapping of disease resistance gene analogs in maize. Mol Plant-Microbe Interact 11: 968–978.
Dahlberg, J.A., X. Zhang, G.E. Hart & J.E. Mullet, 2002. Comparative assessment of variation among sorghum germplasm accessions using seed morphology and RAPD measurements. Crop Sci 42: 291–296.
Dean, R.E., J.A. Dahlberg, M.S. Hopkins, S.E. Mitchell & S. Kresovich, 1999. Genetic redundancy and diversity among ‘orange’ accessions in the U.S. national sorghum collection as assessed with simple sequence repeat (SSR) markers. Crop Sci 39: 1215–1221.
Deu, M., D. Gonzales-de-Leon, J.-C. Glaszmann, I. Degremont, J. Chantereau, C. Lanaud & P. Hanon, 1994. RFLP diversity in cultivated sorghum in relation to racial differentiation. Theor Appl Genet 88: 834–844.
Feuillet, C., S. Travella, N. Stein, L. Albar, A. Nublat & B. Keller, 2003. Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc Natl Acad Sci USA 100: 15253–15258.
Felsenstein, J., 2004. PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle.
Frederiksen, R.A., 2000. Diseases and disease management in sorghum. In: C.W. Smith & R.A. Frederiksen (Eds.), Sorghum origin, history, technology, and production, pp. 497–533. John Wiley & Sons, Inc., New York.
Huang, L., S.A. Brooks, W. Li, J.P. Fellers, H.N. Trick and B.S. Gill, 2003. Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploidy genome of bread wheat. Genetics 164: 655–664.
Minch, E., A. Ruiz-Linares, D. Goldstein, M. Feldman & L.L. Cavalli-Sforza, 1995, 1996. Microsat (version 1.5b): A computer program for calculation various statistics on microsatellite allele data. WWW: http://lotka.stanford.edu/microsat.html.
Morden, C.W., J. Doebley & Schertz K. F. 1989. Allozyme variation in old world races of Sorghum bicolor (Poaceae). Am J Bot 76: 247–255.
Oliveira, A.C., T. Richter & J.L. Bennetzen, 1996. Regional and racial specificities in sorghum germplasm assessed with DNA markers. Genome 39: 579–587.
Page, R.D.M., 1996. TREEVIEW: An application to display phylogenetic trees on personal computers. Computer Applications in Biosciences 12: 357–358.
Panday, S., A. Sindhu & K.S. Boora, 2002. RAPD based DNA markers linked to anthracnose disease resistance in Sorghum bicolor (L.) Moench. Indian J Exp Biol 40: 206–211.
Rana, B.S., D.P. Tripathi & N.G. Rao, 1976. Genetic analysis of some exotic x Indian crosses in sorghum. XV. Inheritance of resistance to sorghum rust. Indian J Genet Plant Breed 36: 244–249.
Reichardt, M. & S. Rogers, 1997. Preparation of genomic DNA from plant tissue. In: F.M. Ausubel, R. Brent, R.E. Kingston, D.D. Moore, J.G. Seidman, J.A. Smith & K. Struhl (Eds.), Current protocols in molecular biology, pp 2.3.3–7. John Wiley & Sons, Inc., New York.
Schlötterer, C., 2004. The evolution of molecular markers – just a matter of fashion? Nature Rev Genet 5: 63–69.
Schloss, S.J., S.E. Mitchell, G.M. White, R. Kukatla, J.E. Browers, A.H. Paterson & S. Kresovich, 2002. Characterization of RFLP probe sequences for genetic discovery and SSR development in Sorghum bicolor (L.) moench. Theor Appl Genet 105: 912–920.
Shinde, D., Y. Lai, F. Sun & N. Arnheim, 2003. Taq DNA polymerase slippage mutation rates measured by PCR and quasi-likelihood analysis: (CA/GT)n and (A/T)n microsatellites. Nucleic Acids Res 31: 974–980.
Thomas, M.D., I. Sissoko & M. Sacko, 1996. Development of leaf anthracnose and its effect on yield and grain weight of sorghum in West Africa. Plant Dis 79: 151–153.
Tao, Y.Z., D.R. Jordan, R.G. Henzell & C.L. McIntyre, 1998. Identification of genomic regions for rust resistance in sorghum. Euphytica 103: 287–292.
Tao, Y., J. M. Manners, M.M. Ludlow & R.G. Henzell, 1993. DNA polymorphisms in grain sorghum (Sorghum bicolor (L.) Moench). Theor Appl Genet 86: 679–688.
Yang, W., A.C. de Oliveira, I. Godwin, K. Schertz & J.L. Bennetzen, 1996. Comparison of DNA marker technologies in characterizing plant genome diversity: Variability in Chinese sorghums. Crop Sci 36: 1669–1676.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, M.L., Dean, R., Erpelding, J. et al. Molecular genetic evaluation of sorghum germplasm differing in response to fungal diseases: Rust (Puccinia purpurea) and anthracnose (Collectotrichum graminicola). Euphytica 148, 319–330 (2006). https://doi.org/10.1007/s10681-005-9040-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-005-9040-0