Abstract
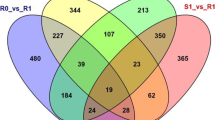
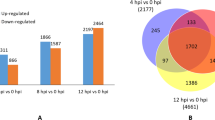
Plasmopara viticola is a plant pathogenic oomycete that causes downy mildew in grapevines. Shine Muscat, a hybrid grapevine cultivar of Vitis vinifera and V. labrascana produced in Japan, exhibits resistance to P. viticola. Transcriptome analyses were performed to elucidate the mechanism of defense against P. viticola infection in Shine Muscat. Using two sets of RNA seq data and V. vinifera genome information, transcriptome analyses reproducibly identified 269 and 55 grapevine genes that were upregulated and downregulated, respectively, in response to P. viticola infection. Gene ontology analysis of the differentially expressed genes suggested that Shine Muscat recognizes P. viticola invasion at the cell membrane, with subsequent transmission of signals into the cells to initiate a response. Furthermore, P. viticola–responsive paralogous genes were distributed among 8 clusters located on chromosomes 2, 5, 6, 9, 12, and 16 in the V. vinifera genome. These gene clusters encode PR5, PR10, dirigent protein, pleiotropic drug resistance protein, an uncharacterized protein, phenylalanine ammonia-lyase, ethylene-responsive transcription factor, and stilbene synthase, respectively. The gene cluster encoding ethylene-responsive transcription factor was downregulated by P. viticola infection, whereas the other 7 gene clusters were upregulated. With the exception of the gene cluster encoding the uncharacterized protein, the genes were associated with plant defense mechanisms, suggesting that synchronized transcription of these gene clusters plays an important role in the grapevine defense mechanism against P. viticola. This is the first study to identify multiple infection-responsive gene clusters in the grapevine genome.




Similar content being viewed by others
References
Ali, S., Ganai, B. A., Kamili, A. N., Bhat, A. A., Mir, Z. A., Bhat, J. A., Tyagi, A., Islam, S. T., Mushtaq, M., Yadav, P., Rawat, S., & Grover, A. (2018). Pathogenesis-related proteins and peptides as promising tools for engineering plants with multiple stress tolerance. Microbiological Research, 212–213, 29–37.
Çakır, B., & Kılıçkaya, O. (2013). Whole-genome survey of the putative ATP-binding cassette transporter family genes in Vitis vinifera. PLoS One, 8, e78860.
Chitarra, W., Perrone, I., Avanzato, C. G., Minio, A., Boccacci, P., Santini, D., Gilardi, G., Siciliano, I., Gullino, M. L., Delledonne, M., Mannini, F., & Gambino, G. (2017). Grapevine grafting: scion transcript profiling and defense-related metabolites induced by rootstocks. Frontiers in Plant Science, 8, 654.
De Miccolis Angelini, R. M., Rotolo, C., Gerin, D., Abate, D., Pollastro, S., & Faretra, F. (2019). Global transcriptome analysis and differentially expressed genes in grapevine after application of the yeast-derived defense inducer cerevisane. Pest Management Science, 75, 2020–2033.
Dufour, M. C., Magnin, N., Dumas, B., Vergnes, S., & Corio-Costet, M. F. (2016). High-throughput gene-expression quantification of grapevine defense responses in the field using microfluidic dynamic arrays. BMC Genomics, 17, 957.
Eisenmann, B., Czemmel, S., Ziegler, T., Buchholz, G., Kortekamp, A., Trapp, O., Rausch, T., Dry, I., & Bogs, J. (2019). Rpv3-1 mediated resistance to grapevine downy mildew is associated with specific host transcriptional responses and the accumulation of stilbenes. BMC Plant Biology, 19, 343.
Gauthier, A., Trouvelot, S., Kelloniemi, J., Frettinger, P., Wendehenne, D., Daire, X., Joubert, J. M., Ferrarini, A., Delledonne, M., Flors, V., & Poinssot, B. (2014). The sulfated laminarin triggers a stress transcriptome before priming the SA- and ROS-dependent defenses during grapevine's induced resistance against Plasmopara viticola. PLoS One, 9, e88145.
Gessler, C., Pertot, I., & Perazzolli, M. (2011). Plasmopara viticola: a review of knowledge on downy mildew of grapevine and effective disease management. Phytopathologia Mediterranea, 50, 3–44.
Giannuzzi, G., D'Addabbo, P., Gasparro, M., Martinelli, M., Carelli, F. N., Antonacci, D., & Ventura, M. (2011). Analysis of high-identity segmental duplications in the grapevine genome. BMC genomics, 12, 436.
Hasan, M., & Bae, H. (2017). An overview of stress-induced resveratrol synthesis in grapes: perspectives for resveratrol-enriched grape products. Molecules, 22, 294.
He, R., Wu, J., Zhang, Y., Agüero, C. B., Li, X., Liu, S., Wang, C., Walker, M. A., & Lu, J. (2017). Overexpression of a thaumatin-like protein gene from Vitis amurensis improves downy mildew resistance in Vitis vinifera grapevine. Protoplasma, 254, 1579–1589.
Jaillon, O., Aury, J. M., Noel, B., Policriti, A., Clepet, C., Casagrande, A., Choisne, N., Aubourg, S., Vitulo, N., Jubin, C., Vezzi, A., Legeai, F., Hugueney, P., Dasilva, C., Horner, D., Mica, E., Jublot, D., Poulain, J., Bruy`ere, C., Billault, A., Segurens, B., Gouyvenoux, M., Ugarte, E., Cattonaro, F., Anthouard, V., Vico, V., Fabbro, C. D., Alaux, M., Gaspero, G. D., Dumas, V., Felice, N., Paillard, S., Juman, I., Moroldo, M., Scalabrin, S., Canaguier, A., Clainche, I. L., Malacrida, G., Durand, E., Pesole, G., Laucou, V., Chatelet, P., Merdinoglu, D., Delledonne, M., Pezzotti, M., Lecharny, A., Scarpelli, C., Artiguenave, F., P`e, M. E., Valle, G., Morgante, M., Caboche, M., Adam-Blondon, A. F., Weissenbach, J., Qu´etier, F., & Wincker, P. (2007). The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature, 449, 463–467.
Khalil-Ur-Rehman, M., Sun, L., Li, C., Faheem, M., Wang, W., & Tao, J. (2017a). Comparative RNA-seq based transcriptomic analysis of bud dormancy in grape. BMC Plant Biology, 17, 18.
Khalil-Ur-Rehman, M., Wang, W., Xu, Y., Haider, M., Li, C., & Tao, J. (2017b). Comparative study on reagents involved in grape bud break and their effects on different metabolites and related gene expression during winter. Frontiers in Plant Science, 8, 1340.
Kono, A., Ban, Y., Mitani, N., Fujii, H., Sato, S., Suzaki, K., Azuma, A., Onoue, N., & Sato, A. (2018). Development of SSR markers linked to QTL reducing leaf hair density and grapevine downy mildew resistance in Vitis vinifera. Molecular Breeding, 38, 138.
Lebel, S., Schellenbaum, P., Walter, B., & Maillot, P. (2010). Characterisation of the Vitis vinifera PR10 multigene family. BMC Plant Biology, 10, 184.
Legay, G., Marouf, E., Berger, D., Neuhaus, J. M., Mauch-Mani, B., & Slaughter, A. (2011). Identification of genes expressed during the compatible interaction of grapevine with Plasmopara viticola through suppression subtractive hybridization (SSH). European Journal of Plant Pathology, 129, 281–301.
Li, X., Wu, J., Yin, L., Zhang, Y., Qu, J., & Lu, J. (2015). Comparative transcriptome analysis reveals defense-related genes and pathways against downy mildew in Vitis amurensis grapevine. Plant Physiology and Biochemistry, 95, 1–14.
Liu, R., Weng, K., Dou, M., Chen, T., Yin, X., Li, Z., et al. (2019). Transcriptomic analysis of Chinese wild Vitis pseudoreticulata in response to Plasmopara viticola. Protoplasma. https://doi.org/10.1007/s00709-019-01387-x
Malacarne, G., Vrhovsek, U., Zulini, L., Cestaro, A., Stefanini, M., Mattivi, F., Delledonne, M., Velasco, R., & Moser, C. (2011). Resistance to Plasmopara viticola in a grapevine segregating population is associated with stilbenoid accumulation and with specific host transcriptional responses. BMC Plant Biology, 11, 114.
Monaghan, J., & Zipfel, C. (2012). Plant pattern recognition receptor complexes at the plasma membrane. Current Opinion in Plant Biology, 15, 349–357.
Paniagua, C., Bilkova, A., Jackson, P., Dabravolski, S., Riber, W., Didi, V., Houser, J., Gigli-Bisceglia, N., Wimmerova, M., Budínská, E., Hamann, T., & Hejatko, J. (2017). Dirigent proteins in plants: modulating cell wall metabolism during abiotic and biotic stress exposure. Journal of Experimental Botany, 68, 3287–3301.
Parage, C., Tavares, R., Réty, S., Baltenweck-Guyot, R., Poutaraud, A., Renault, L., Heintz, D., Lugan, R., Marais, G. A., Aubourg, S., & Hugueney, P. (2012). Structural, functional, and evolutionary analysis of the unusually large stilbene synthase gene family in grapevine. Plant Physiology, 160, 1407–1419.
Perazzolli, M., Moretto, M., Fontana, P., Ferrarini, A., Velasco, R., Moser, C., et al. (2012). Downy mildew resistance induced by Trichoderma harzianum T39 in susceptible grapevines partially mimics transcriptional changes of resistant genotypes. BMC Genomics, 13, 660.
Polesani, M., Bortesi, L., Ferrarini, A., Zamboni, A., Fasoli, M., Zadra, C., Lovato, A., Pezzotti, M., Delledonne, M., & Polverari, A. (2010). General and species-specific transcriptional responses to downy mildew infection in a susceptible (Vitis vinifera) and a resistant (V. riparia) grapevine species. BMC genomics, 11, 117.
Polesani, M., Desario, F., Ferrarini, A., Zamboni, A., Pezzotti, M., Kortekamp, A., & Polverari, A. (2008). cDNA-AFLP analysis of plant and pathogen genes expressed in grapevine infected with Plasmopara viticola. BMC Genomics, 9, 142.
Rienth, M., Crovadore, J., Ghaffari, S., & Lefort, F. (2019). Oregano essential oil vapour prevents Plasmopara viticola infection in grapevine (Vitis Vinifera) and primes plant immunity mechanisms. PLoS ONE, 14, e0222854.
Shimizu, T., Kono, A., & Suzaki, K. (2019). Transcriptional analysis of defense-related genes induced by infection with the causal agent of downy mildew, Plasmopara viticola, in grapevine cultivar Shine Muscat. Journal of General Plant Pathology, 85, 182–188.
Shirasawa, K., Azuma, A., Taniguchi, F., Yamamoto, T., Sato, A., Hirakawa, H., & Isobe, S. (2019). De novo whole-genome assembly in interspecific hybrid table grape, ‘Shine Muscat’. bioRxiv, 730762; https://doi.org/10.1101/730762.
Suehiro, Y., Mochida, K., Itamura, H., & Esumi, T. (2014). Skin browning and expression of PPO, STS, and CHS genes in the grape berries of ‘Shine Muscat’. Journal of the Japanese Society for Horticultural Science, 83, 122–132.
Toffolatti, S. L., De Lorenzis, G., Costa, A., Maddalena, G., Passera, A., Bonza, M. C., Pindo, M., Stefani, E., Cestaro, A., Casati, P., Failla, O., Bianco, P. A., Maghradze, D., & Quaglino, F. (2018). Unique resistance traits against downy mildew from the center of origin of grapevine (Vitis vinifera). Scientific Reports, 8, 12523.
Vannozzi, A., Dry, I. B., Fasoli, M., Zenoni, S., & Lucchin, M. (2012). Genome-wide analysis of the grapevine stilbene synthase multigenic family: genomic organization and expression profiles upon biotic and abiotic stresses. BMC Plant Biology, 12, 130.
Wang, W., Khalil-Ur-Rehman, M., Feng, J., & Tao, J. (2017). RNA-seq based transcriptomic analysis of CPPU treated grape berries and emission of volatile compounds. Journal of Plant Physiology, 218, 155–166.
Xu, Y., Hou, X., Feng, J., Khalil-Ur-Rehman, M., & Tao, J. (2019). Transcriptome sequencing analyses reveals mechanisms of eliminated russet by applying GA3 and CPPU on ‘Shine Muscat’ grape. Scientia Horticulturae, 250, 94–103.
Yamada, M., & Sato, A. (2016). Advances in table grape breeding in Japan. Breeding Science, 66, 34–45.
Yan, X., Qiao, H., Zhang, X., Guo, C., Wang, M., Wang, Y., & Wang, X. (2017). Analysis of the grape (Vitis vinifera L.) thaumatin-like protein (TLP) gene family and demonstration that TLP29 contributes to disease resistance. Scientific Reports, 7, 4269.
Yin, L., An, Y., Qu, J., Li, X., Zhang, Y., Dry, I., Wu, H., & Lu, J. (2017). Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism. Scientific Reports, 7, 46553.
Yu, Y., Zhang, Y., Yin, L., & Lu, J. (2012). The mode of host resistance to Plasmopara viticola infection of grapevines. Phytopathology, 102, 1094–1101.
Zhu, Y., Li, Y., Zhang, S., Zhang, X., Yao, J., Luo, Q., Sun, F., & Wang, X. (2019). Genome-wide identification and expression analysis reveal the potential function of ethylene responsive factor gene family in response to Botrytis cinerea infection and ovule development in grapes (Vitis vinifera L.). Plant Biology, 21, 571–584.
Acknowledgements
This work was supported in part by the Cabinet Office, Government of Japan, Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for Smart Bio-industry and Agriculture” (funding agency: Bio-oriented Technology Research Advancement Institution, NARO).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This manuscript is original and complies with the ethical rules applicable to this journal.
Conflict of interest
The authors declare that there are no conflicts of interest regarding the publication of this paper.
Human and animal studies
The research did not involve any human participants and/or animals.
Electronic supplementary material
ESM 1
(DOCX 45.1 KB)
Rights and permissions
About this article
Cite this article
Shimizu, T., Suzaki, K. Multiple gene clusters responsive to Plasmopara viticola infection in grapevines. Eur J Plant Pathol 158, 681–691 (2020). https://doi.org/10.1007/s10658-020-02110-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-020-02110-w