Abstract
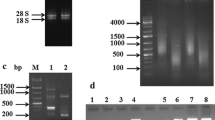
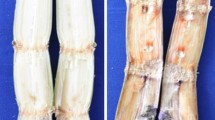
Mustard clubroot, caused by Plasmodiophora brassicae, is one of the most serious diseases affecting Brassica juncea var. tumida Tsen, a mustard plant which is the raw material of a traditional fermented food manufactured in the Chongqing Municipality, P. R. China. We used suppression subtractive hybridization (SSH) to better understand the interaction between B. juncea var. tumida and P. brassicae, and the complex regulation of resistance mechanisms occurring in B. juncea var. tumida after infection by P. brassicae. A total of 1,842 different gene clones were selected from the forward subtracted library (using diseased roots as tester and healthy roots as driver), and 224 positive spots were identified following cDNA array dot blotting. Elimination of polyA tails and sequences shorter than 100 bp generated 196 high-quality gene sequences with an average length of 332 bp. Bioinformatic analysis showed that these 196 sequences represented 173 unigenes, comprising 14 contigs and 159 singlets. Of these, 146 ESTs (84.4 % of the total) were significantly similar to known sequences in plants, the remaining 23 (13.3 % of the total) were of P. brassicae origin. We used quantitative reverse transcription-PCR to analyze the six genes most likely to be involved in disease resistance or the stress response to evaluate the efficiency of SSH, and the results showed that our library data is reliable. Further study of these genes might be helpful for breeding resistance of mustard plants to P. brassicae.


Similar content being viewed by others
References
Agarwal, A., Kaul, V., Faggian, R., Rookes, J. E., Ludwig-Muller, J., & Cahill, D. M. (2011). Analysis of global host gene expression during the primary phase of the Arabidopsis thaliana–Plasmodiophora brassicae interaction. Functional Plant Biology, 38, 462–478.
Ando, S., Asano, T., Tsushima, S., Kamachi, S., Hagio, T., & Tabei, Y. (2005). Changes in gene expression of putative isopentenyltransferase during clubroot development in Chinese cabbage (Brassica rapa L.). Physiological and Molecular Plant Pathology, 67, 59–67.
Belbahri, L., Chevalier, L., Bensaddek, L., Gillet, F., Fliniaux, M. A., Boerjan, W., et al. (2000). Different expression of an S-adenosylmethionine synthetase gene in transgenic tobacco callus modifies alkaloid biosynthesis. Biotechnology and Bioengineering, 69, 11–20.
Brehm, A., Miska, E. A., Mccance, D. J., Reid, J. L., Bannister, A. J., & Kouzarides, T. (1998). Retinoblastoma protein recruits histone deacetylase to repress transcription. Nature, 391, 597–601.
Bulman, S., Siemens, J., Ridgeway, H., Eady, C., & Conner, A. (2006). Identification of genes from the obligate intracellular plant pathogen, Plasmodiophora brassicae. FEMS Microbiology Letters, 264, 198–204.
Burki, F., Kudryavtsev, A., Matz, M. V., Aglyamova, G. V., Bulman, S., Fiers, M., et al. (2010). Evolution of Rhizaria: new insights from phylogenomic analysis of uncultivated protists. BMC Evolutionary Biology, 10, 377.
Cao, T., Srivastava, S., Rahman, M. H., Kav, N. N. V., Hotte, N., Deyholos, M. K., et al. (2008). Proteome-level changes in the roots of Brassica napus as a result of Plasmodiophora brassicae infection. Plant Science, 174, 97–115.
Chiang, P. K., Gordon, R. K., Tal, J., Zeng, G. C., Doctor, B. P., Pardhasaradhi, K., et al. (1996). S-Adenosylmethionine and methylation. The FASEB Journal, 10, 471–480.
Dekhuijzen, H. M. (1979). Electron microscopic studies on the root hairs and cortex of a susceptible and a resistant variety of Brassica campestris infected with Plasmodiophora brassicae. European Journal of Plant Pathology, 85, 1–17.
Delaure, S. L., Van Hemelrijck, W., Bolle, D., Miguel, F. C., Cammue, B. P. A., Coninck, D., et al. (2008). Building up plant defenses by breaking down proteins. Plant Science, 174, 375–385.
Devos, S., Vissenberg, K., Verbelen, J. P., & Prinsen, E. (2005). Infection of Chinese cabbage by Plasmodiophora brassicae leads to a stimulation of plant growth: impacts on cell wall metabolism and hormone balance. The New Phytologist, 166, 241–250. doi:10.1111/j.1469-8137.2004.01304.x.
Devos, S., Laukens, K., Deckers, P., Van-Der-Straeten, D., Beeckman, T., Inze, D., et al. (2006). A hormone and proteome approach to picturing the initial metabolic events during Plasmodiophora brassicae infection on Arabidopsis. Molecular Plant-Microbe Interactions, 19, 1431–1443.
Donald, C., & Porter, I. (2009). Integrated control of clubroot. Plant Growth Regulation, 28, 289–303.
Feng, J., Hwang, S. F., & Strelkov, S. E. (2012). Analysis of expressed sequence tags derived from a compatible Plasmodiophora brassicae–canola interaction. Canadian Journal of Plant Pathology. doi:10.1080/07060661.2012.722128.
Ito, S., Tanaka, S., Miyanaga, C., Takabayashi, S., Yano, S., & Kameya-Iwaki, M. (1996). Constitutive and inducible proteins in the root of Chinese cabbage infected with Plasmodiophora brassicae. Journal of Phytopathology, 144, 89–95.
Jones, J. D., & Dangl, J. L. (2006). The plant immune system. Nature, 444, 323–329.
Kakimoto, T. (2001). Identification of plant cytokinin biosynthesis enzymes as dimethylallyl diphosphate: ATP/ADP isopentenyltransferases. Plant & Cell Physiology, 42, 677–685.
Knaust, A., & Ludwig-Muller, J. (2012). The ethylene signaling pathway is needed to restrict root gall growth in Arabidopsis after Infection with the obligate biotrophic protist Plasmodiophora brassicae. Journal of Plant Growth Regulation. doi:10.1007/s00344-012-9271-y.
Kobelt, P., Siemens, J., & Sacristan, M. D. (2000). Histological characterisation of the incompatible interaction between Arabidopsis thaliana and the obligate biotrophic pathogen Plasmodiophora brassicae. Mycological Research, 104, 220–225.
Kroll, T. K., Lacy, G. H., & Moore, L. D. (1983). A quantitative description of the colonization of susceptible and resistant radish plants by Plasmodiophora brassicae. Journal of Phytopathology, 108, 97–105.
Kuginuki, Y., Yoshikawa, H., & Hirai, M. (1999). Variation in virulence of Plasmodiophora brassicae in Japan tested with clubroot-resistant cultivars of Chinese cabbage (Brassica rapa L. ssp. pekinensis). European Journal of Plant Pathology, 105, 327–332.
Laskowski, M. J., Dreher, K. A., Gehring, M. A., Abel, S., Gensler, A. L., & Sussex, I. M. (2002). FQR1, a novel primary auxin-response gene, encodes a flavin mononucleotide-binding quinone reductase. Plant Physiology, 128, 578–590.
Ludwig-Muller, J. (2008). Glucosinolates and the clubroot disease: defense compounds or auxin precursors? Phytochemistry Reviews, 8, 135–148.
Ludwig-Muller, J., & Schuller, A. (2008). What can we learn from clubroots: alterations in host roots and hormone homeostasis caused by Plasmodiophora brassicae. European Journal of Plant Pathology, 121, 291–302.
Ludwig-Muller, J., Thermann, P., Pieper, K., & Hilgenberg, W. (1994). Peroxidase and chitinase isoenzyme activities during root infection of Chinese cabbage with Plasmodiophora brassicae. Physiologia Plantarum, 90, 661–670.
Ludwig-Muller, J., Prinsen, E., Rolfe, S. A., & Scholes, J. D. (2009). Metabolism and plant hormone action during clubroot disease. Journal of Plant Growth Regulation, 28, 229–244.
Moldenhauer, J., Moerschbacher, B. M., & Van der Westhuizen, A. J. (2006). Histological investigation of stripe rust (Puccinia striiformis f.sp tritici) development in resistant and susceptible wheat cultivars. Plant Pathology, 55, 469–474.
Murashige, T., & Skoog, F. (1962). A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiologia Plantarum, 15, 473–497.
Nagaoka, T., Doullah, M. A. U., Matsumoto, S., Kawasaki, S., Ishikawa, T., Hori, H., et al. (2010). Identification of QTLs that control clubroot resistance in Brassica oleracea and comparative analysis of clubroot resistance genes between B. rapa and B. oleracea. Theoretical and Applied Genetics, 120, 1335–1346.
Paz-Ares, J., Ghosal, D., Wienand, U., Peterson, P., & Saedler, H. (1987). The regulatory c1 locus of Zea mays encodes a protein with homology to myb oncogene products and with structural similarities to transcriptional activators. The EMBO Journal, 6, 3553–3558.
Ramonell, K., Berrocal-Lobo, M., Koh, S., Wan, J., Edwards, H., Stacey, G., et al. (2005). Loss-of-function mutations in chitin responsive genes show increased susceptibility to the powdery mildew pathogen Erysiphe cichoracearum. Plant Physiology, 138, 1027–1036.
Rizhsky, L., Liang, H., & Mittler, R. (2002). The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiology, 130, 1143–1151.
Rizhsky, L., Liang, H., Shuman, J., Shulaev, V., Davletova, S., & Mittler, R. (2004). When defense pathways collide. The response of Arabidopsis to a combination of drought and heat stress. Plant Physiology, 134, 1683–1696.
Siemens, J., Keller, I., Sarx, J., Kunz, S., Schuller, A., Nagel, W., et al. (2006). Transcriptome analysis of Arabidopsis clubroots and disease resistance of cytokinin oxidase/dehydrogenase gene overexpressing plants indicate a key role for cytokinin in disease development. Molecular Plant-Microbe Interactions, 19, 480–494. doi:10.1094/MPMI-19-0480.
Siemens, J., Bulman, S., Rehn, F., & Sundelin, T. (2009). Molecular biology of Plasmodiophora brassicae. Journal of Plant Growth Regulation, 28, 245–251.
Smolen, G., & Bender, J. (2002). Arabidopsis cytochrome P450 cyp83B1 mutations activate the tryptophan biosynthetic pathway. Genetics, 160, 323–332.
Sundelin, T., Jensen, D., & Lubeck, M. (2011). Identification of expressed genes during infection of Chinese cabbage (Brassica rapa subsp. pekinensis) by Plasmodiophora brassicae. Journal of Eukaryotic Microbiology, 58, 310–314.
Terasaka, K., Blakeslee, J. J., Titapiwatanakun, B., et al. (2005). PGP4, an ATP Binding Cassette P-Glycoprotein, Catalyzes Auxin Transport in Arabidopsis thaliana Roots. The Plant Cell, 17, 2922–2939.
Torres, M. A., & Dangl, J. L. (2005). Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Current Opinion in Plant Biology, 8, 397–403.
Webster, M. A., & Dixon, G. R. (1991). Calcium, pH and inoculum concentration influencing colonization by Plasmodiophora brassicae. Mycological Research, 95, 65–73.
Woodward, A. W., & Bartel, B. (2005). Auxin: regulation, action and interaction. Annals of Botany, 95, 707–735.
Xiao, C., & Guo, X. (2002). Biological characteristic of Plasmodiophora Brassicae. Mycosystema, 21, 597–603.
Zhao, J., Davis, L. C., & Verpoorte, R. (2005). Elicitor signal transduction leading to production of plant secondary metabolites. Biotechnology Advances, 23, 283–333.
Acknowledgments
This research was financially supported by the Natural Science Foundation of Chongqing Science and Technology Commission (CSTC, 2008BB1370). We also gratefully appreciate the support from Chongqing Fuling Institute of Agricultural Science.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 2
Differentially expressed genes in the SSH library (DOC 169 kb) (DOC 169 kb)
Rights and permissions
About this article
Cite this article
Luo, Y., Yin, Y., Liu, Y. et al. Identification of differentially expressed genes in Brassica juncea var. tumida Tsen following infection by Plasmodiophora brassicae . Eur J Plant Pathol 137, 43–53 (2013). https://doi.org/10.1007/s10658-013-0215-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-013-0215-6