Abstract
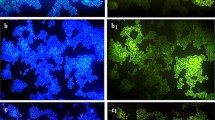
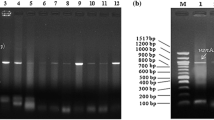
The glycopeptide antibiotics teicoplanin and vancomycin are common to treat severe Gram-positive bacterial infections. The gene vanA confers high-level resistance to these antibiotics, and these phenomena have been shown to be transferable. Release of vancomycin- and teicoplanin-resistant bacteria to surface waters is, therefore, of particular concern since they might proliferate and spread in different environments. Monitoring of the fate of vanA gene in these waters provides information on the exposure and potential threats of those bacteria for the environment and public health. Therefore, this study aimed at preparing a 25-mer-oligonucleotide DNA probe based on the 909 bp BamHI-ClaI fragment from Enterococcus faecium plasmids pVEF1 and pVEF2 through the use of Vector NTI Express Software. The newly designed vanA probe displayed highly specific hybridization with vanA-positive Enterococcus faecalis tested at 46 °C, 55 % formamide, and 0.020 M NaCl stringency conditions. In situ fluorescein hybridizations under the same stringency conditions were also used to monitor river water samples by using fluorescein microscopy. The results showed that the vanA-targeted oligonucleotide DNA probe prepared was not only highly specific but also quantitative tool for monitoring vancomycin- and teicoplanin-resistant bacteria in surface waters.



Similar content being viewed by others
References
Amann, R., Fuchs, B. M., & Behrens, S. (2001). The identification of microorganisms by fluorescence in situ hybridisation. Current Opinion in Biotechnology, 12, 231–236.
Amann, R. I., Ludwig, W., & Schleifer, K. H. (1995). Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiology Reviews, 59, 143–169.
Arthur, M., Molinas, C., Depardieu, F., & Courvalin, P. (1993). Characterization of Tn1546, a Tn3-related transposon conferring glycopeptide resistance by synthesis of depsipeptide peptidoglycan precursors in Enterococcus faecium BM4147. Journal of Bacteriology, 175, 117–127.
Arthur, M., Reynolds, P., & Courvalin, P. (1996). Glycopeptide resistance in enterococci. Trends in Microbiology, 4, 401–407.
Benson, D. A., & Karsch-Mizrachi, I. (2000). GenBank. Nucleic Acids Research, 28, 15–18.
Courvalin, P. (2006). Vancomycin resistance in gram-positive cocci. Clinical Infectious Diseases, 42, 25–34.
Daims, H., Brühl, A., Amann, R., Schleifer, K. H., & Wagner, M. (1999). The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Systematic and Applied Microbiology, 22, 434–444.
Daims, H., Stoecker, K., & Wagner, M. (2005). Fluorescence in situ hybridization for the detection of prokaryotes. In A. M. Osborn & C. J. Smith (Eds.), Advanced methods in molecular microbial ecology (pp. 213–239). Abingdon, UK: Bios-Garland.
Daria, M., Maren, W., Mashal, A., Martin, Z., Andrea, V., Michael, Z., & Hilke, W. (2010). Monitoring of the microbial community composition in saline aquifers during CO2 storage by fluorescence in situ hybridisation. International Journal of Greenhouse Gas Control, 4, 981–989.
Eilers, H., Pernthaler, J., Glöckner, F. O., & Amann, R. (2000). Culturability and in situ abundance of pelagic bacteria from the North Sea. Applied and Environmental Microbiology, 66, 3044–3051.
Findlay, S. E. G., Sinsabaugh, R. L., Sobczak, W. V., & Hoostal, M. (2003). Metabolic and structural response of hyporheic microbial communities to variations in supply of dissolved organic matter. Limnology and Oceanography, 48, 1608–1617.
Glöckner, F. O., Amann, R., Alfreider, A., Pemthaler, J., Psewner, R., Trebesius, K., & Schleifer, K. H. (1996). An in situ hybridization protocol for detection and identification of planktonic bacteria. Systematic and Applied Microbiology, 19, 403–406.
Glöckner, F. O., Zaichikov, E., Belkova, N., Denissowa, L., Pernthaler, J., Pernthaler, A., & Amann, R. (2000). Comparative 16S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria. Applied and Environmental Microbiology, 66, 5053–5065.
Handwerger, S., & Skoble, J. (1995). Identification of chromosomal mobile element conferring high-level vancomycin resistance in Enterococcus faecium. Antimicrobial Agents and Chemotherapy, 39, 2446–2453.
Icgen, B., & Yilmaz, F. (2016). Design a cadA-targeted DNA probe for screening of potential bacterial cadmium biosorbents. Environmental Science and Pollution Research, 23, 5743–5752.
Kalmbach, S., Manz, W., & Szewzyk, U. (1997). Isolation of new bacterial species from drinking water biofilms and proof of their in situ dominance with highly specific 16S rRNA probes. Applied and Environmental Microbiology, 63, 4164–4170.
Kenzaka, T., Yamaguchi, N., Tani, K., & Nasu, M. (1998). rRNA-targeted fluorescent in situ hybridization analysis of bacterial community structure in river water. Microbiology, 144, 2085–2093.
Klammer, S., Posh, T., Sonntag, B., Griebler, C., Mindl, B., & Psenner, R. (2002). Dynamics of bacterial abundance, biomass, activity, and community composition in the oligotrophic Traunsee and the Traun River Austria. Water, Air and Soil Pollution, 2, 137–163.
Kümmerer, K. (2004). Resistance in the environment. Journal of Antimicrobial Chemotherapy, 54, 311–320.
Levsky, J. M., & Singer, R. H. (2003). Fluorescence in situ hybridization: past, present and future. Journal of Cell Science, 116, 2833–2838.
Li, B., Irvin, S., & Baker, B. (2007). The variation of nitrifying bacterial population sizes in a sequencing batch reactor (SBR) treating low, mid, high concentrated synthetic wastewater. Journal of Environmental Engineering and Science, 6, 651–663.
Loy, A., Horn, M., & Wagner, M. (2003). ProbeBase: an online resource for rRNA-targeted oligonucleotide probes. Nucleic Acids Research, 31, 514–516.
Murray, B. E., & Nannini, E. C. (2009). Glycopeptides (vancomycin and teicoplanin), streptogramins (quinupristin-dalfopristin), and lipopeptides (daptomycin). In G. L. Mandell, J. E. Bennett, & R. Dolin (Eds.), Mandell, Douglas, and Bennett’s principles and practice of infectious diseases (7th ed., pp. 449–465). Philadelphia, PA: Elsevier Churchill Livingston.
Nielsen, P., Daims, H., & Lemmer, H. (2009). FISH handbook for biological wastewater treatment: identification and quantification of microorganisms in activated sludge and biofilms by FISH, London. New York: IWA Publishing.
Pernthaler, J., Alfreider, A., Posch, T., Andreatta, S., & Psenner, R. (1997). In situ classification and image cytometry of pelagic bacteria from a high mountain lake (Gossenköllesee, Austria). Applied and Environmental Microbiology, 63, 4778–4783.
Pernthaler, J., Glöckner, F. O., Unterholzner, S., Alfreider, A., Psenner, R., & Amann, R. (1998). Seasonal community and population dynamics of pelagic bacteria and Archaea in a high mountain lake. Applied and Environmental Microbiology, 64, 4299–4306.
Pinhassi, J., & Hagström, A. (2000). Seasonal succession in marine bacterioplankton. Aquatic Microbial Ecology, 21, 245–256.
Rice, L. B. (2001). Emergence of vancomycin-resistant enterococci. Emerging Infectious Diseases, 7, 183–187.
Sletvold, H., Johnsen, P. J., Simonsen, G. S., Aasnaes, B., Sundsfjord, A., & Nielsen, K. M. (2007). Comparative DNA analysis of two vanA plasmids from Enterococcus faecium strains isolated from poultry and a poultry farmer in Norway. Antimicrobial Agents and Chemotherapy, 51, 736–739.
Sletvold, H., Johnsen, P. J., Wikmark, O. G., Simonsen, G. S., Sundsfjord, A., & Nielsen, K. M. (2010). Tn1546 is part of a larger plasmid-encoded genetic unit horizontally disseminated among clonal Enterococcus faecium lineages. Journal of Antimicrobial Chemotherapy, 65, 1894–1906.
Vergis, E. N., Hayden, M. K., Chow, J. W., Snydman, D. R., Zervos, M. J., Linden, P. K., Wagener, M. M., Schmitt, B., & Muder, R. R. (2001). Determinants of vancomycin resistance and mortality rates in enterococcal bacteremia. A prospective multicenter study. Annals of Internal Medicine, 135, 484–492.
Volpi, E. V., & Bridger, J. M. (2008). FISH glossary: an overview of the fluorescence in situ hybridization technique. BioTechniques, 45, 385–409.
Wagner, M., Amann, R., Lemmer, H., & Schleifer, K. H. (1993). Probing activated sludge with oligonucleotides specific for proteobacteria: inadequacy of culture-dependent methods for describing microbial community structure. Applied and Environmental Microbiology, 59, 1520–1525.
Acknowledgments
The authors are thankful for the support given by TUBITAK-Turkey through the project 114Z973.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nakipoglu, M., Yilmaz, F. & Icgen, B. vanA-targeted oligonucleotide DNA probe designed to monitor vancomycin- and teicoplanin-resistant bacteria in surface waters. Environ Monit Assess 188, 569 (2016). https://doi.org/10.1007/s10661-016-5578-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-016-5578-7