Abstract
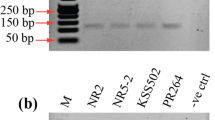
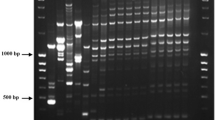
Three new species-specific primer sets were designed to detect C. truncatum, C. gloeosporioides species complex, and C. scovillei, which were reported as the cause of chili anthracnose disease. These new primers were designed based on the sequence data of the nuclear ribosomal DNA (rDNA) region to amplify and identify short size target DNA (<200 bp). The short size target DNA is more efficient for DNA hybridization, and will improve the sensitivity and specificity of DNA assay. The amplicon size obtained for C. gloeosporioides species complex, C. truncatum, and C. scovillei, was 66, 112, and 168 bp, respectively. Each primer set could identify its specific species of Colletotrichum as confirmed by the absence of PCR product with DNA of two other Colletotrichum species and DNA from different fungal genera, indicating high specificity. The sensitivity test was employed, and the results showed that these new primer sets could strongly amplify DNA at the concentration in femto- to picogram levels. Furthermore, these primer sets can be used to amplify and identify Colletotrichum species DNA directly from infected seeds and fruit tissue. The short PCR products obtained by the new primers have the potential to improve the sensitivity in DNA biosensors applications.






Similar content being viewed by others
References
Abang, M. M., Winter, S., Green, K. R., Hoffmann, P., Mignouna, H. D., & Wolf, G. A. (2002). Molecular identification of Colletotrichum gloeosporioidescausing yam anthracnose in Nigeria. Plant Pathology, 51, 63–71. https://doi.org/10.1046/j.0032-0862.2001.00655.x.
Alam, M. Z., Hamim, I., Ali, M. A., & Ashrafuzzaman, M. (2015). Effect of seed treatment on seedling health of chili. Journal of Environmental Science and Natural Resources, 7, 177–181. https://doi.org/10.3329/jesnr.v7i1.22167.
Alexander, P. J., Rajanikanth, G., Bacon, C. D., & Bailey, C. D. (2007). Recovery of plant DNA using a reciprocating and silica-based columns. Molecular Ecology Resources, 7, 5–9. https://doi.org/10.1111/j.1471-8286.2006.01549.x.
Ali, M. E., Raifana Abdul Rashid, N., Bee Abd Hamid, S., Hossain, S. M. A., Asing, A., Hossain, M. A. M., & Zaidul, I. S. M. (2016). Development and validation of short-amplicon length PCR assay for macaques meat detection under complex matrices. International Journal of Food Properties, 20, 231–245. https://doi.org/10.1080/10942912.2016.1154573.
Chen, W., Hoy, J. W., & Schneider, R. W. (1992). Species-specific polymorphism in transcribed ribosomal DNA of five Pythium species. Experimental Mycology, 16, 22–34.
de Silva, D. D., Groenewald, J. Z., Crous, P. W., Ades, P. K., Nasruddin, A., Mongkolporn, O., & Taylor, P. W. J. (2019). Identification, prevalence and pathogenicity of Colletotrichum species causing anthracnose of Capsicum annuum in Asia. International Mycological Association Fungus, 10, 1–32. https://doi.org/10.1186/s43008-019-0001-y.
Dean, R., et al. (2012). The top 10 fungal pathogens in molecular plant pathology. Molecular Plant Pathology, 13, 414–430. https://doi.org/10.1111/j.1364-3703.2011.00783.x.
FAOSTAT (2016) Production data.
Gardes, M., White, T. J., Fortin, J. A., Bruns, T. D., & Taylor, J. W. (1991). Identification of indigenous and introduced symbiotic Fungi in ectomycorrhizae by amplification of nuclear and mitochondrial ribosomal DNA. Canadian Journal of Botany, 69, 180–190.
Garg, R., Loganathan, M., Saha, S., & Roy, B. K. (2014). Chili anthracnose: A review of causal organism, resistance source and mapping of gene. Microbial diversity and biotechnology in food security. New Delhi: Springer. https://doi.org/10.1007/978-81-322-1801-2_53.
Gibriel, A. (2014). Effect of target length on specificity and sensitivity of oligonucleotide microarrays: A comparison between dendrimer and modified PCR based labelling methods. Open Biochemistry Journal, 8, 11–20. https://doi.org/10.2174/1874091X01408010011.
Gutierrez, P., Yepes, M. S., Restrepo, J. F. A., Berrouet, K. V., & Montoya, M. M. (2014). The CD1/CD2 marker for specific detection of Colletotrichum lindemuthianum is an iron transporter pseudogene. Tropical Plant Pathology, 39, 275–283. https://doi.org/10.1590/S1982-56762014000400002.
Hasyim, A., Setiawati, W., & Sutarva, R. (2014). Screening for resistance to anthracnose caused by Colletotrichum acutatum in chili pepper (Capsicum annum L.) in Kediri. East Java AAB Bioflux, 6, 104–118.
Imjit N, Rattanakreetakul C, Pongpisutta R (2013) Polymerase Chain Reaction Based Detection of Chilli Anthracnose Disease I International Conference on Postharvest Pest and Disease Management in Exporting Horticultural Crops - Ppdm2012 973:199–206 https://doi.org/10.17660/ActaHortic.2013.973.27
Johnston, P. R., & Jones, D. (1997). Relationship among Colletotrichum isolates from fruit rots assessed using rDNA sequences. Mycologia, 89, 420–430.
Kamel, A. A.-E. (2003). Bioinformatic tools and guideline for PCR primer design. African Journal of Biotechnology, 2, 91–95. https://doi.org/10.5897/ajb2003.000-1019.
Kamle, M. (2013). A species-specific PCR based assay for rapid detection of mango anthracnose pathogen Colletotrichum gloeosporioides penz. and sacc. Journal of Plant Pathology and Microbiology, 04. https://doi.org/10.4172/2157-7471.1000184.
Kim, C. H., & Park, K. S. (1988). A predictive model of disease progression of red pepper anthracnose. Korean Journal Plant Pathology, 4, 325–331.
Lee, S. P., & Lee, Y. S. (2002). Development of rapid molecular detection marker for Colletotrichum spp. in leaf and fruit tissues of sweet persimmon. Journal of Microbiology and Biotechnology, 12, 989–992.
Liu, W. T., Guo, H., & Wu, J. H. (2007). Effects of target length on the hybridization efficiency and specificity of rRNA-based oligonucleotide microarrays. Applied and Environmental Microbiology, 73, 73–82. https://doi.org/10.1128/AEM.01468-06.
Liu, F., Tang, G., Zheng, X., Li, Y., Sun, X., Qi, X., Zhou, Y., Xu, J., Chen, H., Chang, X., Zhang, S., & Gong, G. (2016). Molecular and phenotypic characterization of Colletotrichum species associated with anthracnose disease in peppers from Sichuan Province, China. Scientific Reports, 6, 32761. https://doi.org/10.1038/srep32761.
Manandhar, J. B., Hartman, G. L., & Wang, T. C. (1995). Anthracnose development on pepper fruits inoculated with Colletotrichum gloeosporioides. Plant Disease, 79, 380–383. https://doi.org/10.1094/Pd-79-0380.
Martinez-Culebras, P. V., Querol, A., Suarez-Fernandez, M. B., Garcia-Lopez, M. D., & Barrio, E. (2003). Phylogenetic relationships among Colletotrichum pathogens of strawberry and design of PCR primers for their identification. Journal of Phytopathology-Phytopathologische Zeitschrift, 151, 135–143. https://doi.org/10.1046/j.1439-0434.2003.00694.x.
Mills, P. R., Sreenivasaprasad, S., & Brown, A. E. (1992). Detection and differentiation of Colletotrichum gloeosporioides isolates using PCR. FEMS Microbiology Letters, 98, 137–143. https://doi.org/10.1111/j.1574-6968.1992.tb05503.x.
Mongkolporn, O., & Taylor, P. W. J. (2018). Chili anthracnose: Colletotrichum taxonomy and pathogenicity. Plant Pathology, 67, 1255–1263. https://doi.org/10.1111/ppa.12850.
Noireung, P., Phoulivong, S., Liu, F., et al. (2012). Novel species of Colletotrichum revealed by morpholgoy and molecular analysis. Cryptogamie Mycologie, 33, 347–362. https://doi.org/10.7872/crym.v33.iss3.2012.347.
Noor, N. M., & Zakaria, L. (2018). Identification and characterization of Colletotrichum spp. associated with chili anthracnose in peninsular Malaysia. European Journal of Plant Pathology, 151, 961–973. https://doi.org/10.1007/s10658-018-1431-x.
Poonpolgul S, Kumphai S (2007) Chili pepper anthracnose in Thailand. Paper presented at the The First International Symposium on Chili Anthracnose, Convention Center, Seoul National University, Korea
Rahman, M., Islam, T., Schwegel, R., & Louws, F. J. (2019). Simultaneous detection of Colletotrichum acutatum and C. gloeosporioides from quiescently infected strawberry foliage by real-time PCR based on high resolution melt curve analysis. American Journal of Plant Sciences, 10, 382–401. https://doi.org/10.4236/ajps.2019.103028.
Raja, H. A., Miller, A. N., Pearce, C. J., & Oberlies, N. H. (2017). Fungal identification using molecular tools: A primer for the natural products research community. Journal of Natural Products, 80, 756–770. https://doi.org/10.1021/acs.jnatprod.6b01085.
Roberts, P. D., Pernezny, K., & Kucharek, T. A. (2001). Anthracnose caused by Colletotrichum sp. on pepper. Gainesville: University of Florida Press.
Safenkova, I. V., Ivanov, A. V., Slutskaya, E. S., Samokhvalov, A. V., Zherdev, A. V., & Dzantiev, B. B. (2020). Key significance of DNA-target size in lateral flow assay coupled with recombinase polymerase amplification. Analytica Chimica Acta, 1102, 109–118. https://doi.org/10.1016/j.aca.2019.12.048.
Saxena, A., Raghuwanshi, R., Gupta, V. K., & Singh, H. B. (2016). Chilli anthracnose: The epidemiology and management. Frontiers in Microbiology, 7. https://doi.org/10.3389/fmicb.2016.01527.
Schoch, C. L., et al. (2012). Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proceedings of the National Academy of Sciences, 109, 6241–6246. https://doi.org/10.1073/pnas.1117018109.
Sreenivasaprasad, S., Brown, A. E., & Mills, P. R. (1992). DNA-sequence variation and interrelationships among Colletotrichum species causing strawberry anthracnose. Physiological and Molecular Plant Pathology, 41, 265–281. https://doi.org/10.1016/0885-5765(92)90026-R.
Srinivasan, M., Kothandaraman, S. V., Vaikuntavasan, P., & Rethinasamy, V. (2014). Development of conventional and real-time PCR protocols for specific and sensitive detection of Colletotrichum capsici in chilli (Capsicum annuum L.). Phytoparasitica, 42, 437–444. https://doi.org/10.1007/s12600-013-0380-3.
Sutton, B. C. (1980). The coelomycetes. Kew: Commeonwealth Mycological Institute.
Sutton, B. C. (1992). The genus Glomerella and its anamorph Colletotrichum. Colletotrichum biology, pathology and control. Wallingford: CAB International.
Suwannarat, S., Steinkellner, S., Songkumarn, P., & Sangchote, S. (2017). Diversity of Colletotrichum spp. isolated from chili pepper fruit exhibiting symptoms of anthracnose in Thailand. Mycological Progress, 16, 677–686. https://doi.org/10.1007/s11557-017-1304-2.
Torres-Calzada, C., Tapia-Tussell, R., Quijano-Ramayo, A., Martin-Mex, R., Rojas-Herrera, R., Higuera-Ciapara, I., & Perez-Brito, D. (2011). A species-specific polymerase chain reaction assay for rapid and sensitive detection of Colletotrichum capsici. Molecular Biotechnology, 49, 48–55. https://doi.org/10.1007/s12033-011-9377-7.
Villafana, R. T., Ramdass, A. C., & Rampersad, S. N. (2019). Development of a new methodology for the detection of Colletotrichum truncatum and Fusarium sp. in bell pepper seed. Phytoparasitica, 47, 543–555. https://doi.org/10.1007/s12600-019-00751-0.
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. InnisMA. PCR Protocols. Academic, San Diego.
Whitelaw-Weckert MA, Curtin SJ, Huang R, Steel CC, Blanchard CL, Roffey PE (2007) Phylogenetic relationships and pathogenicity of Colletotrichum acutatum isolates from grape in subtropical Australia Plant Pathology 56:448–463 https://doi.org/10.1111/j.1365-3059.2007.01569.x.
Widodo, & Hidayat, S. H. (2018). Identification of Colletotrichum species associated with chili anthracnose in Indonesia by morphological characteristics and species-specific primers. Asian Journal of Plant Pathology, 12, 7–15. https://doi.org/10.3923/ajppaj.2018.7.15.
Acknowledgments
We gratefully acknowledge the financial support provided by Agricultural Research Development Agency and the King Mongkut’s University of Technology Thonburi through the “KMUTT 55th Anniversary Commemorative Fund”.
Author information
Authors and Affiliations
Contributions
All authors listed above have made a substantial and direct contribution to the work, and approved it for publication.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical statement
Authors declared that this manuscript has not published elsewhere. All authors read and approved the final version of this manuscript. The authors declare that the present work was developed without any potential conflict of interest, with no human or animal participants.
Rights and permissions
About this article
Cite this article
Osman Abdelrazig, A., Suwannarat, S., Rijiravanich, P. et al. Identification and detection of chili anthracnose using three new species-specific PCR primers. Eur J Plant Pathol 158, 571–582 (2020). https://doi.org/10.1007/s10658-020-02103-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-020-02103-9