Abstract
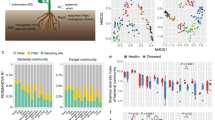
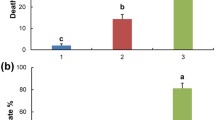
Goji berry (Lycium barbarum) is an important cash crop in China. However, in recent years its yield has been threatened by root rot caused by fungal pathogens. The infection significantly alters the structure and function of the plant microbiome. Thus, knowledge of the responses of the plant microbiome is a prerequisite for the bio-control of fungal pathogens. In the present study, the microbial communities of healthy, diseased, and wild goji berries were studied by amplicon sequencing. We found different responses of the microbial communities in different goji berry organs to the fungal pathogen. Moreover, higher microbial community abundance and diversity in the wild goji berry plant parts were observed, implying more complex and diverse ecological functions. At the same time, some potential beneficial bacteria were enriched in the diseased goji berry roots, consistent with the ‘cry to help’ hypothesis in plant roots. Moreover, intra-kingdom co-occurrence network analysis revealed higher positive correlations in bacterial than in fungal networks, implying an unstable bacterial network. In the inter-kingdom networks, most of the hubs (with high degree and closeness centrality values) were bacterial, indicating the important role of bacteria in the interaction under stress. In addition, of the 54 fungal strains isolated from the diseased goji berry roots, 42.6% were identified as Fusarium spp.; hence, it is a potential pathogen for subsequent research on the control of root rot in goji berries. The results of this study provide new perspectives on the biological control of root rot in goji berries.




Similar content being viewed by others
Data Availability
The data that support the findings of this study are available on request from the corresponding author, [Liu], upon reasonable request.
References
Amagase, H., & Farnsworth, N. R. (2011). A review of botanical characteristics, phytochemistry, clinical relevance in efficacy and safety of Lycium barbarum fruit (goji). Food Research International, 44(7), 1702–1717. https://doi.org/10.1016/j.foodres.2011.03.027
Arif, I., Batool, M., & Schenk, P. M. (2020). Plant microbiome engineering: Expected benefits for improved crop growth and resilience. Trends in Biotechnology, 38(12), 1385–1396. https://doi.org/10.1016/j.tibtech.2020.04.015
Bakker, P. A. H. M., Pieterse, C. M. J., de Jonge, R., & Berendsen, R. L. (2018). The soil-borne legacy. Cell, 172(6), 1178–1180. https://doi.org/10.1016/j.cell.2018.02.024
Bello, M. G. D., Knight, R., Gilbert, J. A., & Blaser, M. J. (2018). Preserving microbial diversity. Science, 362(6410), 33–34. https://doi.org/10.1126/science.aau8816
Berendsen, R. L., Vismans, G., Yu, K., Song, Y., de Jonge, R., Burgman, W. P., Burmølle, M., Herschend, J., Bakker, P. A. H. M., & Pieterse, C. M. J. (2018). Disease-induced assemblage of a plant-beneficial bacterial consortium. The ISME Journal, 12(6), 1496–1507. https://doi.org/10.1038/s41396-018-0093-1
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., & Holmes, S. P. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods, 13(7), 581–583. https://doi.org/10.1038/nmeth.3869
Carrión, V. J., Perez-Jaramillo, J., Cordovez, V., Tracanna, V., de Hollander, M., Ruiz-Buck, D., Mendes, L. W., van Ijcken, W. F. J., Gomez-Exposito, R., Elsayed, S. S., Mohanraju, P., Arifah, A., van der Oost, J., Paulson, J. N., Mendes, R., van Wezel, G. P., Medema, M. H., & Raaijmakers, J. M. (2019). Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science, 366(6465), 606–612. https://doi.org/10.1126/science.aaw9285
Chang, H. M., But, P. H., Yao, S. C., Wang, L. L., & Yeung, C. S. (1986). Pharmacology and applications of chinese materia medica. World Scientific Book.
Chen, S., Zhou, Y., Chen, Y., & Gu, J. (2018). Fastp: An ultra-fast all-in-one fastq preprocessor. Bioinformatics, 34(17), 884–890. https://doi.org/10.1093/bioinformatics/bty560
Coyte, K. Z., Schluter, J., & Foster, K. R. (2015). The ecology of the microbiome: Networks, competition, and stability. Science, 350(6261), 663–666. https://doi.org/10.1126/science.aad2602
Cregger, M. A., Veach, A. M., Yang, Z. K., Crouch, M. J., Vilgalys, R., Tuskan, G. A., & Schadt, C. W. (2018). The Populus holobiont: dissecting the effects of plant niches and genotype on the microbiome. Microbiome, 6. https://doi.org/10.1186/s40168-018-0413-8
de la Lastra, E., Marin-Guirao, J. I., Lopez-Moreno, F. J., Soriano, T., Cara-Garcia, M., & Capote, N. (2021). Potential inoculum sources of fusarium species involved in asparagus decline syndrome and evaluation of soil disinfestation methods by qPCR protocols. Pest Management Science, 77(10), 4749–4757. https://doi.org/10.1002/ps.6519
de Vries, F. T., & Wallenstein, M. D. (2017). Below-ground connections underlying above-ground food production: A framework for optimising ecological connections in the rhizosphere. Journal of Ecology, 105(4), 913–920. https://doi.org/10.1111/1365-2745.12783
de Vries, F. T., Griffiths, R. I., Knight, C. G., Nicolitch, O., & Williams, A. (2020). Harnessing rhizosphere microbiomes for drought-resilient crop production. Science, 368(6488), 270–274. https://doi.org/10.1126/science.aaz5192
Dixon, P. (2003). VEGAN, a package of R functions for community ecology. Journal of Vegetation Science, 14(6), 927–930. https://doi.org/10.1658/1100-9233(2003)014[0927:Vaporf]2.0.Co;2
Edwards, J., Johnson, C., Santos-Medellín, C., Lurie, E., Podishetty, N. K., Bhatnagar, S., Eisen, J. A., & Sundaresan, V. (2015). Structure, variation, and assembly of the root-associated microbiomes of rice. Proceedings of the National Academy of Sciences, 112(8), E911–E920. https://doi.org/10.1073/pnas.1414592112
Fang, X., Zhang, W., Meng, Q., Gao, J., Wang, X., King, J., Song, C., Dai, S., & Miao, Y. (2007). High-resolution magneto stratigraphy of the Neogene Huaitoutala section in the eastern Qaidam Basin on the NE Tibetan Plateau, Qinghai Province, China and its implication on tectonic uplift of the NE Tibetan Plateau. Earth and Planetary Science Letters, 258(1–2), 293–306. https://doi.org/10.1016/j.epsl.2007.03.042
Fang, T., Lu, H., & Bai, L. (2022). Analysis of pathogenesis to Lycium barbarum root rot on transcriptome sequencin. Journal of Arid Land Resources and Environment, 36(11), 133–140. https://doi.org/10.13448/j.cnki.jalre.2022.286
Feng, K., Peng, X., Zhang, Z., Gu, S., He, Q., Shen, W., Wang, Z., Wang, D., Hu, Q., Li, Y., Wang, S., & Deng, Y. (2022). iNAP: An integrated network analysis pipeline for microbiome studies. iMeta, 1(2), e13. https://doi.org/10.1002/imt2.13
Fernie, A. R., & Yang, J. (2019). De novo domestication: An alternative route toward new crops for the future. Molecular Plant, 12(5), 615–631. https://doi.org/10.1016/j.molp.2019.03.016
Fitzpatrick, C. R., Copeland, J., Wang, P. W., Guttman, D. S., Kotanen, P. M., & Johnson, M. T. J. (2018). Assembly and ecological function of the root microbiome across angiosperm plant species. Proceedings of the National Academy of Sciences of the United States of America, 115(6), E1157–E1165. https://doi.org/10.1073/pnas.1717617115
Friedman, J., & Alm, E. J. (2012). Inferring correlation networks from genomic survey data. Plos Computational Biology, 8(9). https://doi.org/10.1371/journal.pcbi.1002687
Gao, H. H., Jia, C. B., Han, Q., Su, J. Y., & Xu, C. Y. (2022). Microbiological mechanism of root rot of Lycium barbarum Ningqi-7. Biotechnology Bulletin, 38(12), 244–251.
Gao, C., Montoya, L., Xu, L., Madera, M., Hollingsworth, J., Purdom, E., Singan, V., Vogel, J., Hutmacher, R. B., Dahlberg, J. A., Coleman-Derr, D., Lemaux, P. G., & Taylor, J. W. (2020). Fungal community assembly in drought-stressed sorghum shows stochasticity, selection, and universal ecological dynamics. Nature Communications, 11(1). https://doi.org/10.1038/s41467-019-13913-9
Gao, M., Xiong, C., Gao, C., Tsui, C. K. M., Wang, M. M., Zhou, X., Zhang, A. M., & Cai, L. (2021). Disease-induced changes in plant microbiome assembly and functional adaptation. Microbiome, 9(1). https://doi.org/10.1186/s40168-021-01138-2
Gardes, M., & Bruns, T. D. (1993). ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Molecular Ecology, 2(2), 113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Garibaldi, A., Bertetti, D., Pensa, P., Poli, A., & Gullino, M. L. (2011). First report of Fusarium oxysporum causing wilt on jade plant (Crassula ovata) in Italy. Protezione Delle Colture, 11(3), 44–47. https://doi.org/10.1094/PDIS-12-10-0897
Giles, J. M., Tordsen, C. L., Rebstock, T. R., Bucciarelli, B., Samac, D. A., & Sathoff, A. E. (2022). Management strategies and distribution of Aphanomyces root rot of alfalfa (Medicago sativa), a continuing threat to forage production in the United States. Plant Pathology, 71(6), 1231–1240. https://doi.org/10.1111/ppa.13563
Han, L., Zhou, X., Zhao, Y., Wu, L., Ping, X., He, Y., Peng, S., He, X., & Du, Y. (2020). First report of Plectosphaerella plurivora causing root rot disease in Panax notoginseng in China. Journal of Phytopathology, 168(7–8), 375–379. https://doi.org/10.1111/jph.12901
Hassan, M. K., McInroy, J. A., & Kloepper, J. W. (2019). The interactions of rhizodeposits with plant growth-promoting rhizobacteria in the rhizosphere: a review. Agriculture-Basel, 9(7). https://doi.org/10.3390/agriculture9070142
Jia, C., Guo, Y., Ma, C., Su, J., & Xu, C. (2021). Difference on soil microbial community and function of healthy and diseased plants of Lycium barbarum ningqi-1. Ecology and Environmental Sciences, 30(9), 1831–1841.
Kuiper, J. J., van Altena, C., de Ruiter, P. C., van Gerven, L. P. A., Janse, J. H., & Mooij, W. M. (2015). Food-web stability signals critical transitions in temperate shallow lakes. Nature Communications, 6. https://doi.org/10.1038/ncomms8727
Laforest-Lapointe, I., Paquette, A., Messier, C., & Kembel, S. W. (2017). Leaf bacterial diversity mediates plant diversity and ecosystem function relationships. Nature, 546(7656), 145–147. https://doi.org/10.1038/nature22399
Lee, S. M., Kong, H. G., Song, G. C., & Ryu, C. M. (2021). Disruption of Firmicutes and Actinobacteria abundance in tomato rhizosphere causes the incidence of bacterial wilt disease. The ISME Journal, 15(1), 330–347. https://doi.org/10.1038/s41396-020-00785-x
Li, J., Pan, L., Naman, C. B., Deng, Y., Chai, H., Keller, W. J., & Kinghorn, A. D. (2014). Pyrrole alkaloids with potential cancer chemopreventive activity isolated from a Goji berry-contaminated commercial sample of African mango. Journal of Agricultural and Food Chemistry, 62(22), 5054–5060. https://doi.org/10.1021/jf500802x
Li, A XZ, & LC, B. (2021). Isolation and identification of pathogen oflycium barbarum root rot disease in nuomuhong farm. Molecular Plant Breeding, 19(4), 1232-1236. https://doi.org/10.13271/j.mpb.019.001232
Li, C., Zhang, M., Li, J., Huang, M., Shao, X., & Yang, Z. (2022). Fusarium redolens causes black rot disease in Gastrodia elata grown in China. Crop Protection, 155. https://doi.org/10.1016/j.cropro.2022.105933
Lindow, S. E., & Brandl, M. T. (2003). Microbiology of the phyllosphere. Applied and Environmental Microbiology, 69(4), 1875–1883. https://doi.org/10.1128/aem.69.4.1875-1883.2003
Liu, H., & Brettell, L. E. (2019). Plant defense by voc-induced microbial priming. Trends in Plant Science, 24(3), 187–189. https://doi.org/10.1016/j.tplants.2019.01.008
Liu, D., Zhao, Q., Cui, X., Chen, R., Li, X., Qiu, B., & Ge, F. (2019). A transcriptome analysis uncovers Panax notoginseng resistance to Fusarium solani induced by methyl jasmonate. Genes & Genomics, 41(12), 1383–1396. https://doi.org/10.1007/s13258-019-00865-z
Liu, H., Li, J., Carvalhais, L. C., Percy, C. D., Prakash Verma, J., Schenk, P. M., & Singh, B. K. (2021). Evidence for the plant recruitment of beneficial microbes to suppress soil-borne pathogens. New Phytologist, 229(5), 2873–2885. https://doi.org/10.1111/nph.17057
Liu, L., & ÖZsu, M. T. (2009).Plots for Qualitative Information. Springerhttps://doi.org/10.1007/978-0-387-39940-9_3304
Liu, J., Yu, X., Qin, Q., Dinkins, R. D., & Zhu, H. (2020). The impacts of domestication and breeding on nitrogen fixation symbiosis in legumes. Frontiers in Genetics, 11. https://doi.org/10.3389/fgene.2020.00973
Nilsson, R. H., Larsson, K.-H., Taylor, A. F. S., Bengtsson-Palme, J., Jeppesen, T. S., Schigel, D., Kennedy, P., Picard, K., Glöckner, F. O., Tedersoo, L., Saar, I., Kõljalg, U., & Abarenkov, K. (2018). The UNITE database for molecular identification of fungi: Handling dark taxa and parallel taxonomic classifications. Nucleic Acids Research, 47(D1), D259–D264. https://doi.org/10.1093/nar/gky1022
Pang, Z., Mao, X., Xia, Y., Xiao, J., Wang, X., Xu, P., & Liu, G. (2022). Multiomics reveals the effect of root rot on polygonati rhizome and identifies pathogens and biocontrol strain. Microbiology Spectrum, 10(2). https://doi.org/10.1128/spectrum.02385-21
Parks, D. H., Tyson, G. W., Hugenholtz, P., & Beiko, R. G. (2014). STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics, 30(21), 3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Porter, S. S., & Sachs, J. L. (2020). Agriculture and the disruption of plant-microbial symbiosis. Trends in Ecology & Evolution, 35(5), 426–439. https://doi.org/10.1016/j.tree.2020.01.006
Potterat, O. (2010). Goji (Lycium barbarum and L-chinense): Phytochemistry, pharmacology and safety in the perspective of traditional uses and recent popularity. Planta Medica, 76(1), 7–19. https://doi.org/10.1055/s-0029-1186218
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., & Glöckner, F. O. (2012). The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Research, 41(D1), D590–D596. https://doi.org/10.1093/nar/gks1219
Raaijmakers, J. M., & Kiers, E. T. (2022). Rewilding plant microbiomes Microbiota of crop ancestors may offer a way to enhance sustainable food production. Science, 378(6620), 599–600. https://doi.org/10.1126/science.abn6350
Raimondo, M. L., & Carlucci, A. (2018). Characterization and pathogenicity assessment of Plectosphaerella species associated with stunting disease on tomato and pepper crops in Italy. Plant Pathology, 67(3), 626–641. https://doi.org/10.1111/ppa.12766
Reese, A. T., Chadaideh, K. S., Diggins, C. E., Schell, L. D., Beckel, M., Callahan, P., Ryan, R., Thompson, M. E., & Carmody, R. N. (2021). Effects of domestication on the gut microbiota parallel those of human industrialization. Elife, 10. https://doi.org/10.7554/eLife.60197
Schulz-Bohm, K., Gerards, S., Hundscheid, M., Melenhorst, J., de Boer, W., & Garbeva, P. (2018). Calling from distance: Attraction of soil bacteria by plant root volatiles. The ISME Journal, 12(5), 1252–1262. https://doi.org/10.1038/s41396-017-0035-3
Sun, D. L., Jiang, X., Wu, Q. L., & Zhou, N. Y. (2013). Intragenomic heterogeneity of 16s rRNA genes causes overestimation of prokaryotic diversity. Applied and Environmental Microbiology, 79(19), 5962–5969. https://doi.org/10.1128/aem.01282-13
de Vries, F. T., Griffiths, R. I., Bailey, M., Craig, H., Girlanda, M., Gweon, H. S., Hallin, S., Kaisermann, A., Keith, A. M., Kretzschmar, M., Lemanceau, P., Lumini, E., Mason, K. E., Oliver, A., Ostle, N., Prosser, J. I., Thion, C., Thomson, B., & Bardgett, R. D. (2018). Soil bacterial networks are less stable under drought than fungal networks. Nature Communications, 9. https://doi.org/10.1038/s41467-018-05516-7
Wagg, C., Schlaeppi, K., Banerjee, S., Kuramae, E. E., & van der Heijden, M. G. A. (2019). Fungal-bacterial diversity and microbiome complexity predict ecosystem functioning. Nature Communications, 10. https://doi.org/10.1038/s41467-019-12798-y
Wang X, Sun J Y, GE S H, & H, Z. L. (2015). Analysis of the influence of meteorological conditions on the growth and development of rice in Xuzhou.Anhui Agricultural Sciences, 43(01), 135–136. https://doi.org/10.13989/j.cnki.0517-6611.2015.01.096
Ware, I. M., Van Nuland, M. E., Yang, Z. K., Schadt, C. W., Schweitzer, J. A., & Bailey, J. K. (2021). Climate-driven divergence in plant-microbiome interactions generates range-wide variation in bud break phenology. Communications Biology, 4(1). https://doi.org/10.1038/s42003-021-02244-5
Wei, Z., Gu, Y., Friman, V.-P., Kowalchuk, G. A., Xu, Y., Shen, Q., & Jousset, A. (2019). Initial soil microbiome composition and functioning predetermine future plant health. Science Advances, 5(9). https://doi.org/10.1126/sciadv.aaw0759
Wen, T., Yuan, J., He, X., Lin, Y., Huang, Q., & Shen, Q. (2020). Enrichment of beneficial cucumber rhizosphere microbes mediated by organic acid secretion. Horticulture Research, 7(1). https://doi.org/10.1038/s41438-020-00380-3
White, T., Bruns, T., Lee, S., Taylor, J., Innis, M., Gelfand, D., & Sninsky, J. (1990). Amplification and direct sequencing of fungal ribosomal Rna genes for phylogenetics. Academic Press.
Xing, R., Gao, Q. B., Zhang, F. Q., Wang, J. L., & Chen, S. L. (2019). Large-scale distribution of bacterial communities in the Qaidam Basin of the Qinghai-Tibet Plateau. Microbiologyopen, 8(10). https://doi.org/10.1002/mbo3.909
Xing, R., Gao, Q. B., Zhang, F. Q., Wang, J. L., & Chen, S. L. (2021). Environmental filtering affects fungal communities more than dispersal limitation in a high-elevation hyperarid basin on Qinghai-Tibet Plateau. FEMS Microbiol Lett, 368(6). https://doi.org/10.1093/femsle/fnab033
Xiong, L., Deng, N., Zheng, B., Li, T., & Liu, R. H. (2021). Goji berry (Lycium spp.) extracts exhibit antiproliferative activity via modulating cell cycle arrest, cell apoptosis, and the p53 signaling pathway. Food & Function, 12(14), 6513–6525. https://doi.org/10.1039/d1fo01105g
Yuan, J., Zhao, J., Wen, T., Zhao, M., Li, R., Goossens, P., Huang, Q., Bai, Y., Vivanco, J. M., Kowalchuk, G. A., Berendsen, R. L., & Shen, Q. (2018). Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome, 6. https://doi.org/10.1186/s40168-018-0537-x
Acknowledgements
This study received Special Fund for Qilian Mountain National Park (QHTX-2020-004); Long-Term National Scientifc Research Base of Qilian Mountain National Park, Xining 810000, Qinghai, China; financial support from the Sanjiangyuan National Park Joint Program (LHZX-2020-02-01), Construction Project for Innovation Platform of Qinghai province (2022-ZJ-Y04) and Central Aisan Drug Discovery and Development Center of Chinese Academy of Sciences (CAM202102).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the study was conducted in the absence of any commercial or financial relationships that could be constructed as a potential conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Z., Xiao, Y., Li, N. et al. Effects of root rot on microbial communities associated with goji berry (Lycium barbarum) in the Qaidam Basin, China. Eur J Plant Pathol 167, 853–866 (2023). https://doi.org/10.1007/s10658-023-02723-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02723-x